

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.8 + phase: 0
(268 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
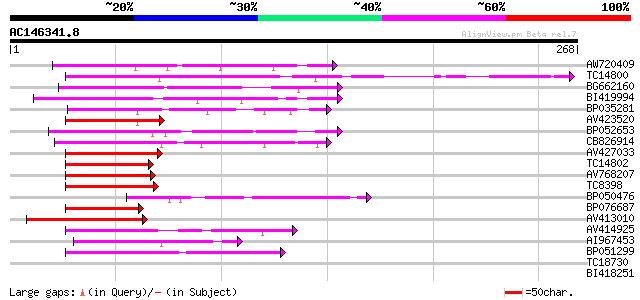
Score E
Sequences producing significant alignments: (bits) Value
AW720409 75 9e-15
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 72 1e-13
BG662160 66 7e-12
BI419994 52 8e-08
BP035281 50 3e-07
AV423520 49 9e-07
BP052653 48 2e-06
CB826914 47 3e-06
AV427033 47 4e-06
TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (... 47 4e-06
AV768207 46 6e-06
TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific... 46 6e-06
BP050476 44 2e-05
BP076687 44 3e-05
AV413010 43 5e-05
AV414925 41 2e-04
AI967453 41 2e-04
BP051299 41 2e-04
TC18730 similar to UP|Q963J2 (Q963J2) Voltage-dependent calcium ... 36 0.006
BI418251 31 0.25
>AW720409
Length = 575
Score = 75.5 bits (184), Expect = 9e-15
Identities = 56/148 (37%), Positives = 75/148 (49%), Gaps = 13/148 (8%)
Frame = +3
Query: 21 LTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLI-SDPTFVQKQLKRPPR------ 73
+ + LP EL +ILS L VKT++R C+SKSW +LI D F + L R P+
Sbjct: 75 MEIEFVLPFELWIEILSWLPVKTLMRFSCVSKSWKSLIYQDRDFKKLHLDRSPKNNNQVI 254
Query: 74 LILKPPCWVYPMRTIQSLPITRLLK-----NPSITVSGDFCYDGGGLNDNCKVVI-SCNG 127
L L+ P ++ RT P+ RLL+ + SI F + DN I SCNG
Sbjct: 255 LTLQKP--LFSGRTFFPFPVRRLLQDQEPSSSSIIDDIPFEEEDEDEEDNYHDTIGSCNG 428
Query: 128 LLCFIFCSDNKEYHNYWFRIWNPATGTR 155
L+C +D H WFR+WNPAT R
Sbjct: 429 LVCLYGIND----HGLWFRLWNPATRFR 500
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5, TAC
clone:K14B20, partial (7%)
Length = 782
Score = 71.6 bits (174), Expect = 1e-13
Identities = 69/252 (27%), Positives = 104/252 (40%), Gaps = 11/252 (4%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK----------RPPRLIL 76
LP+ELI +ILS L VK++L+ +SK+W + ISDP FV+ L L++
Sbjct: 48 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTSLLI 227
Query: 77 KPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSD 136
K + I S ++ LL++PS V+ C G C ++S L D
Sbjct: 228 KCHTDDFGRPYISSRTVSSLLESPSAIVASRSCISGYDFIGTCNGLVSLRKL-----NYD 392
Query: 137 NKEYHNY-WFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLG 195
+N+ R WNPAT T S+ + + R+L FG
Sbjct: 393 ESNTNNFSQVRFWNPATRTMSQ----DSPPSWSPRNLHLGFG------------------ 506
Query: 196 CLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDSFPVIPLIS 255
+ C S TYK V P + V + ++ D+CWR I P P+
Sbjct: 507 -----YDCS--SDTYKVVGMI----------PGLTMVNVYNMGDNCWRTIQISPHAPMHL 635
Query: 256 PFNNGVYLSGTI 267
+ VY+S T+
Sbjct: 636 Q-GSAVYVSNTL 668
>BG662160
Length = 445
Score = 65.9 bits (159), Expect = 7e-12
Identities = 45/135 (33%), Positives = 66/135 (48%), Gaps = 1/135 (0%)
Frame = +2
Query: 24 SMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVY 83
+ FLPEEL +ILS L VK+++R C+SKSW ++ISD F++ L R + + Y
Sbjct: 14 AQFLPEELRLEILSWLPVKSLVRFRCVSKSWKSIISDSQFIKLHLHRSSS-TTRNTDFAY 190
Query: 84 PMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCS-DNKEYHN 142
I S +R +I DFC +CNGL+C D K+ +
