

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
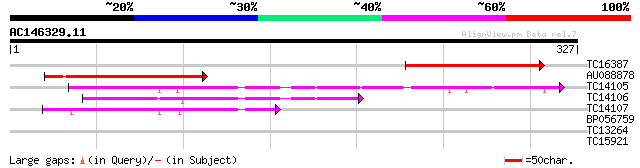
Query= AC146329.11 + phase: 0
(327 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16387 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, part... 149 6e-37
AU088878 133 5e-32
TC14105 similar to UP|Q42966 (Q42966) Nitrilase , partial (91%) 76 9e-15
TC14106 similar to UP|Q42966 (Q42966) Nitrilase , partial (87%) 53 6e-08
TC14107 homologue to UP|Q42966 (Q42966) Nitrilase , partial (39%) 42 1e-04
BP056759 27 3.6
TC13264 UP|Q9SWS9 (Q9SWS9) Ribosomal protein S26, partial (93%) 27 6.1
TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp ... 27 6.1
>TC16387 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (28%)
Length = 530
Score = 149 bits (376), Expect = 6e-37
Identities = 73/80 (91%), Positives = 75/80 (93%)
Frame = +3
Query: 229 EAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIV 288
EAHWEILLRARAIE+QCYVIAAAQAG HN+KRESYGDTLIIDPWGTVV RLPDR STG
Sbjct: 3 EAHWEILLRARAIESQCYVIAAAQAGIHNEKRESYGDTLIIDPWGTVVSRLPDRSSTGFA 182
Query: 289 VADIDLSLVDSVREKMPIAK 308
VADIDLSLVDSVREKMPIAK
Sbjct: 183 VADIDLSLVDSVREKMPIAK 242
>AU088878
Length = 302
Score = 133 bits (334), Expect = 5e-32
Identities = 72/94 (76%), Positives = 76/94 (80%)
Frame = +2
Query: 21 SIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLC 80
SI RASLS ES ATNS+RVAAA MTSI DLA+NF+TCSRLVKEA AGAKLLC
Sbjct: 23 SIPLSXRASLS-GAAESIXATNSLRVAAAXMTSINDLAANFTTCSRLVKEAXLAGAKLLC 199
Query: 81 FPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLA 114
FPEAFSFVGAKDGDSV A PLDGPIM+ YCSLA
Sbjct: 200 FPEAFSFVGAKDGDSVXXAXPLDGPIMEXYCSLA 301
>TC14105 similar to UP|Q42966 (Q42966) Nitrilase , partial (91%)
Length = 1446
Score = 75.9 bits (185), Expect = 9e-15
Identities = 80/330 (24%), Positives = 137/330 (41%), Gaps = 44/330 (13%)
Frame = +2
Query: 35 GESTMATNSVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAF-------- 85
G + A +VR Q ++I D + RLV EAA G++L+ FPEAF
Sbjct: 137 GSDSNAPTTVRATVVQASTIFYDTPATLDKAERLVAEAAGNGSQLVVFPEAFIGGYPRGS 316
Query: 86 ---SFVGAKDGDS-------VSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRH 135
+G++ S A + GP +D+ ++A + + L +G + G
Sbjct: 317 NFGVIIGSRTAKGREDFRKYHSSAIDVPGPEVDKLAAMAGKYKVHLVIGVIERDGYT--- 487
Query: 136 LFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSV 195
L+ T + D G +RKI +P F + G I D+P+G++G +
Sbjct: 488 LYCTALFFDSQGHYLGKHRKI------MPTAMERVMWGFGD-GSTIPVFDTPLGKIGAVI 646
Query: 196 CYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ--- 252
C++ + P L + + G +I P A + T W+ + A+E C+V++A Q
Sbjct: 647 CWENKMP-LLRTAMYAKGVEIYCAPTADDRKT----WQASMTHIALEGGCFVLSANQFIR 811
Query: 253 ------------AGTHNDKRES----YGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSL 296
+GT D G ++II P G V+ P+ ++ AD+DL
Sbjct: 812 RKDYPPPPEYVFSGTEEDLTPDSVVCVGGSVIISPLGNVLAG-PNYEGEALISADLDLGE 988
Query: 297 VDSVREKMPIA------KVIVCNVYIHPTN 320
+ + + +V+ +V HPTN
Sbjct: 989 IARAKLDFDVVGHYSRPEVLSLSVKDHPTN 1078
>TC14106 similar to UP|Q42966 (Q42966) Nitrilase , partial (87%)
