

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.6 + phase: 2 /partial
(753 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
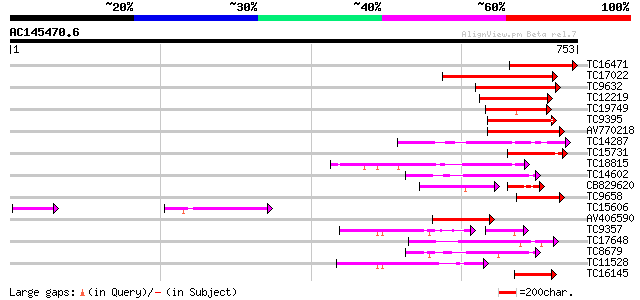
Score E
Sequences producing significant alignments: (bits) Value
TC16471 similar to UP|Q41384 (Q41384) Protein kinase (Fragment) ... 162 1e-40
TC17022 homologue to UP|Q39183 (Q39183) Serine/threonine protein... 152 2e-37
TC9632 similar to PIR|T47546|T47546 protein kinase-like - Arabid... 124 6e-29
TC12219 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like protei... 115 2e-26
TC19749 weakly similar to UP|KPK1_PHAVU (P15792) Protein kinase ... 96 2e-20
TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1... 95 5e-20
AV770218 92 3e-19
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 89 3e-18
TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Pr... 88 4e-18
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 88 6e-18
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 84 8e-17
CB829620 66 1e-16
TC9658 similar to UP|Q8H934 (Q8H934) Phototropin, partial (8%) 80 2e-15
TC15606 weakly similar to GB|AAL15406.1|16323302|AY058232 At2g02... 79 3e-15
AV406590 77 1e-14
TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein k... 52 2e-12
TC17648 similar to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (51%) 70 2e-12
TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partia... 69 2e-12
TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial... 65 4e-11
TC16145 similar to PIR|T00410|T00410 protein kinase homolog T13E... 64 9e-11
>TC16471 similar to UP|Q41384 (Q41384) Protein kinase (Fragment) , partial
(12%)
Length = 617
Score = 162 bits (411), Expect = 1e-40
Identities = 78/90 (86%), Positives = 82/90 (90%)
Frame = +3
Query: 664 FPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDV 723
FPSSIP SLAARQLINALLQRDPASRLGS TG+NEIKQHPFFR INWPLIRNMSPPPLDV
Sbjct: 3 FPSSIPVSLAARQLINALLQRDPASRLGSTTGANEIKQHPFFREINWPLIRNMSPPPLDV 182
Query: 724 PLQFIGKDPTAKDKKWEDDGVLNTSIDMDI 753
PLQ IGKDP AK+ WEDDGVL +S+DMDI
Sbjct: 183 PLQLIGKDPVAKNINWEDDGVLVSSVDMDI 272
>TC17022 homologue to UP|Q39183 (Q39183) Serine/threonine protein kinase
(Protein kinase 5) (AT5G47750/MCA23_7) , partial (32%)
Length = 775
Score = 152 bits (383), Expect = 2e-37
Identities = 73/153 (47%), Positives = 97/153 (62%), Gaps = 1/153 (0%)
Frame = +1
Query: 576 KQSLPGNRRRSRSQPPPIFVSEPV-TQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGIL 634
K P N ++ P P ++EP +S SFVGT EY+APEII G H SA+DWWT GI
Sbjct: 61 KDRKPKNEIGNQVSPLPELIAEPTDARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIF 240
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYE+L+G+TPF+G + T N++ + L FP + S AAR LI LL ++P RL
Sbjct: 241 LYELLFGKTPFKGSGNRATLFNVVGQPLRFPEAPVVSFAARDLIRGLLVKEPQHRLAYKR 420
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQF 727
