

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
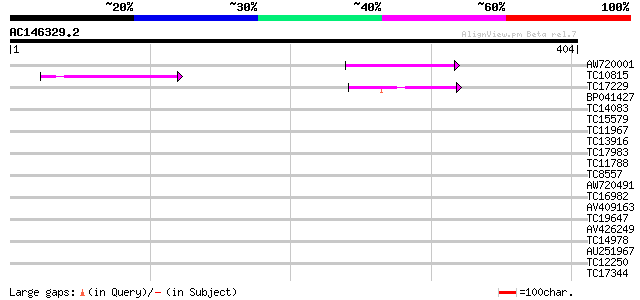
Query= AC146329.2 + phase: 0
(404 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW720001 61 3e-10
TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g1... 44 5e-05
TC17229 similar to UP|O49595 (O49595) HMG protein (AT3G51880/ORF... 40 7e-04
BP041427 37 0.006
TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor... 35 0.022
TC15579 weakly similar to UP|Q8L7T9 (Q8L7T9) AT5g54930/MBG8_20, ... 34 0.038
TC11967 homologue to UP|AAS09978 (AAS09978) MYB transcription fa... 32 0.19
TC13916 homologue to UP|Q9FK48 (Q9FK48) Similarity to transcript... 29 1.2
TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , part... 29 1.2
TC11788 weakly similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 prote... 29 1.2
TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete 29 1.6
AW720491 28 2.1
TC16982 weakly similar to GB|AAO64893.1|29029028|BT005958 At1g54... 28 2.1
AV409163 28 2.7
TC19647 weakly similar to UP|N551_SOYBN (Q05544) Early nodulin 5... 28 2.7
AV426249 28 2.7
TC14978 weakly similar to UP|Q7S392 (Q7S392) Predicted protein, ... 28 2.7
AU251967 28 2.7
TC12250 weakly similar to GB|AAP21346.1|30102856|BT006538 At3g60... 28 3.6
TC17344 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/F... 27 4.7
>AW720001
Length = 542
Score = 61.2 bits (147), Expect = 3e-10
Identities = 31/82 (37%), Positives = 46/82 (55%), Gaps = 1/82 (1%)
Frame = +3
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
+KK + KK+DP PK SG+ FF + LK + G ++ R +GE W K+ EK
Sbjct: 138 KKKKQKKKKDPNAPKRAMSGFMFFSQMERENLKKTNPGISFTDVGRVLGEKWKKMSTEEK 317
Query: 299 AVYQDKAVKDKERYITEMEYYR 320
Y+ KA DK+RY+ E+ Y+
Sbjct: 318 EPYEVKARADKKRYMDEISGYK 383
>TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g17400),
partial (18%)
Length = 1027
Score = 43.9 bits (102), Expect = 5e-05
Identities = 27/101 (26%), Positives = 46/101 (44%)
Frame = +2
Query: 23 EYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFE 82
+Y P A+ E + F+ LE + +F P G L+ +L+ V GG++
Sbjct: 506 DYDEPGASLER-----EAFMKELENFYRERSLEFKPPKFYGEPLNCLKLWRAVIRLGGYD 670
Query: 83 KIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQ 123
+ + W++V F+ P T T S+ R +Y L YE+
Sbjct: 671 VVTGSKLWRQVGESFHPPKTCTTVSWTFRIFYEKALLEYEK 793
>TC17229 similar to UP|O49595 (O49595) HMG protein (AT3G51880/ORF13),
partial (66%)
Length = 540
Score = 40.0 bits (92), Expect = 7e-04
Identities = 26/88 (29%), Positives = 40/88 (44%), Gaps = 7/88 (7%)
Frame = +3
Query: 242 KSEMKKRDPAHPKPNRSGYNFF-------FAEQHPRLKPLHRGKDREISRTIGELWNKLP 294
K++ K+DP PK S + F F ++P +K + + + GE W L
Sbjct: 231 KAKKAKKDPNKPKRPPSAFFVFLEDFRKTFKAENPNVKGVSA-----VGKAGGEKWKSLT 395
Query: 295 ESEKAVYQDKAVKDKERYITEMEYYREK 322
++EKA Y+ KA K K Y + Y K
Sbjct: 396 KAEKAPYEAKAAKRKAEYEKLINAYNNK 479
>BP041427
Length = 527
Score = 37.0 bits (84), Expect = 0.006
Identities = 15/32 (46%), Positives = 22/32 (67%)
Frame = -1
Query: 175 GSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLY 206
G++V G +DG FDSG L+T T+ +GVL+
Sbjct: 500 GAEVRGKVDGAFDSGLLMTATVNGRVFRGVLF 405
>TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor, partial
(92%)
Length = 1693
Score = 35.0 bits (79), Expect = 0.022
Identities = 25/94 (26%), Positives = 44/94 (46%), Gaps = 1/94 (1%)
Frame = +1
Query: 279 DREISRTIGE-LWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQ 337
D E ++ + E W K ++EKA +++ K +E E+ K D SDA
