

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145767.6 - phase: 0
(533 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
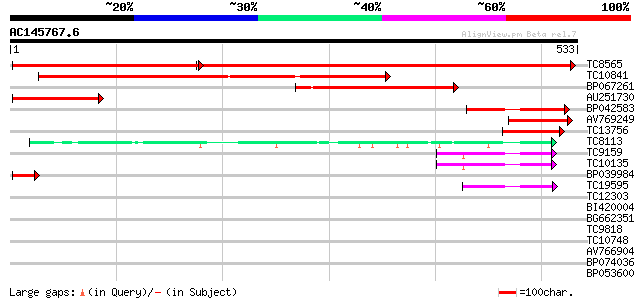
Score E
Sequences producing significant alignments: (bits) Value
TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, pa... 610 0.0
TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1,... 510 e-145
BP067261 251 2e-67
AU251730 129 2e-30
BP042583 109 1e-24
AV769249 90 1e-18
TC13756 similar to UP|Q9ARI9 (Q9ARI9) Phosphate transporter 1, p... 84 6e-17
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 57 7e-09
TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose tr... 50 7e-07
TC10135 44 7e-05
BP039984 44 9e-05
TC19595 43 1e-04
TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transpo... 39 0.002
BI420004 39 0.003
BG662351 38 0.004
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 37 0.006
TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter,... 37 0.008
AV766904 34 0.052
BP074036 32 0.26
BP053600 32 0.26
>TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial
(97%)
Length = 1913
Score = 610 bits (1573), Expect(2) = 0.0
Identities = 295/357 (82%), Positives = 326/357 (90%)
Frame = +2
Query: 176 LGGGIVALTVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMK 235
LG L V++ FDHKY VP+++E+PAAS+++P FDYVWR+ILMFGALPAALTYYWRMK
Sbjct: 620 LGWWNSGLIVSASFDHKYNVPSYQEDPAASMVLPAFDYVWRIILMFGALPAALTYYWRMK 799
Query: 236 MPETARYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHG 295
MPETARYTALVAKNAKQAA+DMSKVLQVELE EEEKVEK+ + +GLFSKQFA+RHG
Sbjct: 800 MPETARYTALVAKNAKQAASDMSKVLQVELEAEEEKVEKILESENQQFGLFSKQFASRHG 979
Query: 296 LALFGTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCS 355
+ L GT +TWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCS
Sbjct: 980 MHLLGTTTTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCS 1159
Query: 356 TVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFF 415
TVPGYWFTVAFID+MGRFAIQ+MGFFFMTVFMFALA+PYDHW+K++NR GFV +Y+LTFF
Sbjct: 1160TVPGYWFTVAFIDYMGRFAIQLMGFFFMTVFMFALALPYDHWTKKDNRFGFVAMYALTFF 1339
Query: 416 FANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKG 475
FANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYA+QSKDP KTD G
Sbjct: 1340FANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYASQSKDPAKTDPG 1519
Query: 476 YPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSR 532
YPTGIG++NSLIMLGVINF+GM+ T LVPESKGKSLEELSGE E +G EA E GSR
Sbjct: 1520YPTGIGVRNSLIMLGVINFLGMVFTFLVPESKGKSLEELSGETEEDGVEAIEAAGSR 1690
Score = 326 bits (836), Expect(2) = 0.0
Identities = 159/180 (88%), Positives = 170/180 (94%)
Frame = +3
Query: 3 GELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPG 62
G +GVLNALDVAKTQ YHFT IVIAGMGFFTDAYDLFCISLVTKLLGRIYYT+ +RPG
Sbjct: 102 GTIGVLNALDVAKTQWYHFTAIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTDITKSRPG 281
Query: 63 TLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSP 122
LP + Q+AVTGVAL GTLAGQLFFGWLGDK+GRK+VYGLTL++MVVCS+ASGLSFG+SP
Sbjct: 282 VLPANVQAAVTGVALCGTLAGQLFFGWLGDKMGRKRVYGLTLMIMVVCSIASGLSFGNSP 461
Query: 123 KSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVA 182
KSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVA
Sbjct: 462 KSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVA 641
>TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(59%)
Length = 991
Score = 510 bits (1314), Expect = e-145
Identities = 251/331 (75%), Positives = 280/331 (83%)
Frame = +3
Query: 28 GMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALVGTLAGQLFF 87
GMGFFTDAYDLFCISLVTKLLGRIYY +PGTLPP+ +AV GVA GTL+GQLFF
Sbjct: 3 GMGFFTDAYDLFCISLVTKLLGRIYYHVDGAAKPGTLPPNVSAAVNGVAFCGTLSGQLFF 182
Query: 88 GWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIGGDYPLS 147
GWLGDKLGRKKVYG+TL++MV+CS+ SGLSFG SPKSVMATLCFFRFWLGFGIGGDYPLS
Sbjct: 183 GWLGDKLGRKKVYGMTLMMMVICSIGSGLSFGHSPKSVMATLCFFRFWLGFGIGGDYPLS 362
Query: 148 ATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYKVPTFEENPAASLL 207
ATIMSEY+NKKTRGAFIAAVFAMQGFGILGGGI A+ +++ F K+ P +E +P S
Sbjct: 363 ATIMSEYSNKKTRGAFIAAVFAMQGFGILGGGIFAIIISAAFKAKFDAPPYEVDPVGS-T 539
Query: 208 VPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQAAADMSKVLQVELEV 267
VPQ DY+WR+I+M GALPAALTYYWRMKMPETARYTALVAKN +QAA DMSKVLQVE++
Sbjct: 540 VPQADYIWRIIVMVGALPAALTYYWRMKMPETARYTALVAKNTEQAAKDMSKVLQVEIQA 719
Query: 268 EEEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCSTWFLLDIAFYSQNLFQKDIFSAIG 327
E K + N++ LFSK+F RHGL L GT STWFLLDIAFYSQNLFQKDIFSAIG
Sbjct: 720 E----PKGDQAQANTFALFSKEFMRRHGLHLLGTASTWFLLDIAFYSQNLFQKDIFSAIG 887
Query: 328 WIPPAKEMNAIHEVYKIARAQTLIALCSTVP 358
WIPPAK MNA+ EVY+IARAQTLIALCS P
Sbjct: 888 WIPPAKTMNALEEVYRIARAQTLIALCSYCP 980
>BP067261
Length = 460
Score = 251 bits (642), Expect = 2e-67
Identities = 121/154 (78%), Positives = 135/154 (87%)
Frame = +1
Query: 269 EEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCSTWFLLDIAFYSQNLFQKDIFSAIGW 328
+EK++K+T N ++S + AL GT STWFLLDIAFYSQNLFQKDIFSAIGW
Sbjct: 1 QEKLQKITEADNNKL-VYSARNCQAPWAALLGTSSTWFLLDIAFYSQNLFQKDIFSAIGW 177
Query: 329 IPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMF 388
IPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVA ID+MGRFAIQ+MGFFFMTVFMF
Sbjct: 178 IPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVALIDYMGRFAIQLMGFFFMTVFMF 357
Query: 389 ALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPN 422
ALAIPYDHW++++NRIGFV +YSLTFFFANFGPN
Sbjct: 358 ALAIPYDHWTEKDNRIGFVAMYSLTFFFANFGPN 459
>AU251730
Length = 336
Score = 129 bits (323), Expect = 2e-30
Identities = 64/86 (74%), Positives = 68/86 (78%)
Frame = +1
Query: 3 GELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPG 62
GELGVL ALD AKTQ YHFT I+I GMGFFTDAYDLF I VTKLLGRIYYT+ +PG
Sbjct: 79 GELGVLTALDTAKTQWYHFTAIIITGMGFFTDAYDLFSIPNVTKLLGRIYYTKEGSPKPG 258
Query: 63 TLPPSAQSAVTGVALVGTLAGQLFFG 88
TLP + AV GVAL GTLAGQLFFG
Sbjct: 259 TLPANVSVAVNGVALCGTLAGQLFFG 336
>BP042583
Length = 459
Score = 109 bits (273), Expect = 1e-24
Identities = 59/97 (60%), Positives = 68/97 (69%)
Frame = -3
Query: 430 AEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIML 489
AEIFP RLRSTCHGISAAAGKAGA+VG+FGFLYA + IG++N LI+L
Sbjct: 457 AEIFPPRLRSTCHGISAAAGKAGAMVGSFGFLYAQNA-------------IGLRNVLILL 317
Query: 490 GVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEAT 526
GV N +G+ T LVPE GKSLEE+SGE E E AT
Sbjct: 316 GVANLLGLFFTFLVPEPNGKSLEEISGEAEEEETAAT 206
>AV769249
Length = 485
Score = 89.7 bits (221), Expect = 1e-18
Identities = 39/60 (65%), Positives = 49/60 (81%)
Frame = -1
Query: 470 TKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQE 529
