

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.15 - phase: 0
(204 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
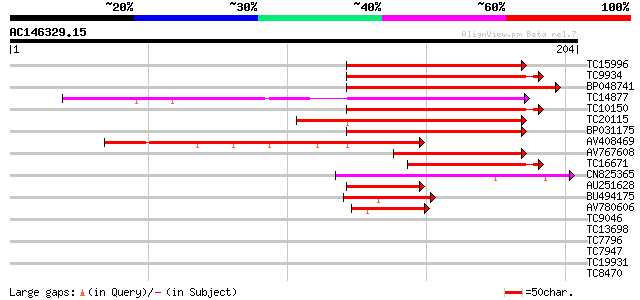
Sequences producing significant alignments: (bits) Value
TC15996 similar to UP|BAC98491 (BAC98491) AG-motif binding prote... 119 3e-28
TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protei... 118 8e-28
BP048741 116 2e-27
TC14877 similar to GB|AAP37701.1|30725358|BT008342 At4g32890 {Ar... 116 3e-27
TC10150 similar to UP|BAC98494 (BAC98494) AG-motif binding prote... 116 3e-27
TC20115 similar to PIR|T52104|T52104 GATA-binding transcription ... 115 5e-27
BP031175 114 1e-26
AV408469 104 1e-23
AV767608 76 5e-15
TC16671 similar to PIR|T52104|T52104 GATA-binding transcription ... 72 5e-14
CN825365 54 2e-08
AU251628 47 3e-06
BU494175 46 4e-06
AV780606 44 3e-05
TC9046 homologue to UP|Q9LRH6 (Q9LRH6) ZIM, partial (10%) 37 0.003
TC13698 similar to GB|AAO63291.1|28950735|BT005227 At4g20000 {Ar... 30 0.22
TC7796 homologue to gb|AF036495.1|AF036495 Hamamelis virginiana ... 30 0.29
TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, parti... 30 0.38
TC19931 weakly similar to UP|Q8L5A7 (Q8L5A7) Steroid sulfotransf... 29 0.50
TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase,... 29 0.65
>TC15996 similar to UP|BAC98491 (BAC98491) AG-motif binding protein-1,
partial (19%)
Length = 555
Score = 119 bits (299), Expect = 3e-28
Identities = 52/65 (80%), Positives = 56/65 (86%)
Frame = +2
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C HCE TKTPQWR GP GPKTLCNACGVRY+SGRL PEYRPAAS TF P +HSN HKK
Sbjct: 161 RKCMHCEVTKTPQWREGPMGPKTLCNACGVRYRSGRLFPEYRPAASPTFVPAVHSNCHKK 340
Query: 182 ILEMR 186
+LEMR
Sbjct: 341 VLEMR 355
>TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (37%)
Length = 1050
Score = 118 bits (295), Expect = 8e-28
Identities = 52/71 (73%), Positives = 61/71 (85%)
Frame = +2
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC+ KTPQWR GP GPKTLCNACGVR+KSGRL PEYRPA S TFS ++HSNSH+K
Sbjct: 569 RRCSHCQVQKTPQWRAGPLGPKTLCNACGVRFKSGRLFPEYRPACSPTFSGEIHSNSHRK 748
Query: 182 ILEMRVMRRKD 192
+LEMR RRK+
Sbjct: 749 VLEMR--RRKE 775
>BP048741
Length = 558
Score = 116 bits (291), Expect = 2e-27
Identities = 50/77 (64%), Positives = 60/77 (76%)
Frame = -1
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC KTPQWRTGP G KTLCNACGVR+KSGRL PEYRPA S TFS +LHSN H+K
Sbjct: 462 RRCSHCGVQKTPQWRTGPHGAKTLCNACGVRFKSGRLLPEYRPACSPTFSSELHSNHHRK 283
Query: 182 ILEMRVMRRKDNKNSGI 198
+LEMR + + +G+
Sbjct: 282 VLEMRRKKEEGEVETGL 232
>TC14877 similar to GB|AAP37701.1|30725358|BT008342 At4g32890 {Arabidopsis
thaliana;}, partial (25%)
Length = 1039
Score = 116 bits (290), Expect = 3e-27
Identities = 71/190 (37%), Positives = 94/190 (49%), Gaps = 22/190 (11%)
Frame = +3
Query: 20 PKSNSSPTCEKTTVRRTRSKRPRLA--------TFSSHHSTMQLIS-------------- 57
P + SP + + + RSKR RL+ + SHH Q +
Sbjct: 138 PLESESPLQKNSVPGKARSKRKRLSAPRTKDPLSMWSHHLNPQNEAFGPDPPLLKQDYWL 317
Query: 58 STSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNA 117
+ S + + + V + + +I K++ GE N + P
Sbjct: 318 ADSELIVKKEEQEVTTKEEQEVVVVVKKEIIVKEE-FEDGEVSNNGQNPMP--------- 467
Query: 118 LGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSN 177
R+CTHC + +TPQWR GP GPKTLCNACGVRYKSGRL PEYRPA S TF LHSN
Sbjct: 468 ----RRCTHCLSQRTPQWRAGPLGPKTLCNACGVRYKSGRLHPEYRPAKSPTFVSFLHSN 635
Query: 178 SHKKILEMRV 187
SHKK++EMR+
Sbjct: 636 SHKKVMEMRM 665