Sbjct: 191 LQSLITSPRKSRSASTIAIPDDYDFCG-------------TCNGLVCLHSSKYDRKDKYT 331
Query: 143 YWFRIWNPATGTRSK 157
R+WNPA + S+
Sbjct: 332 SHVRLWNPAMRSMSQ 376
>BI419994
Length = 549
Score = 52.4 bits (124), Expect = 8e-08
Identities = 48/158 (30%), Positives = 68/158 (42%), Gaps = 12/158 (7%)
Frame = +2
Query: 12 KKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRP 71
K + L+L LP+ELI QIL L V+++LR + KSW LISDP F +
Sbjct: 2 KHPQTSTESLSLFSHLPDELILQILLRLPVRSLLRFKTVCKSWRYLISDPQFTKSHF--- 172
Query: 72 PRLILKPPCWVYPMRT----IQSLPITRLLKNPSITVSGDF--------CYDGGGLNDNC 119
L P ++ T I+SL L + S V F + G
Sbjct: 173 -NLAASPTHRIFLNNTNGF*IESLDTDSSLHDESSLVHLKFPLAPPHPDNHRFGSSVTGL 349
Query: 120 KVVISCNGLLCFIFCSDNKEYHNYWFRIWNPATGTRSK 157
KV+ SC G F+ ++ H +WNP+TG R +
Sbjct: 350 KVLGSCRG---FVLLAN----HQGNVVVWNPSTGVRRR 442
>BP035281
Length = 498
Score = 50.4 bits (119), Expect = 3e-07
Identities = 44/139 (31%), Positives = 67/139 (47%), Gaps = 14/139 (10%)
Frame = +2
Query: 28 PEELIAQILSLLDVKTILRLICLSKSWNTLIS-DPTFVQKQLKRPPRLILKPPCWVYPMR 86
P EL+ +ILS L V +++R C+ KSW +LIS + F + QL+R LK +
Sbjct: 125 PSELVIEILSWLPVLSLIRFKCVCKSWKSLISHNKAFAKLQLERTS---LKINHVILTSS 295
Query: 87 TIQSLP--ITRLLKNPSITVSGDFCYDGGGLNDNC---------KVVISCNGLLCF--IF 133
+P + RLL++PS + D D C +V+ SCNGL+C +
Sbjct: 296 DSSFIPCSLPRLLEDPSSIMDQD--------QDRCLRLDKFEYNEVIGSCNGLICLHADY 451
Query: 134 CSDNKEYHNYWFRIWNPAT 152
C+ + FR NP+T
Sbjct: 452 CAP-----GFCFRPCNPST 493
>AV423520
Length = 468
Score = 48.9 bits (115), Expect = 9e-07
Identities = 24/48 (50%), Positives = 34/48 (70%), Gaps = 1/48 (2%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLIS-DPTFVQKQLKRPPR 73
LP EL+ +IL L VK++L+ C+ KSWN+LIS DP F +K L+ P+
Sbjct: 165 LPFELVVEILCRLPVKSLLQFRCVCKSWNSLISGDPKFARKHLRCSPK 308
>BP052653
Length = 552
Score = 47.8 bits (112), Expect = 2e-06
Identities = 42/143 (29%), Positives = 69/143 (47%), Gaps = 4/143 (2%)
Frame = +1
Query: 19 KKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK--QLKRPP--RL 74
+ L LS LP +LI QIL L V+++LR + KSW +LISDP F + L P R
Sbjct: 163 ESLNLSSILPHDLIVQILLRLPVRSLLRFKSVCKSWLSLISDPQFGKSHFDLAAAPTHRC 342
Query: 75 ILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFC 134
++K + ++SL + + + S V+ F Y G + + +++ SC G + +
Sbjct: 343 LVKINNF-----KLESLDVDASVLDQSSEVNLKFPYTGYVI-FHIQILGSCRGFIALVRG 504
Query: 135 SDNKEYHNYWFRIWNPATGTRSK 157
D +WNP+ + K
Sbjct: 505 GD--------AIVWNPSXCVQRK 549
>CB826914
Length = 575
Score = 47.0 bits (110), Expect = 3e-06
Identities = 43/137 (31%), Positives = 63/137 (45%), Gaps = 6/137 (4%)
Frame = +1
Query: 22 TLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKR-PPRLILKPPC 80
TL + + +LI +IL +L VK+I+R + K W +LISDP F +R P L+
Sbjct: 151 TLHIDMDRDLIIEILLMLPVKSIVRFKAVCKLWRSLISDPLFASLHFQRAAPSLLFADS- 327
Query: 81 WVYPMRTIQ-SLPITRLLKNPSITVSGDFCYDGGGLNDNC-KVVISCNGLLCFIFCSDNK 138
+ +RTI P+ + I Y L NC + SC G L + S N
Sbjct: 328 --HAIRTIDLEGPLQSHRVSQPINCHFLSNYHDLSLPRNCISIHGSCRGFLLITWIS-NI 498
Query: 139 EYHNYW---FRIWNPAT 152
Y + W +WNP+T
Sbjct: 499 GYGHPWNDSLYLWNPST 549
>AV427033
