

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145513.12 + phase: 0
(304 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
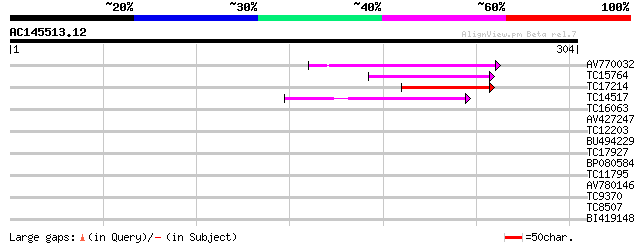
Score E
Sequences producing significant alignments: (bits) Value
AV770032 87 3e-18
TC15764 weakly similar to GB|AAL31246.1|16974485|AY061919 AT4g32... 62 9e-11
TC17214 54 4e-08
TC14517 52 2e-07
TC16063 39 0.001
AV427247 38 0.002
TC12203 similar to UP|Q9SXX9 (Q9SXX9) FAS1 (Fragment), partial (... 30 0.66
BU494229 28 1.5
TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete 28 1.9
BP080584 28 2.5
TC11795 27 5.6
AV780146 27 5.6
TC9370 26 7.3
TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 26 7.3
BI419148 26 9.5
>AV770032
Length = 559
Score = 87.4 bits (215), Expect = 3e-18
Identities = 39/103 (37%), Positives = 61/103 (58%)
Frame = -2
Query: 161 GNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELK 220
G H F+DV+ + S R +EL+F +FE+A+ + Y+ ++ +P +FVG E L
Sbjct: 558 GEHSFLDVID-NTNSKKGVVRVMIELNFRAEFEMAKGSEDYNRLVRKLPEVFVGKVERLS 382
Query: 221 RTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTN 263
+ LC A K C++ + + + PWRK+RYMQ KW GP R T+
Sbjct: 381 NVIKILCMAAKRCMKEKKMHLGPWRKHRYMQAKWLGPCERNTS 253
>TC15764 weakly similar to GB|AAL31246.1|16974485|AY061919
AT4g32480/F8B4_180 {Arabidopsis thaliana;}, partial
(24%)
Length = 608
Score = 62.4 bits (150), Expect = 9e-11
Identities = 26/68 (38%), Positives = 40/68 (58%)
Frame = +1
Query: 193 EIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQN 252
E+AR + Y+ ++ +P ++VG E L + LC A K C + + + PWRK RYM+
Sbjct: 1 ELARASEGYNRLVWRLPEVYVGKVERLSNVIKILCMAAKRCTKENKMHMGPWRKLRYMEA 180
Query: 253 KWFGPYRR 260
KW GP +R
Sbjct: 181 KWLGPCKR 204
>TC17214
Length = 556
Score = 53.5 bits (127), Expect = 4e-08
Identities = 22/50 (44%), Positives = 31/50 (62%)
Frame = +2
Query: 211 IFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRR 260
IFVG ++L+ V + A K LR +G+ +PPWR+ Y+Q KW PY R
Sbjct: 2 IFVGKCDQLQSIVALVSEAAKQSLRKKGMHVPPWRRVEYVQAKWLSPYSR 151
>TC14517
Length = 697
Score = 51.6 bits (122), Expect = 2e-07
Identities = 32/100 (32%), Positives = 50/100 (50%)
Frame = +1
Query: 148 CKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSY 207
CK + + LT ++E+IDV +G Y + A +F+I+ PT Y+ ++
Sbjct: 100 CKYS*EKNARLTVRDYEYIDVNFSGNG-------YIIVCSLATEFKISHPTYHYTFLVEI 258
Query: 208 VPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKN 247
+P IFV EELKR V LC A+K + L + R +
Sbjct: 259 LPLIFVC*MEELKRVVRLLCSALKGSMERMELQLSEMRSS 378
>TC16063
Length = 585
Score = 38.9 bits (89), Expect = 0.001
Identities = 37/158 (23%), Positives = 62/158 (38%), Gaps = 3/158 (1%)
Frame = +3
Query: 13 FNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTG-- 70
+ DE V + L D+ S + P +L DDG ++T
Sbjct: 135 YADEQNTSFVDVEFEFLDDIGEISLGKRASSDEFGPIEMDLDEDHERVDDGRTEENTSFW 314
Query: 71 -NDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSV 129
N F + + S SVE +R + + D Y + + E + +
Sbjct: 315 DNQFQLLQTNLCRTS--SVESRIRNATKEVVHDIYSSGI-----ECGCSRELAASYCRNC 473
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFID 167
R V L++ G N+AIC+T+W +S + +G H F+D
Sbjct: 474 LMREVSRRLQKAGFNSAICQTKWRNS-SVPSGEHTFLD 584
>AV427247
Length = 225
Score = 37.7 bits (86), Expect = 0.002
Identities = 18/45 (40%), Positives = 25/45 (55%)
Frame = +1
Query: 239 LSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCR 283
