

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.12 + phase: 0 /pseudo
(176 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
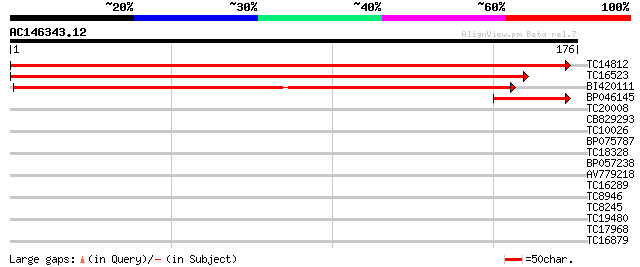
TC14812 homologue to UP|AAS47511 (AAS47511) Ribosomal protein S6... 339 2e-94
TC16523 homologue to UP|AAS47511 (AAS47511) Ribosomal protein S6... 324 6e-90
BI420111 182 4e-47
BP046145 44 2e-05
TC20008 30 0.30
CB829293 27 2.6
TC10026 26 3.4
BP075787 26 4.4
TC18328 weakly similar to UP|HSP2_PANPA (P35299) Sperm histone P... 25 5.8
BP057238 25 5.8
AV779218 25 7.5
TC16289 weakly similar to GB|AAP13385.1|30023704|BT006277 At3g56... 25 7.5
TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine b... 25 7.5
TC8245 25 7.5
TC19480 25 7.5
TC17968 similar to PIR|F86340|F86340 protein F2D10.34 [imported]... 25 9.8
TC16879 similar to UP|Q9LRH6 (Q9LRH6) ZIM, partial (17%) 25 9.8
>TC14812 homologue to UP|AAS47511 (AAS47511) Ribosomal protein S6, complete
Length = 1138
Score = 339 bits (869), Expect = 2e-94
Identities = 163/174 (93%), Positives = 169/174 (96%)
Frame = +3
Query: 1 FNVANPTTGCQKKLEIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKITGGCDKQGFP 60
FN+ANPTTGCQKKLEIDDD KLRAFWDKRISQEV GDALGEEFKGYVFKITGGCDKQGFP
Sbjct: 72 FNIANPTTGCQKKLEIDDDLKLRAFWDKRISQEVLGDALGEEFKGYVFKITGGCDKQGFP 251
Query: 61 MKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGEND 120
MKQGVLTPGRVRLLLHRGTPCFRG+GRRNGERRRKSVRGCIVSPDLSVLNLVIVKKG+ND
Sbjct: 252 MKQGVLTPGRVRLLLHRGTPCFRGYGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGDND 431
Query: 121 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRRTFTTKDGAKLPR 174
LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRR FTTK+G K+ +
Sbjct: 432 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYRRNFTTKNGKKVSK 593
>TC16523 homologue to UP|AAS47511 (AAS47511) Ribosomal protein S6, partial
(66%)
Length = 540
Score = 324 bits (830), Expect = 6e-90
Identities = 156/161 (96%), Positives = 159/161 (97%)
Frame = +1
Query: 1 FNVANPTTGCQKKLEIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKITGGCDKQGFP 60
FN+ANPTTGCQKKLEIDDD KLRAFWDKRISQEV GDALGEEFKGYVFKITGGCDKQGFP
Sbjct: 58 FNIANPTTGCQKKLEIDDDLKLRAFWDKRISQEVLGDALGEEFKGYVFKITGGCDKQGFP 237
Query: 61 MKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGEND 120
MKQGVLTPGRVRLLLHRGTPCFRG+GRRNGERRRKSVRGCIVSPDLSVLNLVIVKKG+ND
Sbjct: 238 MKQGVLTPGRVRLLLHRGTPCFRGYGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGDND 417
Query: 121 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYR 161
LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYR
Sbjct: 418 LPGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYVNTYR 540
>BI420111
Length = 528
Score = 182 bits (461), Expect = 4e-47
Identities = 91/156 (58%), Positives = 117/156 (74%)
Frame = +2
Query: 2 NVANPTTGCQKKLEIDDDQKLRAFWDKRISQEVNGDALGEEFKGYVFKITGGCDKQGFPM 61
N+A P TGCQK + I+++ + R F+D R+ EV D++G EF GYV KI GG DKQGFPM
Sbjct: 14 NIAYPATGCQKSINIEEEIRTRCFYDHRMGAEVAIDSIGPEFAGYVVKIKGGSDKQGFPM 193
Query: 62 KQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGENDL 121
QGVL RVRLLL++ + C+ R +GERRRKSVRGCI+ PD+SVL+LVIVK+GE ++
Sbjct: 194 MQGVLEKKRVRLLLNKNSGCYIPK-RFDGERRRKSVRGCIIGPDISVLDLVIVKRGEAEI 370
Query: 122 PGLTDVEKPRMRGPKRASKIRKLFNLSKDDDVRKYV 157
G+T+ K + GPKRA+KIRK+FNL K DD RKYV
Sbjct: 371 DGVTNNFKAKRLGPKRANKIRKMFNLDKKDDPRKYV 478
>BP046145
Length = 471
Score = 43.5 bits (101), Expect = 2e-05
Identities = 19/24 (79%), Positives = 21/24 (87%)
Frame = -3
Query: 151 DDVRKYVNTYRRTFTTKDGAKLPR 174
DDVRKYVNTYRRTFTTK G K+ +
Sbjct: 469 DDVRKYVNTYRRTFTTKAGKKVSK 398
>TC20008
Length = 520
Score = 29.6 bits (65), Expect = 0.30
Identities = 15/41 (36%), Positives = 19/41 (45%)
Frame = +1
Query: 82 FRGHGRRNGERRRKSVRGCIVSPDLSVLNLVIVKKGENDLP 122
F GRR+ R + GC V P L ++KK E D P
Sbjct: 130 FLVEGRRSSGGRGRGCGGCTVGPKL*PTQRYLIKKNETDKP 252
>CB829293
Length = 547
