

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.3 - phase: 0
(670 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
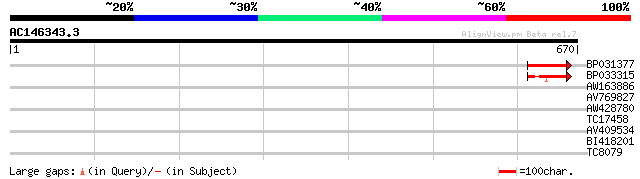
Score E
Sequences producing significant alignments: (bits) Value
BP031377 85 4e-17
BP033315 56 2e-08
AW163886 40 0.001
AV769827 35 0.039
AW428780 33 0.11
TC17458 similar to UP|Q93X17 (Q93X17) Snakin2 precursor, partial... 31 0.56
AV409534 30 1.3
BI418201 28 6.2
TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete 27 8.2
>BP031377
Length = 401
Score = 84.7 bits (208), Expect = 4e-17
Identities = 41/52 (78%), Positives = 44/52 (83%)
Frame = -3
Query: 613 DFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTNTKASILVVQQQ 664
DFIIVGRRNG+KS T L +WTEY ELGV+GDLLASPD TKASILVVQQQ
Sbjct: 390 DFIIVGRRNGVKSEMTAGLENWTEYTELGVIGDLLASPDMETKASILVVQQQ 235
>BP033315
Length = 433
Score = 56.2 bits (134), Expect = 2e-08
Identities = 31/57 (54%), Positives = 39/57 (68%), Gaps = 5/57 (8%)
Frame = -1
Query: 613 DFIIVGRRNGIKSPQTQALA-----SWTEYPELGVLGDLLASPDTNTKASILVVQQQ 664
D ++VGR + PQT ++ W+E PELG +GD+LASPD TKASILVVQQQ
Sbjct: 352 DLVMVGREH----PQTPSVLLKGHEKWSECPELGTIGDMLASPDFVTKASILVVQQQ 194
>AW163886
Length = 177
Score = 40.4 bits (93), Expect = 0.001
Identities = 20/55 (36%), Positives = 36/55 (65%)
Frame = +1
Query: 349 SELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHR 403
S+L++++CI PS+I I +++ TS++ L + I+HL+EL RSS + + R
Sbjct: 13 SQLRVMACIHGPSNIPSIIGLIESTCGTSNSLLKLFIMHLVELTERSSSIMMVQR 177
>AV769827
Length = 438
Score = 35.0 bits (79), Expect = 0.039
Identities = 20/53 (37%), Positives = 31/53 (57%), Gaps = 1/53 (1%)
Frame = -2
Query: 612 HDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTN-TKASILVVQQ 663
+D IIVG+ + + EY ELG +GD+LAS + ++S+LV+QQ
Sbjct: 374 YDLIIVGKGRFPSTMVAELAERHAEYAELGPVGDILASSTGHKMRSSVLVIQQ 216
>AW428780
Length = 330
Score = 33.5 bits (75), Expect = 0.11
Identities = 20/54 (37%), Positives = 30/54 (55%), Gaps = 2/54 (3%)
Frame = +2
Query: 611 QHDFIIVGRRNGI--KSPQTQALASWTEYPELGVLGDLLASPDTNTKASILVVQ 662
+++FIIVG+ I + S E+ ELG +GDLL S +S+LV+Q
Sbjct: 86 EYEFIIVGKGQQILESTMMLNIQDSRLEHAELGPVGDLLTSSGQGITSSVLVIQ 247
>TC17458 similar to UP|Q93X17 (Q93X17) Snakin2 precursor, partial (55%)
Length = 309
Score = 31.2 bits (69), Expect = 0.56
Identities = 16/47 (34%), Positives = 26/47 (55%)
Frame = +3
Query: 511 PFNHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVST 557
PF+H+ +S ++M+ + R + +K +KE VVYHLV T
Sbjct: 63 PFSHHFSSSILSMLLINRDTHRRWVL*SK*IVKEHVVLGVVYHLVQT 203
>AV409534
Length = 427
Score = 30.0 bits (66), Expect = 1.3
Identities = 18/45 (40%), Positives = 24/45 (53%)
Frame = -1
Query: 324 VKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKN 368
VKYL+ P+R G K +I + S V CI + + I PIKN
Sbjct: 241 VKYLWSPNRATHGLTKISIFEIC*TSTQ*QVICISRGTIIAPIKN 107
>BI418201
Length = 637
Score = 27.7 bits (60), Expect = 6.2
Identities = 9/28 (32%), Positives = 17/28 (60%)
Frame = -1
Query: 238 KSDYIYVWLGIIVAVHLFKMLVTIGICW 265
++ + W+G++VA L LV + +CW
Sbjct: 247 RTHVVRAWIGLLVAFILLLRLVHLSLCW 164
>TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete
Length = 1550
Score = 27.3 bits (59), Expect = 8.2
Identities = 15/57 (26%), Positives = 28/57 (48%)
Frame = -1
Query: 190 ICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSDYIYVWL 246
+C++G I L + K +E CS+F+FP F + ++I+ + Y+ L
Sbjct: 1382 VCLIGNI------LNSHTKK*IEN*CSFFIFPCFSSLSILRIECQCFPVDHLFYLLL 1230
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,292,017
Number of Sequences: 28460
Number of extensions: 183255
Number of successful extensions: 1241
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1234
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1241
length of query: 670
length of database: 4,897,600
effective HSP length: 96
effective length of query: 574
effective length of database: 2,165,440
effective search space: 1242962560
effective search space used: 1242962560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146343.3


