

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146549.1 - phase: 0
(476 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
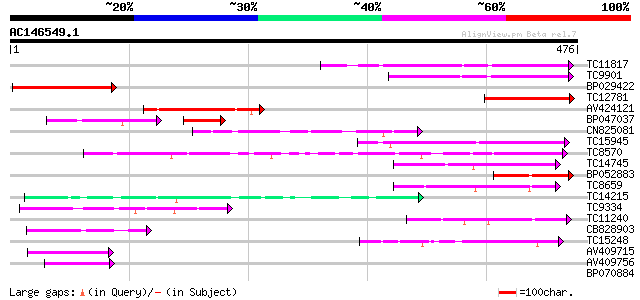
Score E
Sequences producing significant alignments: (bits) Value
TC11817 weakly similar to UP|Q940Z5 (Q940Z5) AT5g39050/MXF12_60,... 151 2e-37
TC9901 weakly similar to UP|ANTA_GENTR (Q9ZWR8) Anthocyanin 5-ar... 132 1e-31
BP029422 124 3e-29
TC12781 weakly similar to UP|Q940Z5 (Q940Z5) AT5g39050/MXF12_60,... 115 2e-26
AV424121 99 2e-21
BP047037 75 2e-20
CN825081 91 4e-19
TC15945 similar to UP|ANTA_GENTR (Q9ZWR8) Anthocyanin 5-aromatic... 89 2e-18
TC8570 similar to UP|Q9FLM5 (Q9FLM5) N-hydroxycinnamoyl/benzoylt... 82 1e-16
TC14745 similar to UP|Q9LFB5 (Q9LFB5) Anthranilate N-benzoyltran... 58 3e-09
BP052883 58 3e-09
TC8659 similar to UP|Q8VWP8 (Q8VWP8) Acyltransferase-like protei... 57 9e-09
TC14215 similar to UP|Q9FYM1 (Q9FYM1) F21J9.8, partial (8%) 56 1e-08
TC9334 similar to UP|Q9MBD4 (Q9MBD4) Acyltransferase homolog, pa... 54 4e-08
TC11240 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transfera... 45 3e-05
CB828903 45 3e-05
TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A:... 41 4e-04
AV409715 40 6e-04
AV409756 40 8e-04
BP070884 40 0.001
>TC11817 weakly similar to UP|Q940Z5 (Q940Z5) AT5g39050/MXF12_60, partial
(12%)
Length = 950
Score = 151 bits (382), Expect = 2e-37
Identities = 85/212 (40%), Positives = 117/212 (55%)
Frame = +2
Query: 262 LTREDLNKIKQMVLSKWELLDTNELTSKPPTLSSFVLTCAYSLVCLAKAIHGVEKEKEKF 321
LT + K+KQ LSK E D LSSF +TCAY L C KA + + +
Sbjct: 2 LTPSHIQKLKQYALSKLETKDR---------LSSFAVTCAYVLACAVKA---EQPQGDTV 145
Query: 322 GFAFTVDCRARLEPPLPNNYFGNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMINEK 381
F+VDCR+RL+PP+ NYFGNC+ G + + + +DG + ++E + + +
Sbjct: 146 MMVFSVDCRSRLDPPIVPNYFGNCIGGQMFGVRIKTLLEKDGFISALEEMNEALNRVKD- 322
Query: 382 GVLDGVSDMFDKFASLASEKLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADRGLTIAL 441
GVL G +M L +L +AGS RF VY IDFGWGRP KV+I S DR +L
Sbjct: 323 GVLKGAKEMASNMEKLPEGRL--YSIAGSPRFEVYSIDFGWGRPKKVDITSIDRTGAFSL 496
Query: 442 AESKDGKGGIEVGLVLNNHVMNLFRTLFVEGL 473
+ES++ GGIE+G+VL H M F FV+ L
Sbjct: 497 SESRNNNGGIEIGMVLKKHEMEAFAASFVQDL 592
>TC9901 weakly similar to UP|ANTA_GENTR (Q9ZWR8) Anthocyanin 5-aromatic
acyltransferase (5AT) , partial (6%)
Length = 599
