

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.8 - phase: 0
(337 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
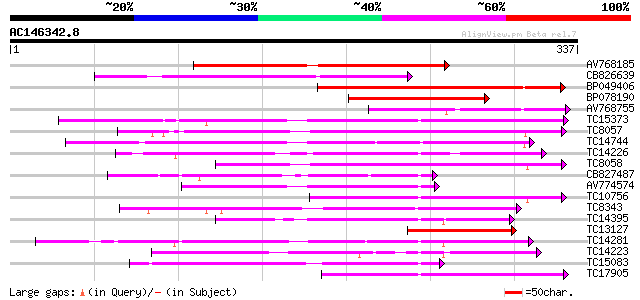
Score E
Sequences producing significant alignments: (bits) Value
AV768185 136 6e-33
CB826639 126 6e-30
BP049406 125 8e-30
BP078190 102 9e-23
AV768755 95 1e-20
TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 hom... 91 3e-19
TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, par... 89 8e-19
TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment)... 88 2e-18
TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete 83 6e-17
TC8058 weakly similar to UP|Q94FM7 (Q94FM7) Elicitor-inducible c... 83 7e-17
CB827487 80 6e-16
AV774574 77 3e-15
TC10756 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 mon... 76 7e-15
TC8343 weakly similar to GB|BAB02401.1|9294391|AB023038 cytochro... 76 7e-15
TC14395 weakly similar to UP|C823_SOYBN (O49858) Cytochrome P450... 76 7e-15
TC13127 weakly similar to UP|O49373 (O49373) Cytochrome P450 - l... 72 1e-13
TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygen... 72 2e-13
TC14223 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethylas... 70 5e-13
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 69 9e-13
TC17905 weakly similar to UP|Q9STH8 (Q9STH8) Flavonoid 3', 5'-hy... 68 2e-12
>AV768185
Length = 446
Score = 136 bits (342), Expect = 6e-33
Identities = 70/152 (46%), Positives = 94/152 (61%)
Frame = +3
Query: 110 SSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESN 169
S N DKYLRDI+++F++AG+DT A+ L+ FF++L +P ++ + +EV V Q S
Sbjct: 9 SVNDDDKYLRDIVVSFLLAGRDTVASGLTCFFWLLSNHPKVEALIREEVDRVMIPKQNSV 188
Query: 170 LNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETV 229
F L +M YL+AA+ E++RLYP V + A E D+LPDG V KG V
Sbjct: 189 QGFASFEQ------LREMQYLNAAIYESMRLYPPVQFDSKFAHEDDVLPDGTFVRKGTRV 350
Query: 230 YYLSYAMGRMPYIWGDDAQEFLPERWLKDGIF 261
Y YAMGRM +WG D EF PERWLKDG+F
Sbjct: 351 TYHPYAMGRMERVWGLDCLEFRPERWLKDGVF 446
>CB826639
Length = 542
Score = 126 bits (316), Expect = 6e-30
Identities = 71/190 (37%), Positives = 112/190 (58%), Gaps = 1/190 (0%)
Frame = +3
Query: 51 KLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDS 110
+ ++ FG+E LK+++KI++++++ + R + S +D+LSRF+ +
Sbjct: 3 RFQKFFCFGAEKKLKESLKIVENYMNEAVSAR--------EASPSDDLLSRFMRKRDTAG 158
Query: 111 SNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINV-TSTSQESN 169
LR I LNF++AG+DTS+ LSWFF+++ +PL++EK+ E+ V T T E
Sbjct: 159 KPFDAAKLRHIALNFVLAGRDTSSVALSWFFWLVMNHPLVEEKIVLELAAVLTDTRGEET 338
Query: 170 LNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETV 229
E + +A D++ YL AAL ETLRLYP+VP + A D+LPDG +V G TV
Sbjct: 339 GKWVEEAVDFEEA--DRLVYLKAALAETLRLYPSVPEDFKYALNDDVLPDGTVVPAGSTV 512
