

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
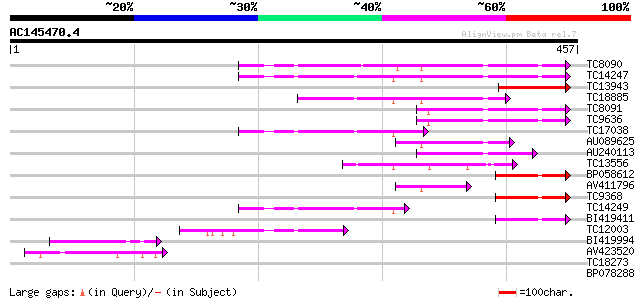
Query= AC145470.4 + phase: 0
(457 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8090 homologue to UP|ACTB_ARATH (P53496) Actin 11, complete 127 4e-30
TC14247 homologue to UP|Q9SLQ6 (Q9SLQ6) Actin isoform B, complete 124 3e-29
TC13943 weakly similar to GB|AAM53248.1|21427471|AF507916 actin-... 103 5e-23
TC18885 homologue to UP|Q96442 (Q96442) Actin (Fragment), partia... 85 2e-17
TC8091 homologue to UP|ACT3_SOLTU (P30167) Actin 58, partial (39%) 75 2e-14
TC9636 homologue to UP|Q9M729 (Q9M729) Actin, partial (41%) 75 2e-14
TC17038 homologue to UP|ACT1_SORBI (P53504) Actin 1, partial (63%) 63 1e-10
AU089625 63 1e-10
AU240113 62 2e-10
TC13556 similar to UP|Q9M729 (Q9M729) Actin, partial (37%) 62 3e-10
BP058612 53 9e-08
AV411796 53 9e-08
TC9368 UP|Q41211 (Q41211) Actin (Fragment), partial (89%) 52 2e-07
TC14249 homologue to UP|Q7XZJ2 (Q7XZJ2) Actin, partial (63%) 51 3e-07
BI419411 49 2e-06
TC12003 homologue to UP|AAQ55798 (AAQ55798) Actin, partial (50%) 46 1e-05
BI419994 45 3e-05
AV423520 44 6e-05
TC18273 similar to AAQ06240 (AAQ06240) Tubby-like protein TULP1,... 40 0.001
BP078288 39 0.002
>TC8090 homologue to UP|ACTB_ARATH (P53496) Actin 11, complete
Length = 1624
Score = 127 bits (319), Expect = 4e-30
Identities = 87/278 (31%), Positives = 141/278 (50%), Gaps = 10/278 (3%)
Frame = +2
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 368 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NTPAMYVAIQA 517
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT FL + + +
Sbjct: 518 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPHAILRL-DLAGRDLTDFLMKILTERG 694
Query: 304 LYFESLYT---VRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL Y+A DYE E+ S S+E +G+ T+ ERF+
Sbjct: 695 YSFTTSAXREIVRDVKEKLAYIALDYEQEIETARTSSSVEKSYELPDGQVITIGDERFRC 874
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP L G+ A +H + C ++ D + +VLSGG+ PG+A+R+
Sbjct: 875 PEVLFQPSLVGMEAAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGTTMFPGIADRMS 1045
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 1046KEISALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 1153
>TC14247 homologue to UP|Q9SLQ6 (Q9SLQ6) Actin isoform B, complete
Length = 1656
Score = 124 bits (311), Expect = 3e-29
Identities = 87/278 (31%), Positives = 141/278 (50%), Gaps = 10/278 (3%)
Frame = +2
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + NVPA+ QA
Sbjct: 416 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NVPAMYVAIQA 565
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 566 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDSLMKILTERG 742
Query: 304 LYFES---LYTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKL YVA DYE EL S ++E +G+ T+ ERF+
Sbjct: 743 YMFTTSAEREIVRDIKEKLAYVALDYEQELETAKSSSSIEKNYELPDGQVITIGAERFRC 922