Score = 132 bits (332), Expect = 1e-31
Identities = 65/155 (41%), Positives = 93/155 (59%)
Frame = +3
Query: 319 EKFGFAFTVDCRARLEPPLPNNYFGNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMI 378
++ F F+VDCR RLEPP+ YFGNC+ HL + + + +DG I +++ +
Sbjct: 3 KRVAFVFSVDCRTRLEPPIKPTYFGNCIMPHLAVAETGEVLGDDGFINALVVISDELSEL 182
Query: 379 NEKGVLDGVSDMFDKFASLASEKLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADRGLT 438
E GV++G D K S+A +K M AGS RF V+ +DFGWGRP KV++ S DR
Sbjct: 183 -ESGVMNGAEDWISKIQSVAGDK--MFSTAGSPRFEVFGVDFGWGRPKKVDVTSVDRTGA 353
Query: 439 IALAESKDGKGGIEVGLVLNNHVMNLFRTLFVEGL 473
+L++S+D GGIE+GL LN M F +F +GL
Sbjct: 354 FSLSDSRDIHGGIEIGLALNKSQMEAFAEVFAQGL 458
>BP029422
Length = 498
Score = 124 bits (312), Expect = 3e-29
Identities = 56/87 (64%), Positives = 68/87 (77%)
Frame = +1
Query: 3 SNNNNNVKIHEHTKVFPPSSTQKTTTPLTYFDIFWLRFHPVERVFFYALPNSHSHPSFFF 62
+NNNN+++IHEH KV PPSS + PL +FD+ WLRFHPVER+FFY L N S PSFFF
Sbjct: 238 NNNNNSIRIHEHCKVAPPSSATEACLPLIFFDLLWLRFHPVERIFFYNLQNPESDPSFFF 417
Query: 63 KKLVPILKSSLSLTLKDFLPLAGNIVW 89
K+VP LK+SLSLTL+ F LAGN+VW
Sbjct: 418 HKVVPKLKNSLSLTLQHFPALAGNVVW 498
>TC12781 weakly similar to UP|Q940Z5 (Q940Z5) AT5g39050/MXF12_60, partial
(12%)
Length = 463
Score = 115 bits (287), Expect = 2e-26
Identities = 55/76 (72%), Positives = 61/76 (79%)
Frame = +1
Query: 399 SEKLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADRGLTIALAESKDGKGGIEVGLVLN 458
SE + +GVAGSNRFGVY DFGWGRP KVEI S DR LTI +AESKDGKGG+EVGLVL+
Sbjct: 1 SEGVVSIGVAGSNRFGVYGTDFGWGRPKKVEITSIDRNLTIGMAESKDGKGGVEVGLVLD 180
Query: 459 NHVMNLFRTLFVEGLC 474
HVM LF T F+ GLC
Sbjct: 181 KHVMGLFGTFFLAGLC 228
>AV424121
Length = 321
Score = 99.0 bits (245), Expect = 2e-21
Identities = 49/105 (46%), Positives = 75/105 (70%), Gaps = 3/105 (2%)
Frame = +2
Query: 113 SDVDFNHVIENSPHDASLSRCFVPHLESTNSFASIISVQITLFPESGFSIGISTHHAALD 172
S+VDFN++ + ++A +PHL +++ AS+I++Q+TLFP SGF IG+++HHA LD
Sbjct: 5 SNVDFNYLCSDL-YEAEKRYHLIPHLTTSHEKASLIALQVTLFPNSGFCIGLTSHHAGLD 181
Query: 173 GKSSTMFIKAWAYLCNKTIETEELPTLLP---ELKPLLDREIIKD 214
GKSST F+K+WA+ C+K ++ E L +L P +L P DR +IKD
Sbjct: 182 GKSSTSFMKSWAFSCSK-LQKEGLSSLFPLPEDLTPFFDRSVIKD 313
>BP047037
Length = 530
Score = 75.1 bits (183), Expect(2) = 2e-20
Identities = 36/98 (36%), Positives = 56/98 (56%), Gaps = 2/98 (2%)
Frame = +1