Query: 230 YYLSYAMGRM 239
Y Y++GRM
Sbjct: 513 TYSIYSVGRM 542
>BP049406
Length = 566
Score = 125 bits (315), Expect = 8e-30
Identities = 68/149 (45%), Positives = 92/149 (61%), Gaps = 2/149 (1%)
Frame = -1
Query: 184 LDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIW 243
+D++ YL AAL+ETLRLYP+VP + D+LPDG V G +V Y Y+ GRM W
Sbjct: 479 VDRLVYLKAALSETLRLYPSVPEDSKHVVADDVLPDGTFVPAGSSVTYSIYSAGRMRSTW 300
Query: 244 GDDAQEFLPERWLK-DGI-FQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFR 301
G+D EF PERWL DG F FKF++F+AGPRICLGKD AY QMK ++ A++ R
Sbjct: 299 GEDCMEFRPERWLSLDGTKFVQHDPFKFVAFNAGPRICLGKDLAYLQMKSIASAVLLNHR 120
Query: 302 FKLENETNDVTYRTMFTLHIDHGLPLYAT 330
++ + V + TL + +GL + T
Sbjct: 119 LEVV-PGHQVEQKMSLTLFMKNGLKVMCT 36
>BP078190
Length = 445
Score = 102 bits (254), Expect = 9e-23
Identities = 48/84 (57%), Positives = 54/84 (64%)
Frame = -3
Query: 202 PAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIF 261
P V + A E D+LPDG V KG V Y YAMGRM IWG D EF PERWLKDG+F
Sbjct: 443 PPVQFDSKYATEDDVLPDGTFVRKGSRVTYHPYAMGRMERIWGPDCLEFKPERWLKDGVF 264
Query: 262 QPESSFKFISFHAGPRICLGKDFA 285
P+ FK+ F AG R+CLGKD A
Sbjct: 263 VPKCPFKYPVFQAGVRVCLGKDLA 192
>AV768755
Length = 488
Score = 95.1 bits (235), Expect = 1e-20
Identities = 50/123 (40%), Positives = 72/123 (57%), Gaps = 3/123 (2%)
Frame = -1
Query: 214 HDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKD---GIFQPESSFKFI 270
H LP G+ V + YL Y+MGRM IWGDD+ EF PERW+ + I P S KFI
Sbjct: 473 HATLPSGHHVGPNTKILYLLYSMGRMEQIWGDDSLEFKPERWISEKGHSIHMPSS--KFI 300
Query: 271 SFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYAT 330
+F+AGPR CLGK + +K+V+ A++ FR ++ E + +T R + + +G + T
Sbjct: 299 TFNAGPRSCLGKGIPFIMLKMVAAAIIPKFRIQV-MEDHPITPRLSVVMGMRYGFKVQVT 123
Query: 331 PRC 333
RC
Sbjct: 122 KRC 114
>TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 homolog,
partial (61%)
Length = 1059
Score = 90.9 bits (224), Expect = 3e-19
Identities = 81/308 (26%), Positives = 139/308 (44%), Gaps = 5/308 (1%)
Frame = +1
Query: 30 FMKAFNDSNAFVFKRYLDPLWK-LKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSM 88
F +A D A + K L + L R G E + K + D +I R ++ S
Sbjct: 40 FREAVADMTALLGKPNLSDFFPGLARFDLQGVEKQMHKVVPRFDRIFEKMIGERVKMESE 219
Query: 89 QKDFSDKEDILSRFLLESKKDSSNMTD---KYLRDIILNFMIAGKDTSANTLSWFFYMLC 145
K S+ +D L +FLL K++ + T +++ ++++ + G DTS+NT+ + +
Sbjct: 220 GKR-SESKDFL-QFLLNLKEEGDSKTPLSITHVKALLMDMLAGGSDTSSNTVEFAMAEMM 393
Query: 146 KNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVP 205
+ P + ++V +E+ V + + ++ + K+ YL A + ETLRL+P +P
Sbjct: 394 QKPEVMKRVQEELEGVVGRD-----------NMVEESHIHKLPYLLAVMKETLRLHPVLP 540
Query: 206 MSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPES 265
+ GY + KG V+ +A+ R P IW ++ EF P R+L +
