

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
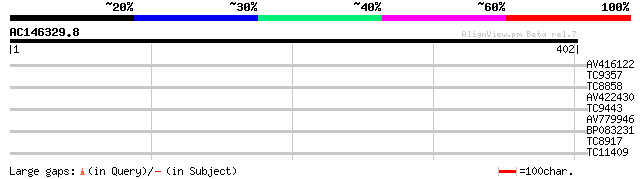
Query= AC146329.8 + phase: 0
(402 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV416122 29 1.2
TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein k... 29 1.6
TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 28 2.1
AV422430 28 2.7
TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting p... 28 2.7
AV779946 27 4.6
BP083231 27 4.6
TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, parti... 27 7.9
TC11409 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific ... 27 7.9
>AV416122
Length = 414
Score = 29.3 bits (64), Expect = 1.2
Identities = 32/82 (39%), Positives = 36/82 (43%), Gaps = 2/82 (2%)
Frame = -3
Query: 17 NSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYG--NCYSQLSKARLSLLVVAT 74
+SSSSSS + S+ SSSPS S S L SLA G S A LL+
Sbjct: 364 SSSSSSSSSSSLSSSSSSSSSPSNWTSLSGSLSSLAASLGWEEQSSVAETAGWLLLLSFE 185
Query: 75 SGTGFVLGSGGAVDLSMLSYTC 96
SG LGSG A L C
Sbjct: 184 SG----LGSGVAASFGALVGGC 131
>TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein kinase,
41K) , partial (71%)
Length = 1181
Score = 28.9 bits (63), Expect = 1.6
Identities = 12/18 (66%), Positives = 17/18 (93%)
Frame = +3
Query: 68 SLLVVATSGTGFVLGSGG 85
SLLV+++SG+GF+ GSGG
Sbjct: 255 SLLVISSSGSGFLDGSGG 308
>TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (65%)
Length = 1272
Score = 28.5 bits (62), Expect = 2.1
Identities = 30/109 (27%), Positives = 48/109 (43%), Gaps = 3/109 (2%)
Frame = -2
Query: 5 NWNCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSAS-KSNDLISLARHYGNCYSQLS 63
NWN + + SSSS+ NF S SS S++ S S S A S
Sbjct: 713 NWNISASAFFSLTILSSSSICNFFFCLSVLSSPASLALSANSRSQASRANRAFLNILIFS 534
Query: 64 KARLSLLVVATSGTGFVL--GSGGAVDLSMLSYTCLGTMMVAASASTLN 110
A +S+ ++A+ T +L S GA+ L +++ + ++S T N
Sbjct: 533 SATISIFLIASWTTSAILLHDSCGAISLPPSAFSRVLGESTSSSWKTTN 387
>AV422430
Length = 415
Score = 28.1 bits (61), Expect = 2.7
Identities = 19/59 (32%), Positives = 29/59 (48%), Gaps = 4/59 (6%)
Frame = +2
Query: 103 AASASTLNQVFEIKNDAIMR--RTSQRPLPSGRITVPH--AVGWASSVGLAGTALLATQ 157
A +N V ++N + R R QR LP+G++ PH + +AS + ALL Q
Sbjct: 5 AGDMRLMNPVSLVENHSACRTNRAGQRVLPNGQVWKPHLLLLSFASKILAEANALLKLQ 181
>TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (41%)
Length = 1025
Score = 28.1 bits (61), Expect = 2.7
Identities = 28/95 (29%), Positives = 44/95 (45%), Gaps = 4/95 (4%)
Frame = -1
Query: 5 NWNCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPS--VSASKSNDLISLARHYGNCYSQL 62
NWN + + SSSS +F S FSS+ S +SAS + R + N
Sbjct: 338 NWNMSASAFLSLTIFSSSSTCDFFFCLSIFSSATSLALSASSRSQASRAKRAFLNILI-F 162
Query: 63 SKARLSLLVVATSGTGFVL--GSGGAVDLSMLSYT 95
S A +S+ ++A+ T +L S GA L + ++
Sbjct: 161 SSATISIFLIAS*TTSAILLQASCGATTLPLSPFS 57
>AV779946
Length = 533
Score = 27.3 bits (59), Expect = 4.6
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 4/37 (10%)
Frame = -1
Query: 184 HPVNTWVGAVVGAIPP----LLGWAAASGDISLNSLI 216
H V T+ GAV A P +LGW +ASG I+ +L+
Sbjct: 467 HMVRTFSGAVAPAGSPHQRSMLGWRSASGTITGITLV 357
>BP083231
Length = 366
Score = 27.3 bits (59), Expect = 4.6
Identities = 12/36 (33%), Positives = 21/36 (58%), Gaps = 1/36 (2%)
Frame = +3
Query: 25 HNFTHIRSTFSSSP-SVSASKSNDLISLARHYGNCY 59
H ++ +S SS+P S+S+ N + + +HY CY
Sbjct: 207 HQCSNCKSYNSSAPISISSGYVNQAVVITKHYEKCY 314
>TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, partial (3%)
Length = 676
Score = 26.6 bits (57), Expect = 7.9
Identities = 26/80 (32%), Positives = 37/80 (45%)
Frame = -2
Query: 17 NSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATSG 76
+S SSS FT + STF+SS +S + S + + + LSK LL ++ G
Sbjct: 429 SSLGSSSGFGFTSLVSTFTSSVVAFSSLGSACTSSSGFFASSSFSLSKVLRDLL--SSLG 256
Query: 77 TGFVLGSGGAVDLSMLSYTC 96
V S L+ LS TC
Sbjct: 255 VSSVFFS----TLASLSCTC 208
>TC11409 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific protein
kinase-like, partial (8%)
Length = 809
Score = 26.6 bits (57), Expect = 7.9
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = -2
Query: 23 SLHNFTHIRSTF 34
SLHNFTH+RS F
Sbjct: 103 SLHNFTHLRSRF 68
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,297,510
Number of Sequences: 28460
Number of extensions: 104237
Number of successful extensions: 744
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 665
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 705
length of query: 402
length of database: 4,897,600
effective HSP length: 92
effective length of query: 310
effective length of database: 2,279,280
effective search space: 706576800
effective search space used: 706576800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146329.8