Query: 32 YFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPILKSSLSLTLKDFLPLAGNIVWPL 91
+ DI W PV+R+FFY P+ H F + +P LK SL++TL+ F P N++ P
Sbjct: 67 FLDIPWFYSPPVQRIFFYEFPHPTQH---FLQVAIPTLKHSLTITLQHFFPFVSNLIVPP 237
Query: 92 ES--QEPIIQYTPNDGVSLIIAESDVDFNHVIENSPHD 127
+ P I+Y D + +AES+ DF+H++ +SP D
Sbjct: 238 QPYVDTPYIRYLHGDSLPFTVAESNSDFDHLVSDSPQD 351
Score = 40.8 bits (94), Expect(2) = 2e-20
Identities = 16/35 (45%), Positives = 25/35 (70%)
Frame = +3
Query: 147 IISVQITLFPESGFSIGISTHHAALDGKSSTMFIK 181
++++Q+T+ P SGFSI ++ HH DG+S FIK
Sbjct: 426 LMAIQVTVLPNSGFSISLTFHHIVADGRSLHHFIK 530
>CN825081
Length = 618
Score = 90.9 bits (224), Expect = 4e-19
Identities = 63/197 (31%), Positives = 101/197 (50%), Gaps = 4/197 (2%)
Frame = +1
Query: 154 LFPESGFSIGISTHHAALDGKSSTMFIKAWAYLCNKTIETEEL-PTLLPELKPLLDREII 212
+FP G I I+ H +D + F+K W+ +C +T + PT L P DRE +
Sbjct: 22 VFPNHGICISITYCHI-MDDFCCSQFMKTWSSIC----QTGGVHPTFLQNSPPCFDREAL 186
Query: 213 KDN-GLGDKFTKNWTEIITMMFPNEKGNERSLKILPFEPKLEDYVRSTFKLTREDLNKIK 271
+D GL F +++ E E+ N + I P +DYV++T +RE + +K
Sbjct: 187 QDTKGLEGVFLRDYFE--------ERSNWKHKFIAPTSDH-DDYVKATIVFSREKIEGLK 339
Query: 272 QMVLSKWELLDTNELTSKPPTLSSFVLTCAYSLVCLAKAIH--GVEKEKEKFGFAFTVDC 329
+ VL++W+ +S FV+TCA+ L K + G ++++E++ F F DC
Sbjct: 340 KWVLNQWK-----------KNVSKFVVTCAFVWASLVKTRYRNGDDEQEEEY-FRFAADC 483
Query: 330 RARLEPPLPNNYFGNCV 346
R RLE P+P YFGNC+
Sbjct: 484 RDRLEYPIPEGYFGNCL 534
>TC15945 similar to UP|ANTA_GENTR (Q9ZWR8) Anthocyanin 5-aromatic
acyltransferase (5AT) , partial (5%)
Length = 923
Score = 88.6 bits (218), Expect = 2e-18
Identities = 57/188 (30%), Positives = 90/188 (47%), Gaps = 10/188 (5%)
Frame = +1
Query: 293 LSSFVLTCAYSLVCLAKAIHGVEKEK----------EKFGFAFTVDCRARLEPPLPNNYF 342
+S+FV+ C+ V + K +E+ K E F F DCR R E P+ YF
Sbjct: 46 MSTFVVACSLVWVSMIK----LEQRKGDYCVAKGSDELCYFLFLADCRGRPELSFPSTYF 213
Query: 343 GNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMINEKGVLDGVSDMFDKFASLASEKL 402
GNC+ V + + E+G+ VA + +I+ + + + + D + L
Sbjct: 214 GNCLAFCTVALNKDEVVGENGLIEVANAVEREIRNWKSDPLQNAETSISD-YRELLKPGK 390
Query: 403 EMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADRGLTIALAESKDGKGGIEVGLVLNNHVM 462
++ + GS VYE DFGWG+P K E V D +++++ +D GG+EVGL L M
Sbjct: 391 SLLTIGGSPSLAVYETDFGWGKPKKSESVHLDTFRIVSMSDCRDKDGGMEVGLALAKVRM 570
Query: 463 NLFRTLFV 470
N F + V
Sbjct: 571 NDFINILV 594