>TC10150 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (22%)
Length = 552
Score = 116 bits (290), Expect = 3e-27
Identities = 51/71 (71%), Positives = 60/71 (83%)
Frame = +3
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC+ KTPQWRTGP G KTLCNACGVRYKSGRLC EYRPA S TF ++HSNSH+K
Sbjct: 63 RRCSHCQVQKTPQWRTGPLGAKTLCNACGVRYKSGRLCSEYRPACSPTFCGEIHSNSHRK 242
Query: 182 ILEMRVMRRKD 192
+LE+R RRK+
Sbjct: 243 VLEIR--RRKE 269
>TC20115 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (31%)
Length = 635
Score = 115 bits (288), Expect = 5e-27
Identities = 53/85 (62%), Positives = 67/85 (78%), Gaps = 2/85 (2%)
Frame = +3
Query: 104 KTKAPLLAALDHNALGL--VRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPE 161
K+++ AA++ + G VR+C+HC + KTPQWRTGP GPKTLCNACGVR+KSGRL PE
Sbjct: 39 KSRSKRAAAVEGSPAGEGGVRRCSHCASEKTPQWRTGPLGPKTLCNACGVRFKSGRLVPE 218
Query: 162 YRPAASSTFSPDLHSNSHKKILEMR 186
YRPAAS TF HSNSH+K++E+R
Sbjct: 219 YRPAASPTFVLSQHSNSHRKVMELR 293
>BP031175
Length = 537
Score = 114 bits (285), Expect = 1e-26
Identities = 50/65 (76%), Positives = 54/65 (82%)
Frame = -2
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C HC KTPQWRTGP GPKTLCNACGVRYKSGRL PEYRPAAS TF HSNSH+K
Sbjct: 524 RRCLHCATDKTPQWRTGPMGPKTLCNACGVRYKSGRLVPEYRPAASPTFMLTKHSNSHRK 345
Query: 182 ILEMR 186
+LE+R
Sbjct: 344 VLELR 330
>AV408469
Length = 429
Score = 104 bits (259), Expect = 1e-23
Identities = 67/127 (52%), Positives = 78/127 (60%), Gaps = 12/127 (9%)
Frame = +2
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGEN--MQDSVISNKGAST--EKFPDSQIAAK 90
R RSKRPR A+F+ S MQLIS SSF+GEN M+ + + S+ E F +SQ
Sbjct: 50 RARSKRPRPASFNPR-SAMQLISPASSFIGENNTMKPNYHHHVRTSSDSENFAESQPVIT 226
Query: 91 KQ---KLSSGESKKNKKTKAPL-LAALDHNALGL----VRQCTHCEATKTPQWRTGPEGP 142
K K +SGE KK KK K PL LA D N + VR+C HCE TKTPQWR GP GP
Sbjct: 227 KMMMPKQASGEPKKKKKIKVPLPLAPSDGNNIQNGSQPVRKCMHCEITKTPQWRAGPMGP 406
Query: 143 KTLCNAC 149
KTLCNAC
Sbjct: 407 KTLCNAC 427
>AV767608
Length = 398
Score = 75.9 bits (185), Expect = 5e-15
Identities = 34/48 (70%), Positives = 40/48 (82%)
Frame = -1
Query: 139 PEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
P GPKTLCNACGVR+KSGRL E RPA+S TF +LHSNS +K++EMR
Sbjct: 398 PHGPKTLCNACGVRFKSGRLVLENRPASSPTFRAELHSNSPRKVMEMR 255
>TC16671 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (20%)
Length = 684
Score = 72.4 bits (176), Expect = 5e-14
Identities = 35/49 (71%), Positives = 40/49 (81%)
Frame = +2
Query: 144 TLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKD 192
TLCNACGVR+KSGRL EYRPAAS TF HSNSH+K+LE+R R+KD
Sbjct: 2 TLCNACGVRFKSGRLVAEYRPAASPTFVSARHSNSHRKVLELR--RQKD 142
>CN825365
Length = 511
Score = 53.5 bits (127), Expect = 2e-08
Identities = 33/97 (34%), Positives = 44/97 (45%), Gaps = 11/97 (11%)
Frame = +3
Query: 118 LGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPD---- 173
+G C HC T TP WR GP LCNACG R+++ Y P S + D
Sbjct: 147 MGKQGPCYHCGVTSTPLWRNGPPEKPVLCNACGSRWRTKGTLANYTPLHSRADNVDNDDQ 326
Query: 174 ----LHSNSHKKILEMRVMRRK---DNKNSGILALEY 203
L + S K E++ ++RK DN SG +Y
Sbjct: 327 RVSRLKNMSLNKNKEVKFVKRKLNHDNAASGGFTPDY 437
>AU251628
Length = 350
Score = 46.6 bits (109), Expect = 3e-06
Identities = 17/28 (60%), Positives = 20/28 (70%)
Frame = +3
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNAC 149
+ C C +KTP WR GP GPK+LCNAC
Sbjct: 267 KTCADCGTSKTPLWRGGPAGPKSLCNAC 350
>BU494175
Length = 475
Score = 46.2 bits (108), Expect = 4e-06
