

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.5 + phase: 0
(639 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
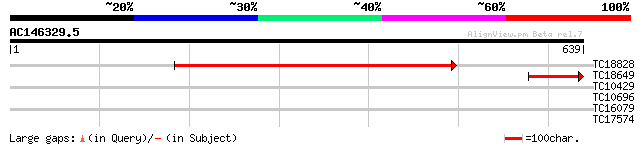
TC18828 homologue to PIR|A43350|A43350 formate-tetrahydrofolate ... 593 e-170
TC18649 homologue to PIR|A43350|A43350 formate-tetrahydrofolate ... 121 3e-28
TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%) 30 1.6
TC10696 weakly similar to UP|SYFB_ARATH (Q9SGE9) Probable phenyl... 28 4.5
TC16079 similar to UP|O04077 (O04077) Sucrose transport protein,... 28 4.5
TC17574 similar to UP|Q8ITA6 (Q8ITA6) Ribosomal protein S15 (Fra... 27 7.7
>TC18828 homologue to PIR|A43350|A43350 formate-tetrahydrofolate ligase -
spinach {Spinacia oleracea;} , partial (49%)
Length = 946
Score = 593 bits (1528), Expect = e-170
Identities = 293/314 (93%), Positives = 305/314 (96%)
Frame = +1
Query: 185 PNKEGKRSFSDVMFRRLKKFGISKTNPDDLTPEEVNKFARLDIDPDSITWRRVMDINDRF 244
PNKEGKRSF+DVMFRRLKK GISKTNPDDLTPEEV KFARLDIDP+SITWRRVMDINDRF
Sbjct: 4 PNKEGKRSFNDVMFRRLKKLGISKTNPDDLTPEEVTKFARLDIDPNSITWRRVMDINDRF 183
Query: 245 LRKITVGQGPDEKGMVRETAFDISVASEIMAVLALTTSLTDMRERLGKMVIGNSKSGDPV 304
LRKITVGQGPDEKGMVRET FDISVASEIMAVLALTTSL DMRERLGKMVIGNSKSGDP+
Sbjct: 184 LRKITVGQGPDEKGMVRETGFDISVASEIMAVLALTTSLADMRERLGKMVIGNSKSGDPI 363
Query: 305 TADDLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVG 364
TADDLG+GGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVG
Sbjct: 364 TADDLGVGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVG 543
Query: 365 PGGFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMHGGGPAVVAGKPLD 424
GGFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAI+VATIRALKMHGGGPAVVAGKPLD
Sbjct: 544 KGGFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIVVATIRALKMHGGGPAVVAGKPLD 723
Query: 425 HAYLTENVALVEAGCANLARHILNSKAYGANVIVAINKFSTDTDAELNAVKKAALDAGAY 484
HAYLTENVALVEAGC N+ARHI+N+KAYG NV+VAINKFSTDT+AELNAV+ AAL AGAY
Sbjct: 724 HAYLTENVALVEAGCVNMARHIVNTKAYGVNVVVAINKFSTDTEAELNAVRNAALAAGAY 903
Query: 485 DAVICSHHAHGGRG 498
DAVICSHHAHGG+G
Sbjct: 904 DAVICSHHAHGGKG 945
>TC18649 homologue to PIR|A43350|A43350 formate-tetrahydrofolate ligase -
spinach {Spinacia oleracea;} , partial (10%)
Length = 404
Score = 121 bits (304), Expect = 3e-28
Identities = 58/61 (95%), Positives = 60/61 (98%)
Frame = +3
Query: 579 AAAKGAPTGFILPIRDVRASIGAGFIYPLVGTMSTMPGLPTRPCFYDIDLDTTTGQVIGL 638
AAAKGAP+GFILPIRDVRASIGAGFIYPLVGTMSTMPGLPTRPCFYDIDLDT TG+VIGL
Sbjct: 3 AAAKGAPSGFILPIRDVRASIGAGFIYPLVGTMSTMPGLPTRPCFYDIDLDTATGKVIGL 182
Query: 639 S 639
S
Sbjct: 183 S 185
>TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%)
Length = 1742
Score = 29.6 bits (65), Expect = 1.6
Identities = 16/49 (32%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Frame = +3
Query: 130 GGYSQVIPMDEFN-LHLTGDIHAITAANNLLAAAIDTRIFHESTQSDKA 177
GG + ++ DE N L+ D+HA+ A + AA I+ + + ST + A
Sbjct: 1311 GGAAMILMNDETNAFSLSADVHALPATHVSYAAGIEIKAYINSTATPTA 1457
>TC10696 weakly similar to UP|SYFB_ARATH (Q9SGE9) Probable phenylalanyl-tRNA
synthetase beta chain (Phenylalanine--tRNA ligase beta
chain) (PheRS) , partial (30%)
Length = 941
Score = 28.1 bits (61), Expect = 4.5
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = -3
Query: 159 LAAAIDTRIFHESTQSDKALFNRLCPPNKE 188
L A +T I+H+ Q + F LCPP+ E
Sbjct: 441 LTAQYNTPIYHQMKQRESHSFPLLCPPSHE 352
>TC16079 similar to UP|O04077 (O04077) Sucrose transport protein, partial
(18%)
Length = 573
Score = 28.1 bits (61), Expect = 4.5
Identities = 14/45 (31%), Positives = 23/45 (51%)
Frame = -1
Query: 263 TAFDISVASEIMAVLALTTSLTDMRERLGKMVIGNSKSGDPVTAD 307
T+ V S+ +A++ALT ++T GK+ N S P+ D
Sbjct: 276 TSAGFGVGSKTIAIMALTAAITAPTTNAGKLPPPNKASQGPLNVD 142
>TC17574 similar to UP|Q8ITA6 (Q8ITA6) Ribosomal protein S15 (Fragment),
partial (87%)
Length = 523
Score = 27.3 bits (59), Expect = 7.7
Identities = 18/62 (29%), Positives = 28/62 (45%), Gaps = 4/62 (6%)
Frame = -3
Query: 22 DIDIANSVQPIHISEIANHL----NLTPNHYDLYGKYKAKVLLSALDELQESKDGYYVVV 77
D+D+ + +H ++ ANHL ++ D G K LL +L EL + G V
Sbjct: 320 DLDLVEHLAVVHTNDGANHLWYDDHVPQVRLDRGGLLKLGALLLSLPELLDESHGLPVQA 141
Query: 78 GG 79
G
Sbjct: 140 AG 135
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,210,062
Number of Sequences: 28460
Number of extensions: 112860
Number of successful extensions: 477
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 477
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 477
length of query: 639
length of database: 4,897,600
effective HSP length: 96
effective length of query: 543
effective length of database: 2,165,440
effective search space: 1175833920
effective search space used: 1175833920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146329.5