Length = 420
Score = 46.6 bits (109), Expect = 4e-06
Identities = 21/46 (45%), Positives = 30/46 (64%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP 72
LP +L+ +IL L V ++L+ C+ KSWN+LISDP F + L P
Sbjct: 124 LPFDLVVEILCRLPVNSLLQFRCVCKSWNSLISDPKFAKNHLHCSP 261
>TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (SLF)-S4
protein, partial (10%)
Length = 551
Score = 46.6 bits (109), Expect = 4e-06
Identities = 22/42 (52%), Positives = 31/42 (73%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL 68
LP+ELI +ILS L VK++L+ +SK+W + ISDP FV+ L
Sbjct: 21 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHL 146
Score = 31.2 bits (69), Expect = 0.19
Identities = 31/122 (25%), Positives = 45/122 (36%)
Frame = +3
Query: 146 RIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGCGI 205
R WNPAT T S+ + + R+L FG + C
Sbjct: 252 RFWNPATRTMSQ----DSPPSWSPRNLHLGFG-----------------------YDCS- 347
Query: 206 LSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDSFPVIPLISPFNNGVYLSG 265
S TYK V P + V + ++ D+CWR I P P+ + VY+S
Sbjct: 348 -SDTYKVVGMI----------PGLTMVNVYNMGDNCWRTIQISPHAPMHLQ-GSAVYVSN 491
Query: 266 TI 267
T+
Sbjct: 492 TL 497
>AV768207
Length = 374
Score = 46.2 bits (108), Expect = 6e-06
Identities = 22/43 (51%), Positives = 30/43 (69%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK 69
LP EL +IL L VK++L+L C+ KSW +LISDP F + L+
Sbjct: 73 LPFELGVEILCRLPVKSLLQLRCVCKSWKSLISDPKFAKNHLR 201
>TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific F-box
protein a, partial (9%)
Length = 1602
Score = 46.2 bits (108), Expect = 6e-06
Identities = 22/44 (50%), Positives = 32/44 (72%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKR 70
L +ELI +ILSLL VK+++R C+ KSW IS+P F++ L+R
Sbjct: 78 LIDELIWEILSLLPVKSLVRFRCVCKSWKLTISNPQFMKLHLRR 209
Score = 36.6 bits (83), Expect = 0.005
Identities = 18/45 (40%), Positives = 24/45 (53%), Gaps = 6/45 (13%)
Frame = +1
Query: 229 RSQVRILSLSDDCWRNIDSFPVIPL------ISPFNNGVYLSGTI 267
+S + ++ D CWR I SFP +P NGVYL+GTI
Sbjct: 724 QSVANVYNMGDGCWRRIQSFPDFTREHWGRGTAPDQNGVYLNGTI 858
>BP050476
Length = 520
Score = 44.3 bits (103), Expect = 2e-05
Identities = 38/129 (29%), Positives = 55/129 (42%), Gaps = 13/129 (10%)
Frame = +2
Query: 56 TLISDPTFVQKQLKRPPRL------ILKPP-------CWVYPMRTIQSLPITRLLKNPSI 102
T D +FV+ L R P+ I P WV P + L+++ S
Sbjct: 5 TYHDDSSFVKLHLNRSPKNTHILLNIANDPYDFENDDTWVVPSS------VCCLIEDLSS 166
Query: 103 TVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWFRIWNPATGTRSKALGTN 162
+ CY L D V+ S NGL+CF D +W ++WNPAT +SK T
Sbjct: 167 MIDAKGCYL---LKDGHLVIGSSNGLICFGNFYDVGPIEEFWVQLWNPATHLKSKKSPT- 334
Query: 163 HDYNLQLRS 171
+NL +R+
Sbjct: 335 --FNLSMRT 355
>BP076687
Length = 389
Score = 43.9 bits (102), Expect = 3e-05
Identities = 22/37 (59%), Positives = 27/37 (72%)
Frame = -3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTF 63
LPEELI QIL L V+++LR + KSW +LISDP F
Sbjct: 381 LPEELIPQILLRLPVRSLLRFKSVCKSWLSLISDPKF 271
>AV413010
Length = 416
Score = 43.1 bits (100), Expect = 5e-05
