

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
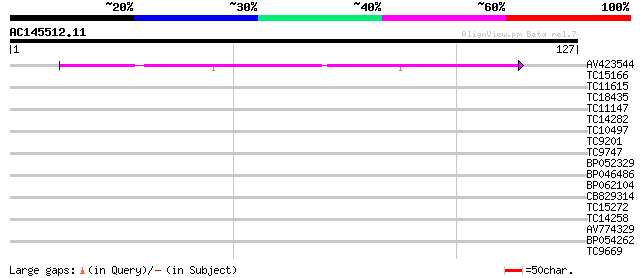
Query= AC145512.11 - phase: 0
(127 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV423544 63 1e-11
TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-bind... 28 0.36
TC11615 28 0.62
TC18435 27 1.1
TC11147 similar to UP|Q9ZSQ7 (Q9ZSQ7) Protein phosphatase 2C hom... 26 2.4
TC14282 25 3.1
TC10497 25 4.0
TC9201 25 4.0
TC9747 similar to GB|AAO63430.1|28951013|BT005366 At3g56900 {Ara... 25 5.2
BP052329 24 6.8
BP046486 24 6.8
BP062104 24 6.8
CB829314 24 6.8
TC15272 weakly similar to UP|Q9FUJ0 (Q9FUJ0) Cytokinin oxidase, ... 24 6.8
TC14258 similar to UP|Q945I3 (Q945I3) Prunasin hydrolase isoform... 24 8.9
AV774329 24 8.9
BP054262 24 8.9
TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%) 24 8.9
>AV423544
Length = 443
Score = 63.2 bits (152), Expect = 1e-11
Identities = 44/108 (40%), Positives = 62/108 (56%), Gaps = 4/108 (3%)
Frame = +1
Query: 12 PSIALGVVSAYTWGYLEHPARVIINDNTFFYTH-GRKIDRCSRPDTYHLHLFHMQVEYFN 70
PSIAL +VS L IIN + + H + R RPD +L+ +Q F
Sbjct: 4 PSIALFLVSKSMESSLG--LNFIINGHEYALAHIFHPVYRTIRPDHAYLYDLQLQGRKFK 177
Query: 71 GNMDKALLENKWNHAE---VDFGFPFMFSGIHVLKEKSNMKDIRFTNP 115
++D+ALLEN+WNHAE V F F+ +GIH+ K+KS+++ IRFT P
Sbjct: 178 -DIDQALLENEWNHAEIILVSFLSTFIGTGIHIFKQKSSIEHIRFTCP 318
>TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-binding
protein 6 {Arabidopsis thaliana;} , partial (26%)
Length = 668
Score = 28.5 bits (62), Expect = 0.36
Identities = 16/54 (29%), Positives = 19/54 (34%)
Frame = +1
Query: 29 HPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKW 82
H R + N H K R S PD HLH H + N + KW
Sbjct: 37 HTTRTLCNSQDRNILHRSKYPRISCPDQPHLHKLHRFIISPNSQVFNCHPSRKW 198
>TC11615
Length = 522
Score = 27.7 bits (60), Expect = 0.62
Identities = 17/60 (28%), Positives = 24/60 (39%), Gaps = 21/60 (35%)
Frame = +2
Query: 76 ALLENKWNHAE---------------------VDFGFPFMFSGIHVLKEKSNMKDIRFTN 114
A ENKWNH E V G ++ + + KE +N +DI+F N
Sbjct: 29 AFSENKWNHVEILCEAKYPRPSSELAMATETWVGTGLSLSWTLVGIYKEGNNKEDIKFEN 208
>TC18435
Length = 505
Score = 26.9 bits (58), Expect = 1.1
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = -1
Query: 56 TYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSG 97
TYH HLF + + F + + + W G+PF+ SG
Sbjct: 274 TYHFHLFSLGISLFFSHHITYHISHFWR------GYPFLTSG 167
>TC11147 similar to UP|Q9ZSQ7 (Q9ZSQ7) Protein phosphatase 2C homolog,
partial (7%)
Length = 583
Score = 25.8 bits (55), Expect = 2.4
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = +1
Query: 44 HGRKIDRCSRPDTYHLHLFH 63
H +++ CS+ +HLH FH
Sbjct: 130 HNKRVSICSKLFIFHLHQFH 189
>TC14282
Length = 647
Score = 25.4 bits (54), Expect = 3.1
Identities = 10/34 (29%), Positives = 15/34 (43%)
Frame = -2
Query: 11 FPSIALGVVSAYTWGYLEHPARVIINDNTFFYTH 44
FP + G WG H ++ + +FFY H
Sbjct: 274 FPFVDAGEQRRRRWGRSSH*PQLAVQRTSFFYAH 173
>TC10497
Length = 542
Score = 25.0 bits (53), Expect = 4.0
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = -2
Query: 45 GRKIDRCSRPDTYHLHLFHMQVEYFN 70
G +D+ S T +L+L+H+ ++Y N
Sbjct: 529 GLILDKSSNHGTDNLYLYHLSLQYLN 452
>TC9201
Length = 1171
Score = 25.0 bits (53), Expect = 4.0
Identities = 10/22 (45%), Positives = 18/22 (81%)
Frame = -1
Query: 104 KSNMKDIRFTNPENDANIVLHS 125
KS+M+D++ TN ++AN++L S
Sbjct: 1132 KSHMRDLQLTNCVSNANVLLVS 1067