+K D GYP GIG+KNSL++LGV+N +G CT LVPE+KGKSLEE+SGENE EG T++E
Sbjct: 482 SKADAGYPAGIGVKNSLLLLGVVNILGFFCTFLVPEAKGKSLEEMSGENEEEGENGTKEE 303
>TC13756 similar to UP|Q9ARI9 (Q9ARI9) Phosphate transporter 1, partial
(11%)
Length = 446
Score = 84.0 bits (206), Expect = 6e-17
Identities = 38/58 (65%), Positives = 48/58 (82%)
Frame = +3
Query: 464 AQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGE 521
AQ++D +K D GYP GIG+KNSL++LGV+N +G L T LVPE+KGKSLEE+SGE E E
Sbjct: 6 AQNQDKSKADAGYPAGIGVKNSLLLLGVVNILGFLFTFLVPEAKGKSLEEMSGEQEEE 179
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (93%)
Length = 1865
Score = 57.0 bits (136), Expect = 7e-09
Identities = 118/531 (22%), Positives = 201/531 (37%), Gaps = 35/531 (6%)
Frame = +2
Query: 19 YHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALV 78
Y +++A M YD +S G + + + + + + Q + G+ +
Sbjct: 113 YALACVIVASMVSIISGYDTGVMS------GALLFIKEDIG----ISDTQQEVLAGILNI 262
Query: 79 GTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGF 138
L G L G D +GR+ L IL +V +V G +G + A L F R G
Sbjct: 263 CALVGCLAAGKTSDYIGRRYTIFLASILFLVGAVFMG--YGPN----FAILMFGRCVCGL 424
Query: 139 GIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGG-------GIVALTVASIFDH 191
G+G + +E ++ TRG + G GI G G +ALT+
Sbjct: 425 GVGFALTTAPVYSAELSSASTRGFLTSLPEVCIGLGIFIGYISNYFLGKLALTLG----- 589
Query: 192 KYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKN-- 249
WRL+L A+P+ + MPE+ R+ + +
Sbjct: 590 -----------------------WRLMLGLAAIPSLGLALGILTMPESPRWLVMQGRLGC 700
Query: 250 AKQAAADMSKVLQ-VELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCSTWFLL 308
AK+ ++S + EL + V +K N + Q + HG ++
Sbjct: 701 AKKVLLEVSNTTEEAELRFRDIIVSAGFDEKCNDEFVKQPQ-KSHHGEGVWK-------- 853
Query: 309 DIAFYSQNLFQKDIFSAIG--WIPPAKEMNAIH----EVYKIARAQTLIALCSTVPGYWF 362
++ ++ + +A+G + A + A+ ++K A T L G
Sbjct: 854 ELFLRPTPPVRRMLIAAVGIHFFEHATGIEAVMLYGPRIFKKAGVTTKDRLLLATIGTGL 1033
Query: 363 T--------VAFIDHMGR---FAIQMMGFFF-MTVFMFALAIPYDHWSKEEN----RIGF 406
T +D +GR I + G F +T+ F+L + +S E+ +
Sbjct: 1034TKITFLTISTFLLDRVGRRRLLQISVAGMIFGLTILGFSLTMV--EYSSEKLVWALSLSI 1207
Query: 407 VVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAAA--GKAGAIVGAFGFLYAA 464
V Y+ FF N G T+V +EIFP RLR+ + I A G AI +F +Y A
Sbjct: 1208VATYTYVAFF-NVGLAPVTWVYSSEIFPLRLRAQGNSIGVAVNRGMNAAISMSFISIYKA 1384
Query: 465 QSKDPTKTDKGYPTGIGIKNSLIMLGVINFVG-MLCTLLVPESKGKSLEEL 514
I I + + ++ V + +PE+KGK+LEE+
Sbjct: 1385---------------ITIGGAFFLFAGMSVVAWVFFYFCLPETKGKALEEM 1492
>TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose transporter
HXT4 (Low-affinity glucose transporter LGT1), partial
(5%)
Length = 728
Score = 50.4 bits (119), Expect = 7e-07
Identities = 34/116 (29%), Positives = 57/116 (48%), Gaps = 3/116 (2%)
Frame = +3
Query: 402 NRIGFVVIYSLTFFFANFGPNATT--FVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFG 459
+++GF+ + L + F P T +VV +EI+P R R C GI++ +V +
Sbjct: 102 SKLGFLALGGLALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIASTTVWVSNLVVSQS 281
Query: 460 FLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLL-VPESKGKSLEEL 514
FL Q+ IG + +M G+I +G+ L+ VPE+KG +EE+
Sbjct: 282 FLSLTQT-------------IGTAWTFMMFGIIAIIGIFFVLIFVPETKGVPMEEV 410
>TC10135
Length = 1501
Score = 43.9 bits (102), Expect = 7e-05
