

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145753.3 - phase: 0
(383 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
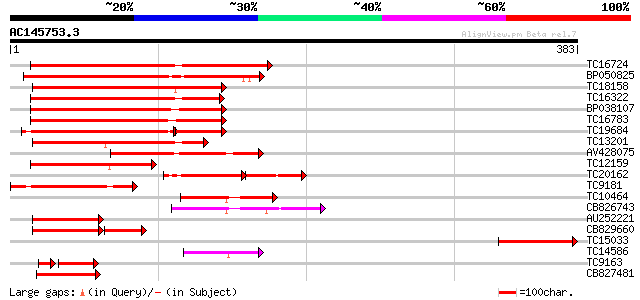
Score E
Sequences producing significant alignments: (bits) Value
TC16724 similar to GB|BAB20599.1|12060424|AB049070 AtNAC3 {Arabi... 184 2e-47
BP050825 173 5e-44
TC18158 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (34%) 170 3e-43
TC16322 homologue to UP|Q9SQL0 (Q9SQL0) Jasmonic acid 2, partial... 166 8e-42
BP038107 165 1e-41
TC16783 similar to UP|AAR88435 (AAR88435) NAC domain protein, pa... 160 3e-40
TC19684 similar to UP|AAR88435 (AAR88435) NAC domain protein, pa... 115 1e-35
TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partia... 121 2e-28
AV428075 101 2e-22
TC12159 similar to UP|Q9M4U3 (Q9M4U3) NAC1, partial (21%) 91 2e-19
TC20162 similar to UP|Q8LKN6 (Q8LKN6) Nam-like protein 18, parti... 89 2e-18
TC9181 similar to UP|Q9FY93 (Q9FY93) NAM-like protein (AT5g13180... 81 3e-16
TC10464 weakly similar to UP|Q8L8G0 (Q8L8G0) Nam-like protein 1,... 62 1e-10
CB826743 61 3e-10
AU252221 60 6e-10
CB829660 59 1e-09
TC15033 55 1e-08
TC14586 similar to UP|Q9FY93 (Q9FY93) NAM-like protein (AT5g1318... 55 1e-08
TC9163 similar to UP|Q9AT45 (Q9AT45) ADP-glucose pyrophosphoryla... 40 1e-05
CB827481 43 1e-04
>TC16724 similar to GB|BAB20599.1|12060424|AB049070 AtNAC3 {Arabidopsis
thaliana;}, partial (53%)
Length = 615
Score = 184 bits (468), Expect = 2e-47
Identities = 90/163 (55%), Positives = 106/163 (64%)
Frame = +2
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L LPPGFRF PTD E++V YL KV FS I E DL K +PW LP KA GEKEWY
Sbjct: 128 LSLPPGFRFFPTDEELLVQYLCRKVAGYHFSLPIIAEIDLYKFDPWVLPGKAIFGEKEWY 307
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NR SGYWKATG DK I +G+ VG+KK LVFY G+APKG
Sbjct: 308 FFSPRDRKYPNGSRPNRVAGSGYWKATGTDKVITTEGR----KVGIKKALVFYIGKAPKG 475
Query: 135 EKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKK 177
KTNW+MHE+RL H L + DEWV+ R++ K + +K
Sbjct: 476 TKTNWIMHEYRLLDSSRKHKLGSSRLDEWVLCRIYKKKSSTQK 604
>BP050825
Length = 539
Score = 173 bits (438), Expect = 5e-44
Identities = 86/172 (50%), Positives = 108/172 (62%), Gaps = 9/172 (5%)
Frame = +2
Query: 10 GEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG 69
G Q +LPPGFRFHPTD E++V YL +K + I E DL K +PW+LP KA G
Sbjct: 17 GSQQQPNLPPGFRFHPTDEELVVHYLKKKAASVPLPVAIIAEVDLYKFDPWELPAKATFG 196
Query: 70 EKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKG 129
E+EWYFF +DRKYP G R NRA SGYWKATG DK + G Q VG+KK LVFY G
Sbjct: 197 EQEWYFFSPRDRKYPNGARPNRAATSGYWKATGTDKPVL-TSNGTQK-VGVKKALVFYGG 370