>TC8570 similar to UP|Q9FLM5 (Q9FLM5)
N-hydroxycinnamoyl/benzoyltransferase-like protein,
partial (91%)
Length = 1551
Score = 82.4 bits (202), Expect = 1e-16
Identities = 96/419 (22%), Positives = 173/419 (40%), Gaps = 13/419 (3%)
Frame = +1
Query: 63 KKLVPILKSSLSLTLKDFLPLAGNIVWPLESQEPIIQYTPNDGVSLIIAESDVDFNHVIE 122
+K ++KS+L L + PLAG + + + +I +G + AE++ + +
Sbjct: 232 EKAGEVVKSALKKVLVHYYPLAGRL--SISPEGKLIVECTGEGALFVEAEANCSLEEIGD 405
Query: 123 -NSPHDASLSRCF--VPHLESTNSFASIISVQITLFPESGFSIGISTHHAALDGKSSTMF 179
P +L + +P + +++ Q+T F GFS+G+ +H DG + F
Sbjct: 406 ITKPDPGTLGKLVYDIPGAKHILQMPPLVA-QVTKFKCGGFSLGLCMNHCMFDGIGAMEF 582
Query: 180 IKAWAYLCNKTIETEELPTLLPELKPLLDREIIKDNGLG--DKFTKNWTEIITMMFPNEK 237
+ +W LP +P P+LDR I+K + + + +I +K
Sbjct: 583 VNSWGE------AARALPLSIP---PILDRSILKARNPPKIEHLHQEFADI------EDK 717
Query: 238 GNERSLKILPFEPKLEDYVRSTFKLTREDLNKIKQMVLSKWELLDTNELTSKPPTLSSFV 297
N +L +E ++ ++ D K+KQ+ K + ++ L S ++F
Sbjct: 718 SNTNTL----YEDEM------LYRSFCFDPEKLKQL---KMKAMEDGALES----CTTFE 846
Query: 298 LTCAYSLVCLAKAIHGVEKEKEKFGFAFTVDCRARLEPPLPNNYFGN--------CVWGH 349
+ A+ + KA+ + +++ K FA VD R PPLP YFGN C G
Sbjct: 847 VLSAFVWIARTKALKLLPEQETKLLFA--VDGRKNFTPPLPKGYFGNGIVLTNSVCQAGE 1020
Query: 350 LVDTKPLDYINEDGVFLVAKCIHEKIKMINEKGVLDGVSDMFDKFASLASEKLEMMGVAG 409
L + K + + I + +KM+ + + + D F+ + S ++ +
Sbjct: 1021LSEKK---------LSHAVRLIQDAVKMVTDSYMRSAI-DYFEVTRARPSLACTLL-ITT 1167
Query: 410 SNRFGVYEIDFGWGRPTKVEIVSADRGLTIALAESKDGKGGIEVGLVLNNHVMNLFRTL 468
+R G + DFGWG P VS I + I V L L VM +F+ L
Sbjct: 1168WSRLGFHTTDFGWGEPVLSGPVSLPEKEVILFLSHGQERRNINVLLGLPASVMKIFQDL 1344
>TC14745 similar to UP|Q9LFB5 (Q9LFB5) Anthranilate
N-benzoyltransferase-like protein (AT5g01210/F7J8_190),
partial (37%)
Length = 842
Score = 58.2 bits (139), Expect = 3e-09
Identities = 40/143 (27%), Positives = 67/143 (45%), Gaps = 3/143 (2%)
Frame = +1
Query: 323 FAFTVDCRARLEPPLPNNYFGNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMINEKG 382
F V+CR RLEP + YFGN + D ++ D F A +H+ + ++
Sbjct: 52 FRMAVNCRHRLEPKMDPFYFGNAIQSIPTVASAGDILSRDLRFC-ADLLHQNVVAHDDAT 228
Query: 383 VLDGVS--DMFDKFASLASEKLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADR-GLTI 439
V G+ + + L + M+ + S RF +Y+ DFGWGRP + A++ I
Sbjct: 229 VRCGIEGWEKAPRLFPLGNFDGAMITMGSSPRFPMYDNDFGWGRPLAIRSGKANKFDGKI 408
Query: 440 ALAESKDGKGGIEVGLVLNNHVM 462
+ +DG G +++ +VL M