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP + G+ + +H + C ++ D + +VLSGGS PG+A+R+
Sbjct: 923 PEVLFQPSMIGMESAGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGSTMFPGIADRMS 1093
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L P M I+V+ PP + W G ++ S+S
Sbjct: 1094KEITALAPSSMK--IKVVAPPERKYSVWIGGSILASLS 1201
>TC13943 weakly similar to GB|AAM53248.1|21427471|AF507916 actin-related
protein 8B {Arabidopsis thaliana;}, partial (16%)
Length = 478
Score = 103 bits (258), Expect = 5e-23
Identities = 46/58 (79%), Positives = 56/58 (96%)
Frame = +1
Query: 395 KTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KTVVLSGG+ACL GLAER++KEL++LLPPYMSNGIRV+PPP+GADT WFGAK+IG++S
Sbjct: 1 KTVVLSGGTACLTGLAERIDKELNALLPPYMSNGIRVVPPPYGADTPWFGAKIIGNLS 174
>TC18885 homologue to UP|Q96442 (Q96442) Actin (Fragment), partial (55%)
Length = 561
Score = 85.1 bits (209), Expect = 2e-17
Identities = 61/180 (33%), Positives = 93/180 (50%), Gaps = 9/180 (5%)
Frame = +3
Query: 233 NVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLT 292
NVPA+ QA L+LYA+ +T+GI ++ G V+ VPI G + + + L LT
Sbjct: 27 NVPAMYVAIQAVLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLT 203
Query: 293 GFLREKMQQNNLYFES---LYTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK 344
+L + + + F + VR +KEKL YVA D+E E+ S S+E +G+
Sbjct: 204 EYLVKILTERGYSFSTSAEKEIVRDVKEKLAYVALDFEQEMDTTKSSSAVEKSYELPDGQ 383
Query: 345 -FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGS 403
T+ ERF+ E+LFQP L G+ A +H + C ++ D + +VLSGGS
Sbjct: 384 VITIGAERFRCPEVLFQPSLIGMEAPGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGS 554
>TC8091 homologue to UP|ACT3_SOLTU (P30167) Actin 58, partial (39%)
Length = 640
Score = 75.5 bits (184), Expect = 2e-14
Identities = 44/128 (34%), Positives = 70/128 (54%), Gaps = 4/128 (3%)
Frame = +1
Query: 329 ELSKDTHA---SFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHS 384
E SK + A S+E +G+ T+ ERF+ E+LFQP L G+ A +H + C
Sbjct: 1 ETSKTSSAVEKSYELPDGQVITIGAERFRCPEVLFQPSLVGMEAAGIHETTYNSIMKC-- 174
Query: 385 AELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFG 444
++ D + +VLSGG+ PG+A+R+ KE+ +L P M I+V+ PP + W G
Sbjct: 175 -DVDIRKDLYGNIVLSGGTTMFPGIADRMSKEISALAPSSMK--IKVVAPPERKYSVWIG 345
Query: 445 AKLIGSVS 452
++ S+S
Sbjct: 346 GSILASLS 369
>TC9636 homologue to UP|Q9M729 (Q9M729) Actin, partial (41%)
Length = 792
Score = 75.1 bits (183), Expect = 2e-14
Identities = 44/128 (34%), Positives = 70/128 (54%), Gaps = 4/128 (3%)
Frame = +3
Query: 329 ELSKDTHA---SFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHS 384
E SK + A S+E +G+ T+ ERF+ E+LFQP + G+ A +H + C
Sbjct: 36 ETSKTSSAVEKSYELPDGQVITIGAERFRCPEVLFQPSMIGMEAPGIHETTYNSIMKC-- 209
Query: 385 AELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFG 444
++ D + +VLSGGS PG+A+R+ KE+ +L P M I+V+ PP + W G
Sbjct: 210 -DVDIRKDLYGNIVLSGGSTMFPGIADRMSKEISALAPSSMK--IKVVAPPERKYSVWIG 380
Query: 445 AKLIGSVS 452
++ S+S