Sbjct: 541 LLVPHCPSETTSAGGYTIPKGSRVFVNVWAIHRDPSIW-ENPLEFDPTRFLDAKWDFSGN 717
Query: 266 SFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKL-ENETNDVTYRTMFTLHIDHG 324
F + F +G RIC G A R + LV F + + E E DV+ + L +
Sbjct: 718 DFNYFPFGSGRRICAGIAMAERSVLYFLATLVHLFDWTVPEGEKLDVSEKFGIVLKKETP 897
Query: 325 LPLYATPR 332
L TPR
Sbjct: 898 LVAIPTPR 921
>TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (29%)
Length = 1148
Score = 89.4 bits (220), Expect = 8e-19
Identities = 74/276 (26%), Positives = 118/276 (41%), Gaps = 9/276 (3%)
Frame = +2
Query: 65 KKNIKIIDDFVHSLIKTRR--ELLSMQK-----DFSDKEDILSRFLLESKKDSSNMTDKY 117
KK + D + +IK LS QK DF+D +LS ++ KD +
Sbjct: 68 KKAHEEFDQMLEQIIKDHEAPSRLSDQKNGQSIDFTDT--LLSH--MKQSKDKHIINKTN 235
Query: 118 LRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVS 177
L+ I+L+ +I DT+ T W L ++P K+ E+ NV +
Sbjct: 236 LKAILLDIIIGSLDTTIMTTDWAMSELLRHPKAMTKLQDELKNVVGMGRV---------- 385
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
+ + + K+ YL+ + ET R++P P+ DI +GY + K V S+ +G
Sbjct: 386 -VEEDDIPKLPYLNMVVKETFRIHPPAPLLVPRECLEDITINGYFIAKKSRVLVNSWTIG 562
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
R P IW D+A++F P+R+ F+F+ F +G R C G + +V LV
Sbjct: 563 RDPKIWSDNAEDFYPDRFQNTDTDSHGLHFQFLPFGSGRRRCPGMQLGLTTVPLVLAQLV 742
Query: 298 RFFRFKLE--NETNDVTYRTMFTLHIDHGLPLYATP 331
F ++L ND+ F L L A P
Sbjct: 743 HCFNWELPLGMSANDLDMTEEFGLTTPRVQNLLAIP 850
>TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment) , complete
Length = 1753
Score = 87.8 bits (216), Expect = 2e-18
Identities = 67/283 (23%), Positives = 134/283 (46%), Gaps = 4/283 (1%)
Frame = +1
Query: 34 FNDSNAFVFKRYLDPLWKLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFS 93
FN ++ F ++L+ L +++ + + + IIDD + +R + ++
Sbjct: 664 FNIADLFPSAKWLENLTRMRSKFEYLHQKMDRILETIIDDHKAN---SRTKEGQVEGGEE 834
Query: 94 DKEDILSRFLLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEK 153
D D+L ++ S ++T + ++ I+ + IAG +TSA T++W + K+P++ +K
Sbjct: 835 DLIDVLLKYENSSTDQDFHLTIRNIKAILFDIFIAGSETSATTINWTMAEMMKDPILLKK 1014
Query: 154 VAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEE 213
EV + + + + ++ YL A + E LRL+P P+ R +
Sbjct: 1015AQDEVREIFQRR-----------GKVDETCIYELKYLKAFINEVLRLHPPGPLVFRECRQ 1161
Query: 214 HDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFH 273
+ +GY + TV ++A+G W + + F PER++ I ++F+ + F
Sbjct: 1162ACEI-NGYHIPAKSTVLVNTFAIGTDSKYWA-EPERFCPERFIDSSIDYQGTNFEHLPFG 1335
Query: 274 AGPRICLGKDFAYRQMKIVSMALVRFFRFKL----ENETNDVT 312
AG RIC G ++ +++V L+ F + L +NE D+T
Sbjct: 1336AGRRICPGINYGMANVELVLALLLYHFDWTLPKGIKNEDLDLT 1464
>TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete
Length = 1878
Score = 83.2 bits (204), Expect = 6e-17
Identities = 71/262 (27%), Positives = 119/262 (45%), Gaps = 6/262 (2%)