+ +PPWR Y+Q KW P+ R T+ GN + D V F +CR
Sbjct: 10 MPLPPWRSLAYLQAKWQSPFERYTHS-EGNNISD-VDCFDHKQCR 138
>TC12203 similar to UP|Q9SXX9 (Q9SXX9) FAS1 (Fragment), partial (24%)
Length = 343
Score = 29.6 bits (65), Expect = 0.66
Identities = 21/65 (32%), Positives = 26/65 (39%), Gaps = 4/65 (6%)
Frame = +2
Query: 29 LSDVSSGSEHSGTGECDSSPSLSE----LVHGFLEEDDGNVCDSTGNDFDSERVDSVSDS 84
LSD E EC S SE + G+L ED+G D D D E +S
Sbjct: 20 LSDCDKDEEECQE-ECSKSDGESEDGFFVPDGYLSEDEGEQVDRMETDIDVEGANSSPSC 196
Query: 85 MDSVE 89
D +E
Sbjct: 197KDDIE 211
>BU494229
Length = 412
Score = 28.5 bits (62), Expect = 1.5
Identities = 15/41 (36%), Positives = 19/41 (45%)
Frame = +1
Query: 64 NVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSY 104
+V D N+ S R+ S D V+ LLRL E D Y
Sbjct: 19 DVIDDHRNEMSSSRLREAEGSEDLVDVLLRLQKEQQEPDQY 141
>TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete
Length = 1793
Score = 28.1 bits (61), Expect = 1.9
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +1
Query: 149 KTRWDSSGGLTAGNHEFIDVVRMRSGS 175
+ + D GG+ G+HE ++VR+R GS
Sbjct: 445 REKMDQHGGVATGSHERNELVRVRHGS 525
>BP080584
Length = 507
Score = 27.7 bits (60), Expect = 2.5
Identities = 12/42 (28%), Positives = 18/42 (42%)
Frame = -1
Query: 244 WRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLV 285
W + + N +F Y R+ +P HG F KC L+
Sbjct: 471 WEIWKCLWNSFFWAYSRSISPSHGTTPSSRSEGFQDTKCELL 346
>TC11795
Length = 577
Score = 26.6 bits (57), Expect = 5.6
Identities = 15/61 (24%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Frame = -2
Query: 240 SIPPWRKNRYMQNKWFGPYRRTTNPVHGNPV-PDVVSSFSGVKCRLVGFDNAMSEIKHGV 298
++ P + R ++ + G + ++T P+ V P + +S L+GF +S++ +GV
Sbjct: 387 TLVPSIRTRAIEKESIGSHSQST*PIMV*KVAPQFILRYSATTSLLLGFSPPLSDLSNGV 208
Query: 299 A 299
A
Sbjct: 207 A 205
>AV780146
Length = 519
Score = 26.6 bits (57), Expect = 5.6
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = -2
Query: 161 GNHEFIDVVRMRSGSSTWQNRYFVELDFAVQ 191
G EFI+V RS S W N E FA++
Sbjct: 407 GQQEFIEVKATRSPSKDWFNITMREWQFAIE 315
>TC9370
Length = 632
Score = 26.2 bits (56), Expect = 7.3
Identities = 9/24 (37%), Positives = 16/24 (66%)
Frame = +3
Query: 177 TWQNRYFVELDFAVQFEIARPTSR 200
+W+NR++V +++A F R SR
Sbjct: 78 SWRNRFYVAINWATTFVFGRDISR 149
>TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (40%)
Length = 620
Score = 26.2 bits (56), Expect = 7.3
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +3
Query: 59 EEDDGNVCDSTGNDFDSERVDSVSDSMDSVED 90
EEDD + D +D D E D+ SDS +S D
Sbjct: 6 EEDDSDDEDDEDDDSDEEMDDADSDSDESDSD 101
>BI419148
Length = 537
Score = 25.8 bits (55), Expect = 9.5
Identities = 11/28 (39%), Positives = 12/28 (42%)
Frame = -2
Query: 233 CLRSRGLSIPPWRKNRYMQNKWFGPYRR 260
C R R PWR Y KW +RR
Sbjct: 329 CCRPRSGGSHPWRAATYRHGKWRERWRR 246
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,829,166
Number of Sequences: 28460
Number of extensions: 64159
Number of successful extensions: 352
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 351
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 352
length of query: 304
length of database: 4,897,600
effective HSP length: 90
effective length of query: 214
effective length of database: 2,336,200
effective search space: 499946800
effective search space used: 499946800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC145513.12