Query: 130 RAPKGEKTNWVMHEFRLEGKFATHNLP-----NKEK----DEWVVSRVFHKN 172
+ P+G KTNW+MHE+RL + P NK+ D+WV+ R++ K+
Sbjct: 371 KPPRGIKTNWIMHEYRLADNKPNNRPPGCDLGNKKNSLRLDDWVLCRIYKKS 526
>TC18158 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (34%)
Length = 476
Score = 170 bits (431), Expect = 3e-43
Identities = 79/134 (58%), Positives = 95/134 (69%), Gaps = 3/134 (2%)
Frame = +3
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYF 75
+LPPGFRFHPTD E+I+ YL +KV + + I E D+ K +PW+LP KA GEKEWYF
Sbjct: 75 NLPPGFRFHPTDEELILHYLRKKVASIPLPVSIIAEVDIYKLDPWELPGKALFGEKEWYF 254
Query: 76 FCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHK---GKGIQNLVGMKKTLVFYKGRAP 132
F +DRKYP G R NRA SGYWKATG DK I G Q+ +G+KK LVFYKG+ P
Sbjct: 255 FSPRDRKYPNGARPNRAAASGYWKATGTDKTIVASLPGGGRAQDSIGVKKALVFYKGKPP 434
Query: 133 KGEKTNWVMHEFRL 146
KG KTNW+MHE+RL
Sbjct: 435 KGVKTNWIMHEYRL 476
>TC16322 homologue to UP|Q9SQL0 (Q9SQL0) Jasmonic acid 2, partial (40%)
Length = 510
Score = 166 bits (419), Expect = 8e-42
Identities = 79/131 (60%), Positives = 91/131 (69%)
Frame = +1
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L LPPGFRF+PTD E++V YL KV F+ I E DL K +PW LP KA GEKEWY
Sbjct: 130 LSLPPGFRFYPTDEELLVQYLCRKVAGHNFTLPIIAEIDLYKFDPWVLPSKAIFGEKEWY 309
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NR SGYWKATG DK I +G+ VG+KK LVFY G+APKG
Sbjct: 310 FFSPRDRKYPNGSRPNRVAGSGYWKATGTDKIITTEGR----KVGIKKALVFYVGKAPKG 477
Query: 135 EKTNWVMHEFR 145
KTNW+MHE+R
Sbjct: 478 TKTNWIMHEYR 510
>BP038107
Length = 469
Score = 165 bits (418), Expect = 1e-41
Identities = 77/132 (58%), Positives = 90/132 (67%)
Frame = +3
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L PPGFRFHPTD E+++ YL K + + I E DL K +PWDLP A GEKEWY
Sbjct: 57 LQWPPGFRFHPTDEELVLHYLCRKCASQPIAVPIIAEIDLYKYDPWDLPGLASYGEKEWY 236
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NRA +GYWKATG DK I H VG+KK LVFY G+APKG
Sbjct: 237 FFSPRDRKYPNGSRPNRAAGTGYWKATGADKPIGH-----PKPVGIKKALVFYAGKAPKG 401
Query: 135 EKTNWVMHEFRL 146
+KTNW+MHE+RL
Sbjct: 402 DKTNWIMHEYRL 437
>TC16783 similar to UP|AAR88435 (AAR88435) NAC domain protein, partial (46%)
Length = 511
Score = 160 bits (406), Expect = 3e-40
Identities = 76/132 (57%), Positives = 89/132 (66%)
Frame = +1
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L+LPPGFRFHPTD E++ YL K + + I E DL K +PW LP A GEKEWY
Sbjct: 91 LELPPGFRFHPTDDELVKHYLCAKCASQPINVPIIKELDLYKFDPWQLPDMALYGEKEWY 270
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NRA +GYWKATG DK I G +G+KK LVFY G+APKG
Sbjct: 271 FFTPRDRKYPNGSRPNRAAGTGYWKATGADKPI-----GKPKTLGIKKALVFYAGKAPKG 435
Query: 135 EKTNWVMHEFRL 146