Sbjct: 409 SAFPGRDGNGSVDLEVVLAPETM 477
>BP052883
Length = 458
Score = 58.2 bits (139), Expect = 3e-09
Identities = 31/67 (46%), Positives = 42/67 (62%)
Frame = -1
Query: 407 VAGSNRFGVYEIDFGWGRPTKVEIVSADRGLTIALAESKDGKGGIEVGLVLNNHVMNLFR 466
V GS FGVY DFG+GRP KVE+V + +G ++LAE D GG+E GLV + F
Sbjct: 377 VTGSPXFGVYGTDFGFGRPVKVEMVHSFKG--VSLAECGDEDGGLEFGLVFRKGEFDCFV 204
Query: 467 TLFVEGL 473
++ +GL
Sbjct: 203 SVIEQGL 183
>TC8659 similar to UP|Q8VWP8 (Q8VWP8) Acyltransferase-like protein
(Fragment), partial (42%)
Length = 866
Score = 56.6 bits (135), Expect = 9e-09
Identities = 43/144 (29%), Positives = 64/144 (43%), Gaps = 4/144 (2%)
Frame = +3
Query: 323 FAFTVDCRARLEPPLPNNYFGNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMINEKG 382
F VDCR R++PP+P YFGN + + + ++ A I + I+ + K
Sbjct: 84 FTVFVDCRKRVDPPMPEAYFGNLIQA-IFTVTAVGLLSAHPPQFGATMIQKAIEAHDAKA 260
Query: 383 VLDGVSD--MFDKFASLASEKLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVSADR--GLT 438
+ + D K + + V S RF VY+IDFGWG+P V ++ G+
Sbjct: 261 IDERNKDWESAPKIFQFKDAGINCVTVGSSPRFKVYDIDFGWGKPEIVRSGPNNKFDGM- 437
Query: 439 IALAESKDGKGGIEVGLVLNNHVM 462
I L K G I+V L L M
Sbjct: 438 IYLYPGKSGGRSIDVELTLEPEAM 509
>TC14215 similar to UP|Q9FYM1 (Q9FYM1) F21J9.8, partial (8%)
Length = 1096
Score = 55.8 bits (133), Expect = 1e-08
Identities = 80/338 (23%), Positives = 131/338 (38%), Gaps = 3/338 (0%)
Frame = +1
Query: 13 EHTKVFPPSSTQKTTTPLTYFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPILKSS 72
E K P+ PL++ D R + +F++ PN S ++ LK+S
Sbjct: 118 EIIKPSTPTPPHLRIYPLSFIDNMMYRNYIPVALFYH--PNESDKQS-----IISNLKNS 276
Query: 73 LSLTLKDFLPLAGNIVWPLESQEPIIQYTPNDGVSLIIAESDVDFNHVIENSPHDASLSR 132
LS L + P AG + ++ + ++GV + E + + +++N P++ L
Sbjct: 277 LSEVLTRYYPFAGRL------RDQLSIECNDEGVPFRVTEIKGEQSTILQN-PNETLLRL 435
Query: 133 CFVPHL--ESTNSFASIISVQITLFPESGFSIGISTHHAALDGKSSTMFIKAWAYLCNKT 190
F L + T SI +VQI F G +I H D ++ F+ WA +
Sbjct: 436 LFPDKLPWKVTEYDESIAAVQINFFSCGGLAITACVCHKMGDATTTLNFVNDWAAM---- 603
Query: 191 IETEELPTLLPELK-PLLDREIIKDNGLGDKFTKNWTEIITMMFPNEKGNERSLKILPFE 249
T+E L +L PLL+ + + K + EI+ KGN+
Sbjct: 604 --TKEKGEALTQLSLPLLNGGV---SVFPHKDMPCFPEIVF-----GKGNK--------- 726
Query: 250 PKLEDYVRSTFKLTREDLNKIKQMVLSKWELLDTNELTSKPPTLSSFVLTCAYSLVCLAK 309
+ V F +N +K+MV S + PT + Y +
Sbjct: 727 ---SNAVCRRFVFPPSKINSLKEMVTSH---------GVQNPTRVEVITAWIY-----MR 855