Frame = +3
Query: 64 LKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKED-----ILSRFLLESKKDSSNMTDKYL 118
L+K +K I KT R L + ++ DK+ ++ L + TD+ +
Sbjct: 759 LEKRVKGISS------KTDRFLRGLLQEHRDKKQRTANTMIDHLLTLQESQPEYYTDQII 920
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSN 178
+ + L ++AG D+SA TL W + P + EK+ E+ ++++ D V
Sbjct: 921 KGLALAMLLAGTDSSAVTLEWSMCNVLNYPEVLEKIKAEL--------DTHVGQDRLVD- 1073
Query: 179 ISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGR 238
++ + K+ YL + ETLRLY P+ + D GY V + V ++A+ R
Sbjct: 1074--ESDIPKLTYLKNVINETLRLYTPAPLLLPHSASDDCTIGGYKVPRDTIVLINAWALHR 1247
Query: 239 MPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVR 298
P +W +A F PER+ K G + K I F G R C G+ A R + + L++
Sbjct: 1248DPQLW-TEATTFKPERFDKKGELE-----KLIPFGLGRRACPGELLAIRAISMTLALLIQ 1409
Query: 299 FFRFK-LENETNDVTYRTMFTL 319
F +K + +E D+ R F L
Sbjct: 1410CFDWKRVSDEEIDMGERDGFVL 1475
>TC8058 weakly similar to UP|Q94FM7 (Q94FM7) Elicitor-inducible cytochrome
P450, partial (28%)
Length = 887
Score = 82.8 bits (203), Expect = 7e-17
Identities = 56/211 (26%), Positives = 94/211 (44%), Gaps = 2/211 (0%)
Frame = +2
Query: 123 LNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDA 182
L+ +I DT+ ++ W L ++P + +K+ E+ NV + + +A
Sbjct: 2 LDIIIGAVDTTIMSVDWNMAELIRHPRVMKKLQDELKNVVGMGRM-----------VEEA 148
Query: 183 TLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYI 242
L K+ YL + ET RL+P P+ DI +GY + K V S+ +GR P +
Sbjct: 149 DLPKLPYLDMVMKETFRLHPPAPLLPPRECTEDITINGYFIAKKSRVLVNSWTLGRDPKM 328
Query: 243 WGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRF 302
W D+ ++F PER+ I +++F+ F +G R C G + + LV F +
Sbjct: 329 WSDNFEDFYPERFHNTDIDSHGHNYQFLPFGSGRRRCPGMLLGLTTVPFLLAQLVHCFNW 508
Query: 303 KLEN--ETNDVTYRTMFTLHIDHGLPLYATP 331
+L T+D+ F L L A P
Sbjct: 509 ELPPGVSTDDMDMTEEFGLTTPRTQNLLAIP 601
>CB827487
Length = 569
Score = 79.7 bits (195), Expect = 6e-16
Identities = 60/199 (30%), Positives = 97/199 (48%), Gaps = 3/199 (1%)
Frame = +3
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSS---NMTD 115
GS A L K I D F ++ + + KD + ++DI+ LL+ + S ++TD
Sbjct: 18 GSLARLDKTINSFDAFFQQVLDEHLDPNRI-KDQTQEDDIVDT-LLQLRNQGSLSIDLTD 191
Query: 116 KYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEF 175
++++ +++ +I DTS W L KNP +KV +E+ N+ +N D
Sbjct: 192 EHIKAFMMDLLIGSTDTSVAASVWLMTGLMKNPTAMKKVQEEIRNLC-------VNKD-- 344
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
I + + K+ YL A + ETLR YP P+ R + +I+ DGY + VY +A
Sbjct: 345 --FIDEVDIQKLEYLKAVIKETLRFYPLAPLIPRETIK-NIIVDGYEIPAKTIVYVNVWA 515
Query: 236 MGRMPYIWGDDAQEFLPER 254
+ R P W D EF P+R
Sbjct: 516 IHRDPEAW-KDPHEFNPDR 569
>AV774574
Length = 510
Score = 77.4 bits (189), Expect = 3e-15
Identities = 47/153 (30%), Positives = 75/153 (48%)