KTNW+MHE+RL
Sbjct: 436 VKTNWIMHEYRL 471
>TC19684 similar to UP|AAR88435 (AAR88435) NAC domain protein, partial (48%)
Length = 535
Score = 115 bits (288), Expect(2) = 1e-35
Identities = 59/106 (55%), Positives = 69/106 (64%)
Frame = +2
Query: 9 KGEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKM 68
KGE L+LPPGFRFHPTD E++ YL K F+ + I E DL K +PW LP+
Sbjct: 95 KGE---LELPPGFRFHPTDEELVNHYLCTKGAGQSFNYSVIKEIDLYKFDPWQLPEMGLD 265
Query: 69 GEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGI 114
GEKEWYFF +DRKYP G R NRA SGY KATG DK + K KG+
Sbjct: 266 GEKEWYFFSPRDRKYPNGSRPNRAAGSGYGKATGADKPM-GKPKGV 400
Score = 50.8 bits (120), Expect(2) = 1e-35
Identities = 21/34 (61%), Positives = 26/34 (75%)
Frame = +3
Query: 113 GIQNLVGMKKTLVFYKGRAPKGEKTNWVMHEFRL 146
G +G+KK LV Y G+APKG KTNW+MHE+RL
Sbjct: 384 GSPRALGIKKALVLYVGKAPKGIKTNWIMHEYRL 485
>TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partial (22%)
Length = 465
Score = 121 bits (304), Expect = 2e-28
Identities = 63/122 (51%), Positives = 76/122 (61%), Gaps = 3/122 (2%)
Frame = +3
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLP--KKAKMGEKEW 73
+L PGFRFHPTD E++ YL +V F IG D+ K EPWDLP K K + EW
Sbjct: 108 NLSPGFRFHPTDEELVGFYLKRRVTGRLFHGDPIGIVDVYKYEPWDLPCLSKMKTRDLEW 287
Query: 74 YFFCQKDRKYPTG-MRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAP 132
YF+ D+KY G RTNRATE GYWK TGKD+ + H + VGMKKTLV++ GRA
Sbjct: 288 YFYSGLDKKYGKGSSRTNRATEKGYWKTTGKDRPVNHANR----TVGMKKTLVYHFGRAS 455
Query: 133 KG 134
G
Sbjct: 456 TG 461
>AV428075
Length = 310
Score = 101 bits (251), Expect = 2e-22
Identities = 50/103 (48%), Positives = 66/103 (63%)
Frame = +2
Query: 69 GEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYK 128
GEKEWYF+ +DRKY G R NR T SGYWKATG D+ I + +G+KKTLVFY
Sbjct: 2 GEKEWYFYVPRDRKYRNGDRPNRVTTSGYWKATGADRMIRTEN---FRSIGLKKTLVFYS 172
Query: 129 GRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHK 171
G+APKG +T+W+M+E+RL H +K E + RV+ +
Sbjct: 173 GKAPKGMRTSWIMNEYRL----PQHETERYQKAEISLCRVYKR 289
>TC12159 similar to UP|Q9M4U3 (Q9M4U3) NAC1, partial (21%)
Length = 630
Score = 91.3 bits (225), Expect = 2e-19
Identities = 42/87 (48%), Positives = 54/87 (61%), Gaps = 2/87 (2%)
Frame = +1
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKA--KMGEKE 72
+ L PGFRFHP D E++ YL KV K I E ++ K PWDLP K+ + G+ E
Sbjct: 370 MKLVPGFRFHPMDVELVKFYLKRKVMGRKLPHDVIAEVNVYKHAPWDLPDKSCLRTGDLE 549
Query: 73 WYFFCQKDRKYPTGMRTNRATESGYWK 99
WYFF ++KY +G R NRATE G+WK
Sbjct: 550 WYFFSPTEKKYGSGARMNRATEIGFWK 630
>TC20162 similar to UP|Q8LKN6 (Q8LKN6) Nam-like protein 18, partial (17%)
Length = 623