Identities = 22/57 (38%), Positives = 36/57 (62%)
Frame = +3
Query: 9 IKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQ 65
+ ++++ + +LS LP+ELI QIL L V ++LR +SKSW + IS+P F +
Sbjct: 42 MNEQRRSCTEENPSLSSILPDELIIQILLQLPVTSLLRFKSVSKSWFSQISNPQFAK 212
>AV414925
Length = 414
Score = 41.2 bits (95), Expect = 2e-04
Identities = 34/119 (28%), Positives = 53/119 (43%), Gaps = 9/119 (7%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
+P E+IA ILS L V ++LR +SKSW +LI F++ L P
Sbjct: 97 IPVEVIADILSRLPVTSLLRFRSISKSWRSLIDSKHFMKLHLNN---------SLTNPNP 249
Query: 87 TIQSLPITRLLKNPSITVSGDFCYDGGGLNDN---------CKVVISCNGLLCFIFCSD 136
+ +L +L++ + DF G +N N ++ SC+GLLC +D
Sbjct: 250 NLTNL----ILRHNTDLYRADFPSIGAAVNLNHPLMCYSNRINILGSCHGLLCICNVAD 414
>AI967453
Length = 456
Score = 40.8 bits (94), Expect = 2e-04
Identities = 28/81 (34%), Positives = 42/81 (51%), Gaps = 1/81 (1%)
Frame = +1
Query: 31 LIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKR-PPRLILKPPCWVYPMRTIQ 89
LI +IL L VK+++R + K W +LISDP F +R PRL+++ + M +
Sbjct: 16 LITEILLRLPVKSLVRFKAVCKFWRSLISDPHFATSHFERAAPRLLIRTDHGIRTMDLEE 195
Query: 90 SLPITRLLKNPSITVSGDFCY 110
SL R+ S + DF Y
Sbjct: 196 SLHPDRI----SELIDYDFPY 246
>BP051299
Length = 539
Score = 40.8 bits (94), Expect = 2e-04
Identities = 28/104 (26%), Positives = 45/104 (42%)
Frame = +2
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
LP+E+ A+IL L KT+++ + KSW +LI+ +F+ P +L C R
Sbjct: 5 LPQEIWAEILHRLPPKTLVKCTSVCKSWRSLITSTSFISLHRNHSPSSLLLQLC---DER 175
Query: 87 TIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLC 130
I + L ++ N VV C+GL+C
Sbjct: 176 AIPNFIHYSLRRDDPFLSESSSLRLPSSFNREFSVVGICHGLVC 307
>TC18730 similar to UP|Q963J2 (Q963J2) Voltage-dependent calcium channel
alpha13 subunit (Fragment), partial (5%)
Length = 515
Score = 36.2 bits (82), Expect = 0.006
Identities = 18/35 (51%), Positives = 24/35 (68%)
Frame = +2
Query: 29 EELIAQILSLLDVKTILRLICLSKSWNTLISDPTF 63
+ELI IL L V+++LR + KSW +LISDP F
Sbjct: 227 KELILHILLRLPVRSLLRFKSVCKSWLSLISDPKF 331
>BI418251
Length = 546
Score = 30.8 bits (68), Expect = 0.25
Identities = 24/86 (27%), Positives = 44/86 (50%), Gaps = 3/86 (3%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTL-ISDPT--FVQKQLKRPPRLILKPPCWVY 83
LP+E++ QILS L + + L K W++L +S PT F + + +L L+ +
Sbjct: 52 LPDEVLCQILSFLPTEDAVATSVLCKRWSSLWLSVPTLDFDDYRYLKGKKLKLQSSFINF 231
Query: 84 PMRTIQSLPITRLLKNPSITVSGDFC 109
I S + + +KN +++V + C
Sbjct: 232 VYAIILSRALHQPIKNFTLSVISEEC 309
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,178,581
Number of Sequences: 28460
Number of extensions: 107278
Number of successful extensions: 632
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 617
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 624
length of query: 268
length of database: 4,897,600
effective HSP length: 89
effective length of query: 179
effective length of database: 2,364,660
effective search space: 423274140
effective search space used: 423274140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146341.8