Frame = +1
Query: 103 LLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVT 162
L E +T L+ ++L+ +AG DTS+ TL W L KNP +KV +EV V
Sbjct: 64 LQEGGMSEFELTQDDLKALLLDMCLAGTDTSSTTLEWAMAELVKNPATMKKVQEEVRRVV 243
Query: 163 STSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYI 222
+ S I D+ +++M Y+ + ETLRL+PA P+ + GY
Sbjct: 244 GSK-----------SRIEDSDVNQMEYMKCVVKETLRLHPAAPLLVPRETISSVKLGGYD 390
Query: 223 VNKGETVYYLSYAMGRMPYIWGDDAQEFLPERW 255
+ VY ++A+ R P +W + + F+PER+
Sbjct: 391 IPSKRMVYINAWAIQRDPELW-ERPEVFIPERF 486
>TC10756 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 monooxygenase
(Fragment), partial (37%)
Length = 887
Score = 76.3 bits (186), Expect = 7e-15
Identities = 50/155 (32%), Positives = 78/155 (50%), Gaps = 2/155 (1%)
Frame = +3
Query: 179 ISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGR 238
+ ++ ++++ YL A + E RL+PAVP+ E D+ GY + KG V +A GR
Sbjct: 24 VEESHIERLPYLQAIVKEIFRLHPAVPLLLPRQAEVDLDMHGYTIPKGSQVLVNVWANGR 203
Query: 239 MPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVR 298
P +W D+ F PER+L I SF+ F AG RIC G A R + ++ L+
Sbjct: 204 DPNLW-DNPNLFSPERFLGSEIDFKGRSFELTPFGAGRRICPGLPLAIRLLFLMLGLLIN 380
Query: 299 FFRFKLEN--ETNDVTYRTMFTLHIDHGLPLYATP 331
F ++LE + D+ F L ++ P+ A P
Sbjct: 381 CFNWELEGGIKPEDMDMDETFGLTLEKTQPVLAVP 485
>TC8343 weakly similar to GB|BAB02401.1|9294391|AB023038 cytochrome P450
{Arabidopsis thaliana;} , partial (60%)
Length = 1784
Score = 76.3 bits (186), Expect = 7e-15
Identities = 64/247 (25%), Positives = 107/247 (42%), Gaps = 8/247 (3%)
Frame = +1
Query: 66 KNIKIIDDFVHSLIKT--RRELLSMQKDFSDKEDILSRFLLESKKDSSNMTD---KYLRD 120
+ +K ID + + R L +++ D+L L + K+S + LR+
Sbjct: 811 RRMKAIDKEIRGSLMAIINRRLKAIKAGEPTNNDLLGILLESNYKESEKSSGGGGMSLRE 990
Query: 121 IILN---FMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVS 177
++ F +AG++ +A L W +L K+P Q K +EV V + + L +
Sbjct: 991 VVEEVKLFYLAGQEANAELLVWTLLLLSKHPDWQAKAREEVFKVFGNEKPDHDKLGQ--- 1161
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
+ + L E+LRLYP V M R + L D I E + +S
Sbjct: 1162 ---------LKIVSMILQESLRLYPPVVMFARYLRKDTKLGDLTIPAGVEIIVPVSMLHQ 1314
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
+ WGDDA EF PER+ + + +I F GPR+C+G++F + KI ++
Sbjct: 1315 EKEF-WGDDAMEFNPERFSEGVSKATKGKVCYIPFGWGPRLCIGQNFGLLEAKIALSMIL 1491
Query: 298 RFFRFKL 304
+ F L
Sbjct: 1492 QHFSLDL 1512
>TC14395 weakly similar to UP|C823_SOYBN (O49858) Cytochrome P450 82A3
(P450 CP6) , partial (38%)
Length = 1079
Score = 76.3 bits (186), Expect = 7e-15
Identities = 57/181 (31%), Positives = 86/181 (47%), Gaps = 3/181 (1%)
Frame = +3
Query: 123 LNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDA 182
L ++AG DT+ +TL+W +L N +K E+ T + I ++
Sbjct: 12 LQLIVAGTDTTTSTLTWALSLLLNNREALKKATHEL----DTQMGGR-------TKIMES 158