G+ EIKQHPFF G+NW LIR +PP + ++F
Sbjct: 421 GATEIKQHPFFEGVNWALIRCATPPEIPKAVEF 519
>TC9632 similar to PIR|T47546|T47546 protein kinase-like - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (12%)
Length = 650
Score = 124 bits (311), Expect = 6e-29
Identities = 57/113 (50%), Positives = 75/113 (65%)
Frame = +3
Query: 619 GARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLI 678
G H +A+DWWT G+ LYE+LYGRTPF+G N ++T +N++ + L FP S S AR LI
Sbjct: 6 GEGHGAAVDWWTFGVFLYELLYGRTPFKGSNNEETLANVVLQGLRFPDSPFVSFQARDLI 185
Query: 679 NALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKD 731
LL ++P +RLG+ G+ EIKQHPFF G+NW LIR PP L +F D
Sbjct: 186 RGLLVKEPENRLGTEKGAAEIKQHPFFEGLNWALIRCAIPPELPDFCEFAFSD 344
>TC12219 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like protein, partial
(11%)
Length = 587
Score = 115 bits (289), Expect = 2e-26
Identities = 52/98 (53%), Positives = 69/98 (70%)
Frame = +3
Query: 624 SAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQ 683
+A+DWWT GI L+E+LYG+TPF+G + T +N++ + L FPS+ S AR LI LL
Sbjct: 3 NAVDWWTFGIFLFELLYGKTPFKGLANEDTLANVVSQSLKFPSAPIVSFHARDLIRGLLI 182
Query: 684 RDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPL 721
+DP +RLGS G+ EIKQHPF G+NW LIR +PP L
Sbjct: 183 KDPENRLGSVKGAAEIKQHPFSEGLNWALIRCAAPPEL 296
>TC19749 weakly similar to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 ,
partial (14%)
Length = 524
Score = 96.3 bits (238), Expect = 2e-20
Identities = 47/91 (51%), Positives = 63/91 (68%), Gaps = 3/91 (3%)
Frame = +3
Query: 632 GILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPAS---LAARQLINALLQRDPAS 688
GI +YEM+YGRTPF G + T +IL K L FP++ P+S + AR LI+ LL +DP
Sbjct: 6 GIFIYEMVYGRTPFAGPSNDVTLRSILKKPLVFPTATPSSALEMHARDLISGLLNKDPNR 185
Query: 689 RLGSATGSNEIKQHPFFRGINWPLIRNMSPP 719
RLGS G+ ++K+HPFF GIN LIR ++PP
Sbjct: 186 RLGSMRGAADVKKHPFFAGINLALIRMVAPP 278
>TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 ,
partial (17%)
Length = 590
Score = 94.7 bits (234), Expect = 5e-20
Identities = 47/92 (51%), Positives = 59/92 (64%)
Frame = +2
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYE+L+GRTPF+G + T N++ + L FP S S AAR LI LL ++P RL
Sbjct: 5 LYELLFGRTPFKGSANRATLFNVVGQPLRFPESPSVSFAARDLIRGLLVKEPQHRLAYRR 184
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQ 726
G+ EIKQHPFF +NW LIR SPP +VP Q
Sbjct: 185 GATEIKQHPFFHNVNWALIRCASPP--EVPRQ 274
>AV770218
Length = 567
Score = 92.0 bits (227), Expect = 3e-19
Identities = 47/102 (46%), Positives = 62/102 (60%)
Frame = -1
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYE+L GRTPF+G + T N++ + L+FP S S AAR LI LL ++P RL
Sbjct: 567 LYELLSGRTPFKGSAYRATLFNVVGQPLSFPVSPAVSFAARDLIRGLLVKEPQHRLAYRR 388
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKD 736
G+ EIKQHPFF +NW LIR +PP + P + PT K+
Sbjct: 387 GATEIKQHPFFHNVNWALIRCTNPPEVPRP-TLMRPSPTEKE 265