Query: 310 AIHGVEKEKEKFGFAFTVDCRARLEPPLPNNYFGNCVW 347
A+H + +K F V R R+ PPLPN GN W
Sbjct: 856 AVHALGLTFDKTSFRQVVSLRRRMTPPLPNKSVGNMFW 969
>TC9334 similar to UP|Q9MBD4 (Q9MBD4) Acyltransferase homolog, partial
(47%)
Length = 817
Score = 54.3 bits (129), Expect = 4e-08
Identities = 48/185 (25%), Positives = 87/185 (46%), Gaps = 6/185 (3%)
Frame = +2
Query: 9 VKIHEHTKVFPPSSTQKTTTPLTYFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPI 68
+K+ + V P K L FD+ +L F+ +++ FY F+ LV
Sbjct: 62 LKVTNKSHVKPEEKIGKKEYQLVTFDLPYLAFYYNQKLLFYKGEG--------FEGLVEK 217
Query: 69 LKSSLSLTLKDFLPLAGNIVWPLESQEPIIQYTPND---GVSLIIAESDVDFNHVIENSP 125
LK L++ LK+F L G I + +E + + +D GV +I A +D +
Sbjct: 218 LKDGLAVVLKEFHQLGGKIG---KDEEGVFRVEFDDDLPGVEVIEAVAD-EVAVADLTLT 385
Query: 126 HDASLSRCFVPH---LESTNSFASIISVQITLFPESGFSIGISTHHAALDGKSSTMFIKA 182
+S+ + +P+ L +++VQ T + G ++G++ +HA LDG S+ F+ +
Sbjct: 386 ESSSILKELIPYSGVLNLEGMHRPLLAVQFTKLKD-GLAMGLAFNHAVLDGTSTWHFMSS 562
Query: 183 WAYLC 187
WA +C
Sbjct: 563 WAEIC 577
>TC11240 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transferase, partial
(31%)
Length = 528
Score = 44.7 bits (104), Expect = 3e-05
Identities = 39/145 (26%), Positives = 67/145 (45%), Gaps = 7/145 (4%)
Frame = +1
Query: 334 EPPLPNNYFGNCVWGHLVDTKPLDYINEDGVFLVAKCIHEKI-KMINE--KGVLDGVSDM 390
+PPLP YFGN ++ D +++ ++ A IH+ + +M NE + LD +
Sbjct: 25 KPPLPQGYFGNVIFTATPIAMAGDLMSKP-IWYAASRIHDALLRMDNEYLRSALDFLELQ 201
Query: 391 FDKFASLASE---KLEMMGVAGSNRFGVYEIDFGWGRPTKVEIVS-ADRGLTIALAESKD 446
D A + + +G+ R +++ DFGWGRP + A GL+ + S
Sbjct: 202 PDLRALVRGAHTFRCPNLGITSWVRLPIHDADFGWGRPIFMGPGGIAYEGLSF-IIPSPV 378
Query: 447 GKGGIEVGLVLNNHVMNLFRTLFVE 471
G + V + L M +F+ LF +
Sbjct: 379 NDGSLSVAIALQPEQMKVFQELFYD 453
>CB828903
Length = 510
Score = 44.7 bits (104), Expect = 3e-05
Identities = 34/105 (32%), Positives = 48/105 (45%)
Frame = +3
Query: 15 TKVFPPSSTQKTTTPLTYFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPILKSSLS 74
T V P ++ L+ D+ L H +++ + P S S PS L+P+LKSSLS
Sbjct: 111 TTVIPDQASTLGNLKLSVSDLPMLSCHYIQKGCLFTKPPSSSLPSH---TLIPLLKSSLS 281
Query: 75 LTLKDFLPLAGNIVWPLESQEPIIQYTPNDGVSLIIAESDVDFNH 119
TL +F PLAG + N V L ++ VDF H
Sbjct: 282 KTLSNFPPLAGRLT-----------TDENGYVHLTCNDAGVDFLH 383
>TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A: benzyl
alcohol benzoyl transferase, partial (52%)
Length = 1085