Query: 183 TLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYI 242
+K+ YL A + ETLRLYP P++ D + GY V G ++ + R P I
Sbjct: 159 DFEKLVYLQAIIKETLRLYPVAPLNVTHMSMEDCVVGGYHVPAGTSLVTNISKIQRDPSI 338
Query: 243 WGDDAQEFLPERWL---KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRF 299
+ D EF PER+L KD + +F+ I F AG RIC G FA + +K+ L+
Sbjct: 339 Y-SDPMEFRPERYLTTHKD-LDMKGKNFELIPFGAGRRICPGISFALQLIKMTLATLLHG 512
Query: 300 F 300
F
Sbjct: 513 F 515
>TC13127 weakly similar to UP|O49373 (O49373) Cytochrome P450 - like protein
(Cytochrome P450-like protein), partial (13%)
Length = 495
Score = 72.0 bits (175), Expect = 1e-13
Identities = 33/66 (50%), Positives = 46/66 (69%), Gaps = 1/66 (1%)
Frame = +3
Query: 237 GRMPYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
GR +WG D EF PERW+ + G S+KFISF+AGPR CLGKD ++ Q+KI++ A
Sbjct: 3 GRSEDVWGKDCLEFKPERWVSERGSIVHAPSYKFISFNAGPRTCLGKDLSFIQLKIIATA 182
Query: 296 LVRFFR 301
++R +R
Sbjct: 183 VLRNYR 200
>TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1,
partial (97%)
Length = 1768
Score = 71.6 bits (174), Expect = 2e-13
Identities = 68/304 (22%), Positives = 135/304 (44%), Gaps = 8/304 (2%)
Frame = +3
Query: 16 ELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGSEAALKKNIKIIDDFV 75
E+ L K +E + +FN + F R LD + +G + K + +D +
Sbjct: 651 EIGELRKVIREIGELLGSFNLGDIIGFMRPLD-------LQGYGKKN--KDMHRKLDGMM 803
Query: 76 HSLIKTRRELLSMQKDFSDKE----DILSRFLLESKKDSSNMTDKYLRDIILNFMIAGKD 131
++K E + + SD++ DIL L+E+ + +T + L+ +AG +
Sbjct: 804 EKVLKEHEEARATEGADSDRKKDLFDILLN-LIEADGADNKLTRDSAKAFALDMFLAGTN 980
Query: 132 TSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLH 191
A+ L W L +NP + +K +E+ V + + ++ + + YL
Sbjct: 981 GPASVLEWSLAELIRNPHVLKKAREEIDTVVGKER-----------LVKESDIPNLPYLQ 1127
Query: 192 AALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFL 251
A + ETLR++P P+ R A DGY + ++ +YA+GR W ++ +
Sbjct: 1128AVVKETLRMHPPTPIFAREA-MRSCQVDGYDIPANSKIFISAYAIGRDSQYW-ENPHVYD 1301
Query: 252 PERWL---KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKL-ENE 307
PER+L + + ++ + F +G R C G A ++ +LV+ F + + + +
Sbjct: 1302PERFLFTDERKVDVRGQYYQLLPFGSGRRSCPGASLALIVIQASLASLVQCFDWVVNDGK 1481
Query: 308 TNDV 311
+N++
Sbjct: 1482SNEI 1493
>TC14223 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethylase, complete
Length = 1917
Score = 70.1 bits (170), Expect = 5e-13
Identities = 60/248 (24%), Positives = 113/248 (45%), Gaps = 16/248 (6%)
Frame = +3
Query: 85 LLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWF-FYM 143
++S + ++D+L F+ KD T+ + +++ + AG+ TS+ T +W Y+
Sbjct: 822 IVSRKSANKSEDDMLQCFIDSKYKDGRPTTEGEVTGLLIAALFAGQHTSSITSTWTGAYL 1001
Query: 144 LCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPA 203