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 88.6 bits (218), Expect = 3e-18
Identities = 68/231 (29%), Positives = 103/231 (44%), Gaps = 1/231 (0%)
Frame = +3
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
L ED AR Y +++ +++ H G+ +RDLKPENLLL ++ + ++DF LS
Sbjct: 714 LNEDDARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDENEDLKVSDFGLS--------- 866
Query: 575 VKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHT-SAIDWWTLGI 633
+LP RR P GT Y+APE++ + S D W+ G+
Sbjct: 867 ---ALPEQRRDDGMLVTP-------------CGTPAYVAPEVLKKKGYDGSKADIWSCGV 998
Query: 634 LLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSA 693
+LY +L G PF+G+N + + + FP I S A+ LI+ LL DP R
Sbjct: 999 ILYALLSGYLPFQGENVMRIYRKAFKAEYEFPEWI--SPQAKNLISNLLVADPEKRYSIP 1172
Query: 694 TGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGV 744
EI P+F+ M P + +G+D T W + V
Sbjct: 1173----EIISDPWFQ------YGFMRPLAFSINESAVGEDSTI*SFWWGGEXV 1295
>TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Probable
serine/threonine-protein kinase PK5) , partial (17%)
Length = 940
Score = 88.2 bits (217), Expect = 4e-18
Identities = 45/80 (56%), Positives = 53/80 (66%)
Frame = +1
Query: 662 LTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPL 721
L FP S SL+ +QL+ LLQRDP+SRLGS G+NEIK+HPFFRGINW L+R PP L
Sbjct: 1 LKFPKSKQVSLSGKQLMYRLLQRDPSSRLGSKEGANEIKRHPFFRGINWALVRCTKPPEL 180
Query: 722 DVPLQFIGKDPTAKDKKWED 741
D PL G K K+ D
Sbjct: 181 DAPL--FGTTEEEKKAKYVD 234
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 87.8 bits (216), Expect = 6e-18
Identities = 74/279 (26%), Positives = 127/279 (44%), Gaps = 15/279 (5%)
Frame = +2
Query: 427 GEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKS------VMLNRNKSVFY 480
G IG + R LG G V+ + +GE A+K M K+ +M + +
Sbjct: 20 GHPIGKYEMG--RVLGKGTLAKVYFAKEITSGEGVAIKVMSKARIKKEGMMDQIKREISI 193
Query: 481 YRFIERA----LKERLYHYWIILFFQHYTPHFKQPMKI----LKEDSARFYAAEVVIGLE 532
R + LKE + I F Y + K+ LK+D AR Y +++ ++
Sbjct: 194 MRLVRHPNIVNLKEVMATKTKIFFIMEYIRGGELFAKVAKGKLKDDLARRYFQQLISAVD 373
Query: 533 YLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPP 592
Y H G+ +RDLKPENLLL ++ ++ ++DF LS + P+ ++Q
Sbjct: 374 YCHSRGVSHRDLKPENLLLDENENLKVSDFGLSGL----PEQLRQD-------------- 499
Query: 593 IFVSEPVTQSNSFVGTEEYIAPEIITGARHTS-AIDWWTLGILLYEMLYGRTPFRGKNRQ 651
++ GT Y+APE++ + D W+ G++LY +L G PF+ +N
Sbjct: 500 -------GLLHTQCGTPAYVAPEVLRKKGYDGFKTDTWSCGVILYALLAGCLPFQHENLM 658
Query: 652 KTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRL 690
++ +L + FP S +++LI+ +L DP R+
Sbjct: 659 TMYNKVLRAEFQFPPWF--SPESKKLISKILVADPNRRI 769
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 84.0 bits (206), Expect = 8e-17
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 2/182 (1%)
Frame = +2
Query: 526 EVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRR 585
+++ + + H LG+ +RD+KP+N+L G + L DF G+ RR