Identities = 29/116 (25%), Positives = 54/116 (46%), Gaps = 3/116 (2%)
Frame = +2
Query: 402 NRIGFVVIYSLTFFFANFGPNATT--FVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFG 459
++ G+ + L + F P T +V+ +EI+P R R C GI++ ++ +
Sbjct: 875 SKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPFRYRGVCGGIASTTVWISNLIVSQS 1054
Query: 460 FLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLL-VPESKGKSLEEL 514
FL Q+ IG + +M G++ V + ++ VPE+KG +EE+
Sbjct: 1055FLSLTQT-------------IGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEV 1183
Score = 26.9 bits (58), Expect = 8.2
Identities = 18/74 (24%), Positives = 33/74 (44%), Gaps = 6/74 (8%)
Frame = +2
Query: 215 WRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQAAADMSKVL------QVELEVE 268
WR +L A PA + + +PE+ R+ L K ++ + + K + E+E
Sbjct: 95 WRWMLGVAAAPAIIQIVLMLSLPESPRW--LYRKGREEESKSILKKIYAPEDVDAEIEAL 268
Query: 269 EEKVEKMTSDKRNS 282
+E VE + + S
Sbjct: 269 KESVESEIEESKTS 310
>BP039984
Length = 217
Score = 43.5 bits (101), Expect = 9e-05
Identities = 20/26 (76%), Positives = 22/26 (83%)
Frame = +3
Query: 3 GELGVLNALDVAKTQLYHFTTIVIAG 28
G + VLNALDVAKTQ YHFT I+IAG
Sbjct: 138 GAIHVLNALDVAKTQWYHFTAIIIAG 215
>TC19595
Length = 500
Score = 43.1 bits (100), Expect = 1e-04
Identities = 28/91 (30%), Positives = 45/91 (48%), Gaps = 1/91 (1%)
Frame = +1
Query: 426 FVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNS 485
+ V +EI+P R C G+SA +++ + FL + + +G+ S
Sbjct: 10 WTVNSEIYPEEFRGVCGGMSATVNWVCSVIMSQSFLSISDA-------------VGLGGS 150
Query: 486 LIMLGVINFVGMLCTLL-VPESKGKSLEELS 515
+LGVI V L LL VPE+KG + EE++
Sbjct: 151 FAILGVIAVVAFLFVLLFVPETKGLTFEEMT 243
>TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transporter like
protein {Arabidopsis thaliana;}, partial (25%)
Length = 545
Score = 38.9 bits (89), Expect = 0.002
Identities = 22/73 (30%), Positives = 37/73 (50%)
Frame = +3
Query: 69 QSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMAT 128
Q A+ A+ G + G GW+ D+ GR+ + L ++ SV ++ +P A
Sbjct: 312 QEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVI--MAAAPNP----AI 473
Query: 129 LCFFRFWLGFGIG 141
L F R ++GFG+G
Sbjct: 474 LLFGRVFVGFGVG 512
>BI420004
Length = 547
Score = 38.5 bits (88), Expect = 0.003
Identities = 37/117 (31%), Positives = 53/117 (44%), Gaps = 2/117 (1%)
Frame = +1
Query: 27 AGMGFFTDAYDLFCISLVTKLLGRIYYTEPN--PTRPGTLPPSAQSAVTGVALVGTLAGQ 84
AG+G YD IS G + Y + + R +L Q + +ALVG + G
Sbjct: 190 AGIGGLLFGYDTGVIS------GALLYIKDDFEEVRRSSL---LQETIVSMALVGAIIGA 342
Query: 85 LFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIG 141
GW+ D GRKK TL VV ++ S L ++P + + F R +G GIG
Sbjct: 343 ATGGWINDAFGRKKA---TLSADVVFTLGS-LVMAAAPDAYF--VIFGRLLVGLGIG 495
>BG662351
Length = 407
Score = 38.1 bits (87), Expect = 0.004
Identities = 30/96 (31%), Positives = 46/96 (47%), Gaps = 1/96 (1%)
Frame = +1
Query: 420 GPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTG 479
G A +VV +EIFP ++ AG +V FG + + + + Y T
Sbjct: 130 GMGAVPWVVMSEIFPVNIKGQ-------AGSLATLVNWFGAWLCSYTFNFLMSWSTYGT- 285
Query: 480 IGIKNSLIMLGVINFVGMLC-TLLVPESKGKSLEEL 514
I+ IN +G+L ++VPE+KGKSLE+L
Sbjct: 286 ------FILYAAINALGILSIVVVVPETKGKSLEQL 375
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 37.4 bits (85), Expect = 0.006
Identities = 50/207 (24%), Positives = 77/207 (37%)