+C N + V +E + +++ + L +M L+ + E LRL+P
Sbjct: 1002 MCNNKYLSAVV-----------EEQKVLMEKHGDRVDHDVLAEMDVLYRCIKEALRLHPP 1148
Query: 204 VPM-----------SGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLP 252
+ M + R +E+DI P G+IV R+P+I+ D + P
Sbjct: 1149 LIMLLRSSHSDFSVTTREGKEYDI-PKGHIVATSPAF------ANRLPHIF-KDPDTYDP 1304
Query: 253 ERWL----KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKLENET 308
+R+ +D + +F +ISF G CLG+ FAY Q+K + L+R F +L +
Sbjct: 1305 DRFAVGREED---KAAGAFSYISFGGGRHGCLGEPFAYLQIKAIWTHLLRNFELELVSPF 1475
Query: 309 NDVTYRTM 316
++ + M
Sbjct: 1476 PEIDWNAM 1499
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 69.3 bits (168), Expect = 9e-13
Identities = 50/188 (26%), Positives = 87/188 (45%), Gaps = 1/188 (0%)
Frame = +1
Query: 72 DDFVHSLIKTRRELLSMQKDFSDKEDILSRFL-LESKKDSSNMTDKYLRDIILNFMIAGK 130
+D + LI+ R+ L K +D + L LE ++ + + + + + AG
Sbjct: 775 EDVLVPLIRARK-LAKEGKPVNDVVSYVDTLLELELPEEKRKLNESEMVTLCSEILNAGT 951
Query: 131 DTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYL 190
DT++ L W L K P +Q+ + E+ +V ++ + ++ V K+ YL
Sbjct: 952 DTTSTALEWVLANLDKYPRVQKNIVDEISDVMKGREDKEVKEEDLV---------KLPYL 1104
Query: 191 HAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEF 250
A + E LR +P A D + DGY+V K TV ++ MG P +W ++ EF
Sbjct: 1105KAVILEGLRRHPPGHFVLPHAVTEDTVLDGYLVPKDGTVNFMVAEMGWDPQVW-EEPMEF 1281
Query: 251 LPERWLKD 258
PER+L +
Sbjct: 1282KPERFLNN 1305
>TC17905 weakly similar to UP|Q9STH8 (Q9STH8) Flavonoid 3', 5'-hydroxylase
like protein (Flavonoid 3, 5-hydroxylase like protein),
partial (30%)
Length = 597
Score = 67.8 bits (164), Expect = 2e-12
Identities = 49/148 (33%), Positives = 70/148 (47%), Gaps = 1/148 (0%)
Frame = +2
Query: 186 KMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGD 245
K+ YL A + ETLRL+P VP+ GY + KG V+ +A+ R P IW +
Sbjct: 5 KLPYLLAVMKETLRLHPTVPLLVPHCPSEATSTGGYTIPKGSRVFVNVWAIHRDPSIW-E 181
Query: 246 DAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKL- 304
EF PER+L + F + F +G RIC+G A R + LV F + +
Sbjct: 182 KPLEFDPERFLDAKWDFCGNDFSYFPFGSGRRICVGIPMAERSVLYFLATLVHMFDWTVP 361
Query: 305 ENETNDVTYRTMFTLHIDHGLPLYATPR 332
E E D++ + TL + L TPR
Sbjct: 362 EGEKLDISEKFGITLKKETPLVAIPTPR 445
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,339,835
Number of Sequences: 28460
Number of extensions: 63876
Number of successful extensions: 376
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 349
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 349
length of query: 337
length of database: 4,897,600
effective HSP length: 91
effective length of query: 246
effective length of database: 2,307,740
effective search space: 567704040
effective search space used: 567704040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146342.8