Score = 88.6 bits (218), Expect = 2e-18
Identities = 44/56 (78%), Positives = 50/56 (88%)
Frame = +2
Query: 105 KEIYHKGKGIQNLVGMKKTLVFYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEK 160
+EI+ KGKG NLVGMKKTLVFY+GRAPKGEKTNWVMHEFRLEGKFA+++LP K
Sbjct: 23 REIF-KGKG--NLVGMKKTLVFYRGRAPKGEKTNWVMHEFRLEGKFASYSLPKIAK 181
Score = 58.5 bits (140), Expect = 2e-09
Identities = 32/42 (76%), Positives = 35/42 (83%), Gaps = 1/42 (2%)
Frame = +2
Query: 160 KDEWVVSRVFHKNTDVKKPQISSGLLRINSIG-HDDLLDYSS 200
+DEWVVSRVFHK+TDVKK I GLLR+NSIG DLLDYSS
Sbjct: 500 QDEWVVSRVFHKSTDVKKSPI-PGLLRMNSIGDQHDLLDYSS 622
>TC9181 similar to UP|Q9FY93 (Q9FY93) NAM-like protein
(AT5g13180/T19L5_140), partial (30%)
Length = 519
Score = 81.3 bits (199), Expect = 3e-16
Identities = 43/86 (50%), Positives = 53/86 (61%)
Frame = +1
Query: 1 MEEPIVVNKGEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPW 60
ME+ V GE L LPPGFRFHPTD E++V YL K+ + A+ I E DL K +PW
Sbjct: 277 MEKVNFVKNGE---LRLPPGFRFHPTDEELVVQYLKRKIFSCPLPASIIPEVDLCKSDPW 447
Query: 61 DLPKKAKMGEKEWYFFCQKDRKYPTG 86
DLP + E+E YFF K+ KYP G
Sbjct: 448 DLPGDS---ERERYFFSMKEAKYPNG 516
>TC10464 weakly similar to UP|Q8L8G0 (Q8L8G0) Nam-like protein 1, partial
(12%)
Length = 935
Score = 62.4 bits (150), Expect = 1e-10
Identities = 33/71 (46%), Positives = 46/71 (64%), Gaps = 5/71 (7%)
Frame = +1
Query: 116 NLVGMKKTLVFYKGRAPKGEKTNWVMHEFR-----LEGKFATHNLPNKEKDEWVVSRVFH 170
+L+GMKKTLVFY GRAPKG++TNWVMHE+R L+G N ++ +V+ R+F
Sbjct: 10 SLIGMKKTLVFYHGRAPKGKRTNWVMHEYRPTLQELDG-------TNPGQNPYVICRLFK 168
Query: 171 KNTDVKKPQIS 181
K + + IS
Sbjct: 169 KQDESIEVSIS 201
>CB826743
Length = 513
Score = 61.2 bits (147), Expect = 3e-10
Identities = 43/115 (37%), Positives = 62/115 (53%), Gaps = 11/115 (9%)
Frame = +2
Query: 110 KGKGIQNLVGMKKTLVFYKGRAPKGEKTNWVMHEFR-----LEGKFATHNLPNKEKDEWV 164
K K L+GMKKTLVFY GRAPKG++T+WVMHE+R L+G N ++ +V
Sbjct: 8 KIKSGSTLIGMKKTLVFYAGRAPKGKRTHWVMHEYRPTLKELDG-------TNP*QNPYV 166
Query: 165 VSRVFHKN------TDVKKPQISSGLLRINSIGHDDLLDYSSLPPLMDPSYTNDD 213
+ R+F KN ++ + QI+S S +++ SL P+ T DD
Sbjct: 167 LCRLFKKNDESLEGSNEEVEQIAS-TPTTASFSPEEIQSDPSLVPVSSSLATEDD 328
>AU252221
Length = 424
Score = 60.1 bits (144), Expect = 6e-10
Identities = 25/48 (52%), Positives = 34/48 (70%)
Frame = +1
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLP 63
+LPPGFRFHPTD E+I+ YL +KV + + I E D+ K +PW+LP
Sbjct: 103 NLPPGFRFHPTDEELILHYLRKKVASIPLPVSIIAEVDIYKLDPWELP 246
>CB829660
Length = 517
Score = 59.3 bits (142), Expect = 1e-09
Identities = 26/48 (54%), Positives = 33/48 (68%)
Frame = +1
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLP 63