Sbjct: 395 QLLEAVAHCHRLGVAHRDVKPDNVLFGGGGDLKLADFG------------SAEWFGDGRR 538
Query: 586 SRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPF 645
+ VGT Y+APE++ G + +D W+ G++LY ML G PF
Sbjct: 539 ----------------MSGVVGTPYYVAPEVLMGREYGEKVDVWSCGVILYIMLSGTPPF 670
Query: 646 RGKNRQKTFSNILHKDLTFPSSI--PASLAARQLINALLQRDPASRLGSATGSNEIKQHP 703
G + + F ++ +L FPS I S AA+ L+ ++ RDP++R+ + + +HP
Sbjct: 671 YGDSAAEIFEAVIRGNLRFPSRIFRNVSPAAKDLLRKMICRDPSNRI----SAEQALRHP 838
Query: 704 FF 705
+F
Sbjct: 839 WF 844
>CB829620
Length = 548
Score = 65.9 bits (159), Expect(2) = 1e-16
Identities = 41/120 (34%), Positives = 56/120 (46%), Gaps = 14/120 (11%)
Frame = +1
Query: 545 KPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSN- 603
KP+NLLL ++GH+ L+DF L C G+ R Q V+ TQ
Sbjct: 1 KPDNLLLDRNGHMKLSDFGLCKPLDCSNLQENDFSTGSNRSGALQSNGRPVAPKRTQQEQ 180
Query: 604 -------------SFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNR 650
S VGT +YIAPE++ + DWW+LG ++YEML G PF N+
Sbjct: 181 LQHWQKNRRTLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGGIMYEMLVGYPPFIQMNQ 360
Score = 38.1 bits (87), Expect(2) = 1e-16
Identities = 22/49 (44%), Positives = 30/49 (60%)
Frame = +3
Query: 662 LTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINW 710
L FP S A+ LI+ LL + RLG+ G++EIK HP+F+GI W
Sbjct: 399 LKFPEEAKLSPEAKDLISRLLC-NVQQRLGTK-GADEIKAHPWFKGIEW 539
>TC9658 similar to UP|Q8H934 (Q8H934) Phototropin, partial (8%)
Length = 536
Score = 79.7 bits (195), Expect = 2e-15
Identities = 40/64 (62%), Positives = 47/64 (72%), Gaps = 1/64 (1%)
Frame = +2
Query: 674 ARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVP-LQFIGKDP 732
A+QLI LL RDP +RLGS G+NEIK+HPFFRG++W LIR M PP LD P LQ +D
Sbjct: 5 AKQLIYRLLHRDPKNRLGSQEGANEIKRHPFFRGVDWALIRCMKPPELDAPLLQETEEDK 184
Query: 733 TAKD 736
AKD
Sbjct: 185 EAKD 196
>TC15606 weakly similar to GB|AAL15406.1|16323302|AY058232
At2g02710/T20F6.15 {Arabidopsis thaliana;}, partial
(25%)
Length = 642
Score = 78.6 bits (192), Expect = 3e-15
Identities = 50/152 (32%), Positives = 82/152 (53%), Gaps = 9/152 (5%)
Frame = +1
Query: 206 DKTIVEPEVLMTKEIEWSKYELRE--------RDIRQGIDLATTLERIEKNFVISDPRLP 257
+++ V LMT E E SK E R R+ +D E + +F I+DP +
Sbjct: 202 ERSTVVATTLMTTEFEDSKLLAVESSFDNRYSRHARESLD-----ELTDTSFTITDPSVS 366
Query: 258 DCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVNRIRDAIKDQREITVQLINY 317
PI+FAS FL++T ++R+E++GR QGP T + +V IR+A+++++E V +NY
Sbjct: 367 GHPIVFASLGFLKMTGFSRDEVVGRGGGMFQGPGTCRRSVMAIREAVREEKEAQVVXLNY 546
Query: 318 TKSGKKFWNLFHLQPM-RDQKGELQYFIGVQL 348
K G F L + P+ G + +F+ VQ+
Sbjct: 547 RKDGTPFXMLLLVCPVFSASSGGVVHFVAVQV 642
Score = 45.4 bits (106), Expect = 3e-05
Identities = 25/62 (40%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +1
Query: 4 QGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPI-KDDRGNTIKFIGM 62
QGP T + V IR+A + K LNY+K+GTPF LL V P+ G + F+ +