Score = 41.2 bits (95), Expect = 4e-04
Identities = 44/185 (23%), Positives = 76/185 (40%), Gaps = 13/185 (7%)
Frame = +1
Query: 294 SSFVLTCAYSLVCLAKAIHGVEKEKEKFGFAFTVDCRARLEPPLPNNYFGNC-------- 345
+++ + +Y C KA+ ++ +E T D R + PP P Y+GNC
Sbjct: 352 TTYEVLTSYIWRCRTKALQ-LDPSQEVRMMCIT-DARGKFNPPFPTGYYGNCFAFPAAVA 525
Query: 346 VWGHLVDTKPLDYINEDGVFLVAKCIHEKIKMINEKGVLDGVSDMFDKFASLASEKLEMM 405
G L + KPL E V L+ K E + + ++D+ +
Sbjct: 526 TAGELCE-KPL----EHAVRLIKKASGEM-----SEEYMHSLADLMVTEGKPLFTVVRSC 675
Query: 406 GVAGSNRFGVYEIDFGWGRPTKVEIVSADRGLTIAL-----AESKDGKGGIEVGLVLNNH 460
V + G E+DFGWG+ + A G A+ +++ G+ GI V + L
Sbjct: 676 VVLDTTYAGFRELDFGWGKAVYGGLAQAGAGAFPAVNFHVPSQNAQGEEGILVLVCLPAK 855
Query: 461 VMNLF 465
+M++F
Sbjct: 856 IMSVF 870
>AV409715
Length = 311
Score = 40.4 bits (93), Expect = 6e-04
Identities = 24/72 (33%), Positives = 33/72 (45%)
Frame = +1
Query: 16 KVFPPSSTQKTTTPLTYFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPILKSSLSL 75
K + P LT +DI L H +++ + P S + F L+ LK SLSL
Sbjct: 94 KPYSPIEDSNQICYLTPWDIALLSVHYIQKGLLFKKPESIDNQQDFMNNLLDNLKHSLSL 273
Query: 76 TLKDFLPLAGNI 87
L F PLAG +
Sbjct: 274 ALDHFYPLAGRL 309
>AV409756
Length = 422
Score = 40.0 bits (92), Expect = 8e-04
Identities = 23/59 (38%), Positives = 31/59 (51%)
Frame = +1
Query: 30 LTYFDIFWLRFHPVERVFFYALPNSHSHPSFFFKKLVPILKSSLSLTLKDFLPLAGNIV 88
LT +DI L H +++ + P S + F L+ LK SLSL L F PLAG +V
Sbjct: 178 LTPWDIAMLSVHYIQKGLLFKNPESIENQPDFMNNLLDNLKHSLSLALVHFYPLAGRLV 354
>BP070884
Length = 493
Score = 39.7 bits (91), Expect = 0.001
Identities = 21/54 (38%), Positives = 30/54 (54%)
Frame = -1
Query: 412 RFGVYEIDFGWGRPTKVEIVSADRGLTIALAESKDGKGGIEVGLVLNNHVMNLF 465
RF +YE+DFGWG+P V +I L +++DG GIE + + M LF
Sbjct: 325 RFPMYEVDFGWGKPIWVTTSECPAKNSIILMDTRDG-DGIEAIVNMEEKDMTLF 167
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,043,783
Number of Sequences: 28460
Number of extensions: 132530
Number of successful extensions: 763
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 742
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 748
length of query: 476
length of database: 4,897,600
effective HSP length: 94
effective length of query: 382
effective length of database: 2,222,360
effective search space: 848941520
effective search space used: 848941520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146549.1