+LPPGFRFHPTD E+IV YL +V + A I E D+ K +PW+LP
Sbjct: 127 ELPPGFRFHPTDEELIVHYLCNQVSSHPSPAPIIPEVDIYKFDPWELP 270
Score = 45.1 bits (105), Expect = 2e-05
Identities = 18/28 (64%), Positives = 20/28 (71%)
Frame = +3
Query: 65 KAKMGEKEWYFFCQKDRKYPTGMRTNRA 92
K GEKEWYFF ++RKYP G R NRA
Sbjct: 432 KTAFGEKEWYFFSPRERKYPNGARPNRA 515
>TC15033
Length = 676
Score = 55.5 bits (132), Expect = 1e-08
Identities = 34/57 (59%), Positives = 41/57 (71%), Gaps = 4/57 (7%)
Frame = +2
Query: 331 QFSSNQSQDTGLSND-TSSAVSKLDMERNRA-LYDDL-EGPSSVA-PLSDLDSFWDY 383
QFSSN SQDTGLSND ++ S + RNRA LY+DL +GP+SVA SDL+ WDY
Sbjct: 2 QFSSNHSQDTGLSNDRNTTETSSVVSGRNRASLYEDLDQGPASVARAFSDLECLWDY 172
>TC14586 similar to UP|Q9FY93 (Q9FY93) NAM-like protein
(AT5g13180/T19L5_140), partial (18%)
Length = 589
Score = 55.5 bits (132), Expect = 1e-08
Identities = 26/57 (45%), Positives = 34/57 (59%), Gaps = 3/57 (5%)
Frame = +3
Query: 118 VGMKKTLVFYKGRAPKGEKTNWVMHEFRL---EGKFATHNLPNKEKDEWVVSRVFHK 171
VGMKKTLVFY G+ P G +T+W+MHE+RL T + WV+ R+F K
Sbjct: 3 VGMKKTLVFYTGKPPHGSRTDWIMHEYRLLTNTNTTTTQGHVQVPMENWVLCRIFLK 173
>TC9163 similar to UP|Q9AT45 (Q9AT45) ADP-glucose pyrophosphorylase large
subunit (Glucose-1-phosphate adenylyltransferase)
(ADP-glucose synthase) , partial (5%)
Length = 663
Score = 39.7 bits (91), Expect(2) = 1e-05
Identities = 17/27 (62%), Positives = 21/27 (76%)
Frame = +2
Query: 34 YLTEKVKNSKFSATAIGEADLNKCEPW 60
YL +KV +S F A AI +ADL+KCEPW
Sbjct: 257 YLPQKVLDSYFYAIAIADADLHKCEPW 337
Score = 25.4 bits (54), Expect(2) = 1e-05
Identities = 9/12 (75%), Positives = 11/12 (91%)
Frame = +3
Query: 20 GFRFHPTDAEII 31
GFRFHPT+ E+I
Sbjct: 216 GFRFHPTNEELI 251
>CB827481
Length = 556
Score = 42.7 bits (99), Expect = 1e-04
Identities = 20/44 (45%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Frame = +2
Query: 19 PGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEA-DLNKCEPWD 61
PGFRFHPTD E++ YL K++ I E+ DL K +PW+
Sbjct: 224 PGFRFHPTDEELVSFYLRRKLEKKSHVFELIKESIDLYKYDPWN 355
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.132 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,707,415
Number of Sequences: 28460
Number of extensions: 93503
Number of successful extensions: 366
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 353
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 354
length of query: 383
length of database: 4,897,600
effective HSP length: 92
effective length of query: 291
effective length of database: 2,279,280
effective search space: 663270480
effective search space used: 663270480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC145753.3