Length = 1286
Score = 53.1 bits (126), Expect = 6e-08
Identities = 45/166 (27%), Positives = 77/166 (46%), Gaps = 4/166 (2%)
Frame = +2
Query: 43 SVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSI--- 98
+VR Q ++I D + RL+ EAA G++L+ FPEAF G G + I
Sbjct: 53 TVRATVVQASTIFYDTPATLDKAERLLAEAAGYGSQLVVFPEAF-IGGYPRGSNFGIIIG 229
Query: 99 AQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHL 158
+ GP +D+ ++A + + L +G + G L+ T + D G +RK+
Sbjct: 230 IRTAKGPEVDRLAAMAGKYKVHLVMGVIERDGYT---LYCTVLFFDSQGHYLGKHRKL-- 394
Query: 159 FDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPEL 204
+P G F + G I D+P+G++G +C++ + P L
Sbjct: 395 ----MPTGMERVMWGFGD-GSTIPVFDTPLGKIGAVICWENKMPLL 517
>TC14107 homologue to UP|Q42966 (Q42966) Nitrilase , partial (39%)
Length = 666
Score = 42.0 bits (97), Expect = 1e-04
Identities = 40/165 (24%), Positives = 73/165 (44%), Gaps = 28/165 (16%)
Frame = +3
Query: 20 TSIRRRLRASLSVTT---------GESTMATNSVRVAAAQMTSIT-DLASNFSTCSRLVK 69
TS ++++ +++S+ T G + A +VR Q ++I D + RL+
Sbjct: 150 TSTQKKMTSNISLVTTPPPPEVDMGSDSNAPTTVRATVVQASTIFYDTPATLDKAERLLA 329
Query: 70 EAASAGAKLLCFPEAF-------SFVGAKDGDSV-----------SIAQPLDGPIMDQYC 111
EAA +G++L+ FPEAF S G G+ S A + GP +D+
Sbjct: 330 EAAGSGSELVVFPEAFIGGYPRGSTFGMAVGNRTAKGREEFRKYHSSAIDVPGPEVDRLA 509
Query: 112 SLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKI 156
++A + + L +G + G L+ + + D G +RK+
Sbjct: 510 AMAGKYKVHLVMGVIERDGYT---LYCSVLFFDSQGHYLGKHRKL 635
>BP056759
Length = 551
Score = 27.3 bits (59), Expect = 3.6
Identities = 14/49 (28%), Positives = 27/49 (54%)
Frame = -3
Query: 15 NNTNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFST 63
+ T+G S+ +RAS + ++ EST+ T + A + SI L + ++
Sbjct: 387 SRTSGPSLLTTIRASNTTSSNESTLRTRLIWGAEKEAKSIPHLTARLAS 241
>TC13264 UP|Q9SWS9 (Q9SWS9) Ribosomal protein S26, partial (93%)
Length = 691
Score = 26.6 bits (57), Expect = 6.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +2
Query: 197 YDLRFPELYQLLRFQHGAQILLVPAAFTKVTG 228
Y+L FP LY+ L+FQ +L T V+G
Sbjct: 512 YELEFPLLYRFLKFQFCVGVL*FLVVVTCVSG 607
>TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp
{Saccharomyces cerevisiae;} , partial (10%)
Length = 580
Score = 26.6 bits (57), Expect = 6.1
Identities = 16/46 (34%), Positives = 20/46 (42%), Gaps = 2/46 (4%)
Frame = -2
Query: 186 SPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFT--KVTGE 229
SP+ + GL+ L F F HG Q + P FT K GE
Sbjct: 447 SPLKKKGLNAILQLFFEHTSHFTFFIHGGQYITTPNQFTIHKYLGE 310
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,504,825
Number of Sequences: 28460
Number of extensions: 66927
Number of successful extensions: 282
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 278
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 279
length of query: 327
length of database: 4,897,600
effective HSP length: 91
effective length of query: 236
effective length of database: 2,307,740
effective search space: 544626640
effective search space used: 544626640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146329.11