Sbjct: 1123 DPEYAKQVAEETWGKQKDAEKAAFEEAEKKREE----------EESKEDPADSDAED--- 1263
Query: 338 RLPEPDTDMLNAEADSLQTPEQSSLDGSDDYEDD 371
E DTD + ++D ++ ++ D +D E+D
Sbjct: 1264 ---EEDTDEASHDSDDAESKTEAGEDSADANEED 1356
>TC15579 weakly similar to UP|Q8L7T9 (Q8L7T9) AT5g54930/MBG8_20, partial
(27%)
Length = 592
Score = 34.3 bits (77), Expect = 0.038
Identities = 21/50 (42%), Positives = 32/50 (64%), Gaps = 10/50 (20%)
Frame = +3
Query: 176 SQVV-GVIDGKFDSGYLVTVTIGSE--KLKGVLYQ-------APQNPVLP 215
SQVV GVI+ FD+GYL++V +G L+GV+++ +P+N V P
Sbjct: 432 SQVVSGVIEAVFDAGYLLSVRVGDSDTTLRGVVFKPGRYVPISPENDVAP 581
>TC11967 homologue to UP|AAS09978 (AAS09978) MYB transcription factor,
partial (32%)
Length = 664
Score = 32.0 bits (71), Expect = 0.19
Identities = 36/139 (25%), Positives = 58/139 (40%), Gaps = 8/139 (5%)
Frame = +3
Query: 87 DRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYFKARDWTNTTSDVLQSQSSI 146
DR WK++ +F+ K + H ++ Y+ K + N TS+ +
Sbjct: 297 DRDWKKIE------------AFIGSKTVIQIRSHAQK-YFLKVQK--NGTSE------HV 413
Query: 147 PAPAPKMQFSHPSPQVQP-------AVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIGSE 199
P P PK + +HP PQ P A P+ SSSA + + DS ++ + S
Sbjct: 414 PPPRPKRKAAHPYPQKAPKNAPTSQATGPLQSSSAFIEPAYIYSS--DSSSVLGTPVTSV 587
Query: 200 KLKGVLYQA-PQNPVLPAS 217
L Y+A P + V P +
Sbjct: 588 PLSSWNYRAIPSSSVPPGT 644
>TC13916 homologue to UP|Q9FK48 (Q9FK48) Similarity to transcription
regulator, partial (17%)
Length = 603
Score = 29.3 bits (64), Expect = 1.2
Identities = 12/47 (25%), Positives = 28/47 (59%)
Frame = +2
Query: 278 KDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLK 324
K +E +WNK+ +++ A ++K D ++ I +++ YR+++K
Sbjct: 203 KVQEGVEVFDSIWNKVYDTDNANQKEKFEADLKKEIKKLQRYRDQIK 343
>TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , partial (42%)
Length = 519
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/43 (39%), Positives = 20/43 (45%)
Frame = +2
Query: 134 NTTSDVLQSQSSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGS 176
+TT S SS +P+ Q S PSP P V SSS S
Sbjct: 239 STTPSTPSSSSSPTSPSTTKQISTPSPSPPPPVATAASSSKNS 367
>TC11788 weakly similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 protein, partial
(15%)
Length = 469
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/45 (37%), Positives = 24/45 (52%)
Frame = +1
Query: 236 RRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDR 280
R RR + R P P+P RS A++HPR + LH+G+ R
Sbjct: 115 RLNRRLTPL*RHRQPPSPQPPRSKVP---AQRHPR*RRLHQGRHR 240
>TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete
Length = 1064
Score = 28.9 bits (63), Expect = 1.6
Identities = 34/151 (22%), Positives = 56/151 (36%), Gaps = 13/151 (8%)
Frame = +3
Query: 234 VHRRRRRKKSEMKKRD--PAHPKPNRSGYNFFFAEQ---------HPRLKPLHRGKDREI 282
+H RRR K + + PA + G F Q H LKP G+D +
Sbjct: 129 LHYRRRLKMTTAARPTWAPAKGGNEQGGTRIFGPSQKYSSRDIASHTTLKPRRDGQDTQD 308
Query: 283 SRTIGELWNKLPESEKAVY--QDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLP 340
L ++L E E+ + ++K+ D + + E K D + D + R
Sbjct: 309 ELKRRNLRDELEERERRHFSSKNKSYSDDRDHGKSSHLFLEGTKRD--VEDHIVARS--- 473
Query: 341 EPDTDMLNAEADSLQTPEQSSLDGSDDYEDD 371
++A+ ++ D DD EDD
Sbjct: 474 ------VDADDSDVEVKSDEESDDDDDDEDD 548
>AW720491
Length = 509
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = +1
Query: 235 HRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRL 271
HR RRR+ +K+ HP P R F +QH ++
Sbjct: 148 HRHRRRRPLRLKENRRRHPPPQRLELPPFPHQQHQQI 258
>TC16982 weakly similar to GB|AAO64893.1|29029028|BT005958 At1g54570
{Arabidopsis thaliana;}, partial (6%)