>TC9747 similar to GB|AAO63430.1|28951013|BT005366 At3g56900 {Arabidopsis
thaliana;}, partial (22%)
Length = 605
Score = 24.6 bits (52), Expect = 5.2
Identities = 14/35 (40%), Positives = 22/35 (62%)
Frame = -1
Query: 32 RVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQV 66
R+ I++ +F Y H R+ D C T+HLHL +Q+
Sbjct: 146 RM*ISEESFLY-HIRRSDPC----TFHLHL*KIQL 57
>BP052329
Length = 441
Score = 24.3 bits (51), Expect = 6.8
Identities = 11/39 (28%), Positives = 18/39 (45%)
Frame = +2
Query: 44 HGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKW 82
H ++ RC + + H +Y GN+ K +L N W
Sbjct: 194 H*VRLSRCVLEHMLPMIILH--ADYLQGNLRKMMLSNVW 304
>BP046486
Length = 452
Score = 24.3 bits (51), Expect = 6.8
Identities = 10/32 (31%), Positives = 15/32 (46%)
Frame = +1
Query: 52 SRPDTYHLHLFHMQVEYFNGNMDKALLENKWN 83
SR H H H+ Y+N N +L+ + N
Sbjct: 292 SRHQCCHQHRVHIHTHYYNHNHQTQMLQQEPN 387
>BP062104
Length = 321
Score = 24.3 bits (51), Expect = 6.8
Identities = 9/29 (31%), Positives = 13/29 (44%)
Frame = +3
Query: 40 FFYTHGRKIDRCSRPDTYHLHLFHMQVEY 68
FF H I C +H H FH+ + +
Sbjct: 213 FFKVHRLHIPFCHSKQQWHAHAFHIWLHF 299
>CB829314
Length = 567
Score = 24.3 bits (51), Expect = 6.8
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = +2
Query: 26 YLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHM 64
+L P ++ + T HG RC+RP+T LH H+
Sbjct: 80 HL*QPLMLLAHSPTHQEDHGDA--RCARPETAPLHRQHL 190
>TC15272 weakly similar to UP|Q9FUJ0 (Q9FUJ0) Cytokinin oxidase, partial
(14%)
Length = 524
Score = 24.3 bits (51), Expect = 6.8
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 77 LLENKWNHAEVDFGFPFMFS 96
LLEN W+ + FG F FS
Sbjct: 341 LLENPWSR*QDSFGIKFSFS 282
>TC14258 similar to UP|Q945I3 (Q945I3) Prunasin hydrolase isoform PH A
(Fragment) , partial (15%)
Length = 602
Score = 23.9 bits (50), Expect = 8.9
Identities = 18/49 (36%), Positives = 26/49 (52%), Gaps = 7/49 (14%)
Frame = +2
Query: 53 RPDTYHLHLFHMQVEYFNGNMDK-----ALLEN-KWNHA-EVDFGFPFM 94
R D Y HLF++Q NG+ K +LL+N +W+ V FG F+
Sbjct: 11 RIDYYFRHLFYLQSAIRNGSNVKGYFAWSLLDNYEWSSGYTVRFGMNFV 157
>AV774329
Length = 443
Score = 23.9 bits (50), Expect = 8.9
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +3
Query: 81 KWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPENDAN 120
K+ ++ V F F+F +L+ S+ +NPEN+ N
Sbjct: 213 KYKNSNVLIIFIFIFFPFILLQSSSSSSSNSSSNPENEKN 332
>BP054262
Length = 579
Score = 23.9 bits (50), Expect = 8.9
Identities = 11/25 (44%), Positives = 13/25 (52%)
Frame = +3
Query: 79 ENKWNHAEVDFGFPFMFSGIHVLKE 103
E++WN D FMF IH L E
Sbjct: 105 ESRWNIQHADNFLVFMFFKIHQLTE 179
>TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%)
Length = 472
Score = 23.9 bits (50), Expect = 8.9
Identities = 8/16 (50%), Positives = 14/16 (87%)
Frame = -2
Query: 58 HLHLFHMQVEYFNGNM 73
+LHL H++VE F+G++
Sbjct: 321 NLHLLHVRVECFHGHL 274
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,875,751
Number of Sequences: 28460
Number of extensions: 42423
Number of successful extensions: 241
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 240
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 240
length of query: 127
length of database: 4,897,600
effective HSP length: 80
effective length of query: 47
effective length of database: 2,620,800
effective search space: 123177600
effective search space used: 123177600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC145512.11