Identities = 18/35 (51%), Positives = 26/35 (73%), Gaps = 2/35 (5%)
Frame = +2
Query: 121 VRQCTHCEATK--TPQWRTGPEGPKTLCNACGVRY 153
+R+C HC ++ TP R GP+GP+TLCNACG+ +
Sbjct: 143 LRRCQHCGVSENNTPAMRRGPDGPRTLCNACGLMW 247
>AV780606
Length = 585
Score = 43.5 bits (101), Expect = 3e-05
Identities = 17/30 (56%), Positives = 21/30 (69%), Gaps = 2/30 (6%)
Frame = -1
Query: 124 CTHC--EATKTPQWRTGPEGPKTLCNACGV 151
CTHC + TP R GP GP++LCNACG+
Sbjct: 435 CTHCGTSSKSTPMMRRGPSGPRSLCNACGL 346
>TC9046 homologue to UP|Q9LRH6 (Q9LRH6) ZIM, partial (10%)
Length = 629
Score = 36.6 bits (83), Expect = 0.003
Identities = 13/20 (65%), Positives = 16/20 (80%)
Frame = +3
Query: 132 TPQWRTGPEGPKTLCNACGV 151
TP R GP GP++LCNACG+
Sbjct: 3 TPMMRRGPSGPRSLCNACGL 62
>TC13698 similar to GB|AAO63291.1|28950735|BT005227 At4g20000 {Arabidopsis
thaliana;}, partial (10%)
Length = 419
Score = 30.4 bits (67), Expect = 0.22
Identities = 21/56 (37%), Positives = 29/56 (51%)
Frame = +2
Query: 99 SKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
S+ +KKT LL A N LV+Q T C +T T + +GP TL G R++
Sbjct: 245 SRSSKKTPTTLLNANTTNFRALVQQFTGCPSTTTMPSPSLYKGPITLNFQQGSRHQ 412
>TC7796 homologue to gb|AF036495.1|AF036495 Hamamelis virginiana large
subunit 26S ribosomal RNA gene, partial sequence,
complete
Length = 6006
Score = 30.0 bits (66), Expect = 0.29
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +3
Query: 126 HCEATKTPQWRTGPEGP 142
+C K PQWRTGP+ P
Sbjct: 2727 NCSLEKRPQWRTGPKSP 2777
>TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, partial (81%)
Length = 1294
Score = 29.6 bits (65), Expect = 0.38
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 4/100 (4%)
Frame = -2
Query: 21 KSNS-SPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGAST 79
K+NS SPTC ++ RR+RSK HH +IS S + + V+S++ T
Sbjct: 867 KNNSHSPTCNNSSTRRSRSK---------HHLCRSIISIRSVW-----ESQVLSHR--HT 736
Query: 80 EKFP---DSQIAAKKQKLSSGESKKNKKTKAPLLAALDHN 116
+ P DS+ ++ L N T L DHN
Sbjct: 735 DNVPPSIDSRFLNRRYNLL---GLANTNTHITFLVTDDHN 625
>TC19931 weakly similar to UP|Q8L5A7 (Q8L5A7) Steroid sulfotransferase-like
protein (At5g07010), partial (29%)
Length = 539
Score = 29.3 bits (64), Expect = 0.50
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 23 NSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSS 61
N S TC + ++R R +PR+ +SH S + + ST S
Sbjct: 155 NRSHTCNRKPIKRRRKFKPRM*GANSHSS*RERLDSTLS 271
>TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(29%)
Length = 807
Score = 28.9 bits (63), Expect = 0.65
Identities = 21/68 (30%), Positives = 30/68 (43%), Gaps = 6/68 (8%)
Frame = -3
Query: 119 GLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPD----- 173
G +C HC ++ P+ P+T S CP PA+ S+ SPD
Sbjct: 367 GCTPRCHHCPSS--------PQPPETTA--------SPHTCPPQ*PASPSSPSPDPATSP 236
Query: 174 -LHSNSHK 180
LHS+SH+
Sbjct: 235 CLHSSSHQ 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.124 0.353
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,535,016
Number of Sequences: 28460
Number of extensions: 47077
Number of successful extensions: 233
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 231
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 231
length of query: 204
length of database: 4,897,600
effective HSP length: 86
effective length of query: 118
effective length of database: 2,450,040
effective search space: 289104720
effective search space used: 289104720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146329.15