Score = 26.6 bits (57), Expect = 2.6
Identities = 16/42 (38%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Frame = +3
Query: 62 KQGVLTPGRVRLLLHRGTPC----FRGHGRRNGERRRKSVRG 99
+QG P +R + R C RGHG R+G+RR S G
Sbjct: 396 QQGRPVPALLRGAIRRVGLCVHQEIRGHGMRSGDRRSLSRHG 521
>TC10026
Length = 599
Score = 26.2 bits (56), Expect = 3.4
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -1
Query: 80 PCFRGHGRRNGERRRKSVRG 99
PC RGH RR G+RRR+ G
Sbjct: 254 PC-RGHTRRRGKRRRRGGGG 198
>BP075787
Length = 504
Score = 25.8 bits (55), Expect = 4.4
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +3
Query: 71 VRLLLHRGTPCFRGHGRRN 89
+ +LLH P FRG+G++N
Sbjct: 378 ISILLHCPNPTFRGNGKKN 434
>TC18328 weakly similar to UP|HSP2_PANPA (P35299) Sperm histone P2 precursor
(Protamine P2), partial (25%)
Length = 517
Score = 25.4 bits (54), Expect = 5.8
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +3
Query: 62 KQGVLTPGRVRLLLHRGTPCFRGHGRRNGER 92
++ VL P R+ LLL G R H RRN ++
Sbjct: 420 RRRVLQPRRLLLLLRAGVSQRRRHPRRNRDK 512
>BP057238
Length = 434
Score = 25.4 bits (54), Expect = 5.8
Identities = 15/35 (42%), Positives = 18/35 (50%), Gaps = 2/35 (5%)
Frame = -1
Query: 46 YVFKITGGC--DKQGFPMKQGVLTPGRVRLLLHRG 78
YVF ITGGC K G+ M V PG + + G
Sbjct: 143 YVFVITGGC*SLKHGYVMLGRVPVPGTYPVRIRYG 39
>AV779218
Length = 443
Score = 25.0 bits (53), Expect = 7.5
Identities = 10/29 (34%), Positives = 18/29 (61%)
Frame = -1
Query: 130 PRMRGPKRASKIRKLFNLSKDDDVRKYVN 158
P +R ++ S+I ++L D D+R Y+N
Sbjct: 308 PWLRRVRKPSQIIFYYSLKNDKDIRSYLN 222
>TC16289 weakly similar to GB|AAP13385.1|30023704|BT006277 At3g56360
{Arabidopsis thaliana;}, partial (25%)
Length = 527
Score = 25.0 bits (53), Expect = 7.5
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = +2
Query: 71 VRLLLHRGTPCFRGHGRRNGERRRKSVRGC 100
VR RG +R RR RRR+ RGC
Sbjct: 23 VRRRCRRGLYSWRSWMRRRRRRRRRGRRGC 112
>TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine
biosynthesis protein-like {Arabidopsis thaliana;} ,
partial (45%)
Length = 503
Score = 25.0 bits (53), Expect = 7.5
Identities = 11/23 (47%), Positives = 13/23 (55%)
Frame = -2
Query: 78 GTPCFRGHGRRNGERRRKSVRGC 100
G C RGH R RRR+ +R C
Sbjct: 295 GRGCRRGHARGP*RRRRRLLRRC 227
>TC8245
Length = 623
Score = 25.0 bits (53), Expect = 7.5
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -3
Query: 62 KQGVLTPGRVRLLLHRGTPCFRG 84
++ ++TPG+ L HRGTP +G
Sbjct: 297 RKEIITPGQPVGLKHRGTPIEKG 229
>TC19480
Length = 511
Score = 25.0 bits (53), Expect = 7.5
Identities = 18/50 (36%), Positives = 22/50 (44%)
Frame = +1
Query: 31 SQEVNGDALGEEFKGYVFKITGGCDKQGFPMKQGVLTPGRVRLLLHRGTP 80
SQ+ G + KG TG C K F MK +LT +L GTP
Sbjct: 34 SQQRTGSLSLKVKKGKSEVETGSCAKNAFIMK*HMLTQKSYKLNKRIGTP 183
>TC17968 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 607
Score = 24.6 bits (52), Expect = 9.8
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = +1
Query: 58 GFPMKQGVLTPGRVRLLLHRGTPCFRGHGRRNGERRR 94
G+ KQGV RL H G+ R H RR R R
Sbjct: 331 GYRRKQGV------RLRCHHGSTPLRPHMRRRPHRHR 423
>TC16879 similar to UP|Q9LRH6 (Q9LRH6) ZIM, partial (17%)
Length = 622
Score = 24.6 bits (52), Expect = 9.8
Identities = 19/39 (48%), Positives = 21/39 (53%), Gaps = 4/39 (10%)
Frame = -3
Query: 74 LLHRGTPCFRGHGRRNG--ERRR--KSVRGCIVSPDLSV 108
L HR RG+G RNG RRR +S RGC S L V
Sbjct: 305 LRHRHRHRHRGYGNRNGCDFRRRCLRSDRGCSWSCTLLV 189
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,749,527
Number of Sequences: 28460
Number of extensions: 33698
Number of successful extensions: 197
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 194
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 196
length of query: 176
length of database: 4,897,600
effective HSP length: 84
effective length of query: 92
effective length of database: 2,506,960
effective search space: 230640320
effective search space used: 230640320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146343.12