Length = 878
Score = 28.5 bits (62), Expect = 2.1
Identities = 22/82 (26%), Positives = 34/82 (40%), Gaps = 8/82 (9%)
Frame = +1
Query: 200 KLKGVLYQAPQNPVLPASHHSV--------PANNNNVTASVGVHRRRRRKKSEMKKRDPA 251
KLK VL+Q+ NP S V PA S G + + + M++ +
Sbjct: 199 KLKSVLFQST-NPRFAVSVDRVSAMAEKGSPAEMKREERSSGAAQSEKNRWEGMEEEETE 375
Query: 252 HPKPNRSGYNFFFAEQHPRLKP 273
K RSG+ +F + +KP
Sbjct: 376 KVKQRRSGWKDYFEQAKELIKP 441
>AV409163
Length = 416
Score = 28.1 bits (61), Expect = 2.7
Identities = 17/62 (27%), Positives = 28/62 (44%), Gaps = 12/62 (19%)
Frame = +2
Query: 210 QNPVLPASHHSVPAN------------NNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNR 257
++PVLP S ++ P++ NN+T+S G+HR + P HP+
Sbjct: 20 ESPVLPHSRNNQPSSPESARSRSFSSPGNNLTSSGGIHR---------EGTSPVHPRSTS 172
Query: 258 SG 259
G
Sbjct: 173 EG 178
>TC19647 weakly similar to UP|N551_SOYBN (Q05544) Early nodulin 55-1
precursor (N-55-1) (Fragment), partial (15%)
Length = 494
Score = 28.1 bits (61), Expect = 2.7
Identities = 14/36 (38%), Positives = 17/36 (46%)
Frame = -1
Query: 146 IPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGV 181
+PAP P F P + Q A F G SS G + V
Sbjct: 326 VPAPPPPAGFCFPPKKQQAATFGEGGSSNGGATLTV 219
>AV426249
Length = 421
Score = 28.1 bits (61), Expect = 2.7
Identities = 15/50 (30%), Positives = 26/50 (52%), Gaps = 4/50 (8%)
Frame = +3
Query: 337 QRLPEPDTDMLNAEADSLQTPEQSSLDGSDDYE----DDKAKEKDFSVDS 382
Q++ + DM+ A SL++ + SDD++ D K K FS+D+
Sbjct: 63 QQIKKEVVDMIKMHATSLESKQPQKAGASDDFQEFGPDGKPITKKFSMDT 212
>TC14978 weakly similar to UP|Q7S392 (Q7S392) Predicted protein, partial
(3%)
Length = 573
Score = 28.1 bits (61), Expect = 2.7
Identities = 11/35 (31%), Positives = 19/35 (53%)
Frame = -1
Query: 134 NTTSDVLQSQSSIPAPAPKMQFSHPSPQVQPAVFP 168
++ + + Q+ SS+P P P PSP +P+ P
Sbjct: 201 SSPNKISQNSSSLPPPPPSSSSHPPSPPAKPSYTP 97
>AU251967
Length = 380
Score = 28.1 bits (61), Expect = 2.7
Identities = 21/59 (35%), Positives = 28/59 (46%), Gaps = 15/59 (25%)
Frame = -3
Query: 224 NNNNVTASVGVHRRRR---------------RKKSEMKKRDPAHPKPNRSGYNFFFAEQ 267
NNNN SV V RRRR RKK + KK++ + R ++F FAE+
Sbjct: 225 NNNNNNNSVWVGRRRRETVNCEL*FLV*VILRKKKKKKKKN*RWIEIERREWSFGFAEK 49
>TC12250 weakly similar to GB|AAP21346.1|30102856|BT006538 At3g60590
{Arabidopsis thaliana;}, partial (58%)
Length = 440
Score = 27.7 bits (60), Expect = 3.6
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = -3
Query: 250 PAHPKPNRSGYNFFFAEQHPRLKPLHRGKD 279
PAH PNR Y + AE +P+L+ H+ +D
Sbjct: 252 PAH*PPNRMHYPCWRAELNPKLQSPHQLQD 163
>TC17344 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/F26K24_12
{Arabidopsis thaliana;}, partial (28%)
Length = 567
Score = 27.3 bits (59), Expect = 4.7
Identities = 17/56 (30%), Positives = 24/56 (42%)
Frame = +3
Query: 204 VLYQAPQNPVLPASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSG 259
V Y A + L H + + +HRRRRR+ P HP+P+R G
Sbjct: 96 VFYAATTDHSLEGRHRHLARESAARQQHQCLHRRRRRR--------PHHPRPSRYG 239
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,065,982
Number of Sequences: 28460
Number of extensions: 104290
Number of successful extensions: 651
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 637
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 647
length of query: 404
length of database: 4,897,600
effective HSP length: 92
effective length of query: 312
effective length of database: 2,279,280
effective search space: 711135360
effective search space used: 711135360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146329.2