Frame = +2
Query: 74 GVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFR 133
G+ + +L G G D +GR+ T++ L G SP L F R
Sbjct: 368 GIINLYSLIGSGLAGRTSDWIGRR----YTIVFAGAIFFVGALLMGFSPNYWF--LMFGR 529
Query: 134 FWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKY 193
F G GIG ++ +E + +RG + GIL G I + +
Sbjct: 530 FIAGIGIGYALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYISNFAFSKL----- 694
Query: 194 KVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQA 253
SL V WR++L GALP+ + + MPE+ R+ + +
Sbjct: 695 -----------SLKVG-----WRMMLGVGALPSVILGVGVLAMPESPRWLVM-----RGR 811
Query: 254 AADMSKVLQVELEVEEEKVEKMTSDKR 280
D KVL + EE ++ KR
Sbjct: 812 LGDAIKVLNKTSDSPEEAQLRLADIKR 892
>TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(21%)
Length = 550
Score = 37.0 bits (84), Expect = 0.008
Identities = 30/111 (27%), Positives = 51/111 (45%), Gaps = 1/111 (0%)
Frame = +2
Query: 407 VVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQS 466
V+ Y TF + G T+V +EIFP RLR+ + + + V + FL ++
Sbjct: 5 VLSYVATF---SIGAGPITWVYSSEIFPLRLRAQGCAMGVVVNRVTSGVISMTFLSLSK- 172
Query: 467 KDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTL-LVPESKGKSLEELSG 516
GI I + + G I G + ++PE++GK+LE++ G
Sbjct: 173 ------------GITIGGAFFLFGGIAICGWIFFYTMLPETRGKTLEDMEG 289
>AV766904
Length = 611
Score = 34.3 bits (77), Expect = 0.052
Identities = 39/155 (25%), Positives = 61/155 (39%)
Frame = +3
Query: 72 VTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCF 131
++G+ + + G G D +GR+ T++L + + G SP A L F
Sbjct: 216 LSGIMNIYSPIGSYIAGRTSDWIGRR----YTIVLAGLIFFVGAILMGFSPN--YAFLMF 377
Query: 132 FRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDH 191
RF G GIG + ++ SE + +RG + GIL G I T +
Sbjct: 378 GRFVAGVGIGFAFLIAPVYTSEISPTLSRGFLTSLPEVFLNAGILIGYISNYTFS----- 542
Query: 192 KYKVPTFEENPAASLLVPQFDYVWRLILMFGALPA 226
K+P WRL+L GA+P+
Sbjct: 543 --KLP--------------LRLGWRLMLGIGAIPS 599
>BP074036
Length = 405
Score = 32.0 bits (71), Expect = 0.26
Identities = 25/101 (24%), Positives = 44/101 (42%), Gaps = 2/101 (1%)
Frame = -2
Query: 416 FANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAG--AIVGAFGFLYAAQSKDPTKTD 473
F + G T++ +EIFP LR+ + A + A++ +F +Y A
Sbjct: 404 FMSIGIGPVTWIYSSEIFPITLRAQGLAVCVAVNRIVNMAMLTSFISIYKA--------- 252
Query: 474 KGYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEEL 514
I + L L +N + +PE+KG+SLE++
Sbjct: 251 ------ITMGGCLFALAGVNVLAFWFYFTLPETKGRSLEDM 147
>BP053600
Length = 400
Score = 32.0 bits (71), Expect = 0.26
Identities = 14/39 (35%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Frame = -2
Query: 480 IGIKNSLIMLGVINFVGMLCTLL-VPESKGKSLEELSGE 517
+G +N ++ G ++ V +L + VPE+KG SLEE+ +
Sbjct: 339 LGAENLFLLFGALSLVALLFVIFSVPETKGLSLEEIESQ 223
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,632,945
Number of Sequences: 28460
Number of extensions: 113925
Number of successful extensions: 919
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 894
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 911
length of query: 533
length of database: 4,897,600
effective HSP length: 95
effective length of query: 438
effective length of database: 2,193,900
effective search space: 960928200
effective search space used: 960928200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145767.6