Sbjct: 457 QGPGTCRRSVMAIREAVREEKEAQVVXLNYRKDGTPFXMLLLVCPVFSASSGGVVHFVAV 636
Query: 63 QV 64
QV
Sbjct: 637 QV 642
>AV406590
Length = 406
Score = 76.6 bits (187), Expect = 1e-14
Identities = 38/83 (45%), Positives = 52/83 (61%), Gaps = 1/83 (1%)
Frame = +3
Query: 562 FDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVT-QSNSFVGTEEYIAPEIITGA 620
F F++ + K+S P ++ P P ++EP +S SFVGT EY+APEI+ G
Sbjct: 156 FTPRFLSGKSKKDKKKSKPKTDIHNQVTPLPELMAEPTNARSMSFVGTHEYLAPEIVKGE 335
Query: 621 RHTSAIDWWTLGILLYEMLYGRT 643
H SA+DWWT GI LYE+L+GRT
Sbjct: 336 GHGSAVDWWTFGIFLYELLFGRT 404
>TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein kinase,
41K) , partial (71%)
Length = 1181
Score = 52.4 bits (124), Expect(2) = 2e-12
Identities = 53/196 (27%), Positives = 88/196 (44%), Gaps = 15/196 (7%)
Frame = +1
Query: 438 IRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN--KSVFYYRFIERA----LKER 491
+R +G G+ G L++ + T EL A+K +E+ ++ N + + +R + KE
Sbjct: 370 VRDIGSGNFGVARLMQDKQTKELVAVKYIERGDKIDENVKREIINHRSLRHPNIVRFKEV 549
Query: 492 LY---HYWIILFFQHYTPHFKQPMKI--LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKP 546
+ H I++ + F++ ED ARF+ +++ G+ Y H + + +RDLK
Sbjct: 550 ILTPTHLAIVMEYASGGELFERICNAGRFTEDEARFFFQQLISGVSYCHAMQVCHRDLKL 729
Query: 547 ENLLLQKDG----HIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQS 602
EN LL DG + + DF S K V+ SQP
Sbjct: 730 ENTLL--DGSPTPRLKICDFGYS-----KSSVL-----------HSQP------------ 819
Query: 603 NSFVGTEEYIAPEIIT 618
S VGT YIAPE+++
Sbjct: 820 KSTVGTPAYIAPEVLS 867
Score = 37.4 bits (85), Expect(2) = 2e-12
Identities = 23/62 (37%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Frame = +3
Query: 632 GILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIP----ASLAARQLINALLQRDPA 687
G+ LY ML G PF N K F + + L+ SIP S R LI+ + DPA
Sbjct: 912 GVTLYVMLVGAYPFEDPNEPKDFRKTIQRVLSVQYSIPDFVQISPECRHLISRIFVFDPA 1091
Query: 688 SR 689
R
Sbjct: 1092ER 1097
>TC17648 similar to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (51%)
Length = 741
Score = 69.7 bits (169), Expect = 2e-12
Identities = 53/212 (25%), Positives = 95/212 (44%), Gaps = 13/212 (6%)
Frame = +3
Query: 530 GLEYLHC-LGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRS 588
GL YLH +I+RD KP +LL+ G + +TDF +S +
Sbjct: 9 GLVYLHNERHVIHRDSKPSHLLVNHQGEVKITDFGVSAM--------------------- 125
Query: 589 QPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGK 648
++ + Q ++FVGT Y++P I+G+ + + D W+LG+++ E GR P+
Sbjct: 126 ------LATSMGQRDTFVGTYNYMSPARISGSTYDYSSDIWSLGMVVLECAIGRFPYIQS 287
Query: 649 NRQKTFSNILHKDLTFPSSIPASLAARQ-------LINALLQRDPASRLGSATGSNEIKQ 701
Q+ + + S P S + Q +++ +Q+DP RL S E+
Sbjct: 288 EDQQAWPSFYELLAAIVESPPPSAPSDQFSPEFCSFVSSCIQKDPQDRLTSL----ELLD 455
Query: 702 HPFF-----RGINWPLIRNMSPPPLDVPLQFI 728
HPF + ++ ++ PP++ P F+
Sbjct: 456 HPFIKKFEDKDLDLEILVGSLEPPINFPR*FL 551
>TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partial (70%)
Length = 972
Score = 69.3 bits (168), Expect = 2e-12
Identities = 57/189 (30%), Positives = 83/189 (43%), Gaps = 9/189 (4%)
Frame = +2
Query: 526 EVVIGLEYLHCLGIIYRDLKPENLLLQKDG----HIVLTDFDLSFITSCKPQVVKQSLPG 581
+++ G+ Y H L I +RDLK EN LL DG + + DF S K SL
Sbjct: 5 QLISGVHYCHALQICHRDLKLENTLL--DGSPAPRLKICDFGYS----------KSSLLH 148
Query: 582 NRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAI-DWWTLGILLYEMLY 640
+R +S VGT YIAPE+++ + + D W+ G+ LY ML
Sbjct: 149 SRPKST------------------VGTPAYIAPEVLSRREYDGKLADVWSCGVTLYVMLV 274
Query: 641 GRTPFRG----KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGS 696
G PF +N +KT I+ P + S R L++ + +P R+
Sbjct: 275 GAYPFEDHDDPRNFRKTIQRIMAVQYKIPDYVHISQDCRHLLSRIFVANPLRRI----SL 442
Query: 697 NEIKQHPFF 705
EIK HP+F
Sbjct: 443 KEIKNHPWF 469
>TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial (56%)
Length = 640
Score = 65.1 bits (157), Expect = 4e-11
Identities = 57/213 (26%), Positives = 93/213 (42%), Gaps = 12/213 (5%)
Frame = +3
Query: 435 FSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN--KSVFYYRFIERA----L 488
+ P++ LG G+ G L + TGEL A+K +E+ ++ N + + +R +
Sbjct: 78 YEPLKELGSGNFGVARLARDKNTGELVAVKYIERGKKIDENVQREIINHRSLRHPNIIRF 257
Query: 489 KERLY---HYWIILFFQHYTPHFKQPMKI--LKEDSARFYAAEVVIGLEYLHCLGIIYRD 543
KE L H I+L + F++ ED AR++ +++ G+ Y H + I +RD
Sbjct: 258 KEVLLTPTHLAIVLEYASGGELFERICSAGRFSEDEARYFFQQLISGVSYCHSMEICHRD 437
Query: 544 LKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSN 603
LK EN LL + L D + S I S+P
Sbjct: 438 LKLENTLLDGNPSPRLKICDFGYSKSA----------------------ILHSQP----K 539
Query: 604 SFVGTEEYIAPEIITGARHTSAI-DWWTLGILL 635
S VGT YIAPE+++ + + D W+ G+ L
Sbjct: 540 STVGTPAYIAPEVLSRKEYDGKVADVWSCGVTL 638
>TC16145 similar to PIR|T00410|T00410 protein kinase homolog T13E15.16 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(10%)
Length = 535
Score = 63.9 bits (154), Expect = 9e-11
Identities = 29/56 (51%), Positives = 37/56 (65%)
Frame = +2
Query: 671 SLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQ 726
S A R LI LL ++P RLG G+ EIKQHPFF G+NW LIR +PP + P++
Sbjct: 14 SYAGRDLIRGLLVKEPQHRLGVKRGATEIKQHPFFEGVNWALIRCSTPPEVPRPVE 181
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.136 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,254,685
Number of Sequences: 28460
Number of extensions: 162870
Number of successful extensions: 1063
Number of sequences better than 10.0: 156
Number of HSP's better than 10.0 without gapping: 1025
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1044
length of query: 753
length of database: 4,897,600
effective HSP length: 97
effective length of query: 656
effective length of database: 2,136,980
effective search space: 1401858880
effective search space used: 1401858880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC145470.6