Sbjct: 381 GSILASLS 404
>TC17038 homologue to UP|ACT1_SORBI (P53504) Actin 1, partial (63%)
Length = 761
Score = 62.8 bits (151), Expect = 1e-10
Identities = 47/157 (29%), Positives = 77/157 (48%), Gaps = 4/157 (2%)
Frame = +2
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 323 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NAPAMYVAIQA 472
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT FL + + +
Sbjct: 473 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDFLMKILTERG 649
Query: 304 LYFES---LYTVRTLKEKLCYVAYDYEAELSKDTHAS 337
F + VR +KEKL Y+A DYE E+ +S
Sbjct: 650 YSFTTSAEREIVRDMKEKLAYIALDYEQEIETSKTSS 760
>AU089625
Length = 353
Score = 62.8 bits (151), Expect = 1e-10
Identities = 38/102 (37%), Positives = 53/102 (51%), Gaps = 6/102 (5%)
Frame = +3
Query: 312 VRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQTGEILFQPRLAG 365
VR +KEKL Y+A DYE E+ S SFE +G+ T+ ERF+ E+LFQP G
Sbjct: 57 VRDMKEKLAYIALDYEQEIETSNTSSSVEKSFELPDGQVITIGAERFRCPEVLFQPSTIG 236
Query: 366 VRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLP 407
+ A +H + C ++ D + +VLSGGS P
Sbjct: 237 MEAPGIHETTYNSIMKC---DVDIRKDLYGNIVLSGGSTMFP 353
>AU240113
Length = 300
Score = 62.4 bits (150), Expect = 2e-10
Identities = 34/98 (34%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Frame = +3
Query: 329 ELSKDTHASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAEL 387
E S S+E +G+ T+ ERF+ E+LFQP + G+ A+ +H + C ++
Sbjct: 6 ETSSAVEKSYELPDGQVITIGAERFRCPEVLFQPSMIGMEAVGIHETTYNSIMKC---DV 176
Query: 388 AGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYM 425
D + +VLSGGS PG+A+R+ KE+ +L P M
Sbjct: 177 DIRKDLYGNIVLSGGSTMFPGIADRMSKEITALAPSSM 290
>TC13556 similar to UP|Q9M729 (Q9M729) Actin, partial (37%)
Length = 497
Score = 61.6 bits (148), Expect = 3e-10
Identities = 48/155 (30%), Positives = 78/155 (49%), Gaps = 14/155 (9%)
Frame = +2
Query: 269 PILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYFES---LYTVRTLKEKLCYVAYD 325
P+ G V+ ++ + G +T +L + + + F + VR +KEKL Y+A +
Sbjct: 44 PVYEGYVLPHA-IQSLNFGGSDITKYLIDILGERGYSFTNSSECKIVRDMKEKLAYLALN 220
Query: 326 YEAELSKDTHAS---FEAAEGK-FTLSKERFQTGEILFQPRLAGVR-------AMSLHHA 374
YE EL+ +S +E +G T+ ERF+ E+LFQP L G++ A +H
Sbjct: 221 YEHELATAKTSSCVDYELPDGHVITIGDERFKCPEVLFQPSLIGMKVPHTNDPAHGIHEI 400
Query: 375 IALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGL 409
I + C S E+ D++ +VLSGGS PG+
Sbjct: 401 INESIMKCDS-EIR--KDFYGNIVLSGGSTMFPGI 496
>BP058612
Length = 368
Score = 53.1 bits (126), Expect = 9e-08
Identities = 25/61 (40%), Positives = 38/61 (61%)
Frame = -2
Query: 392 DWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSV 451
D + VVLSGG+ PG+A+R++KEL +L P M I++I PP + W G ++ S+
Sbjct: 337 DLYGNVVLSGGTTMFPGIADRMQKELTALAPSTMK--IKIIAPPERKYSVWIGGSILASL 164
Query: 452 S 452
S
Sbjct: 163 S 161
>AV411796
Length = 365
Score = 53.1 bits (126), Expect = 9e-08
Identities = 28/67 (41%), Positives = 40/67 (58%), Gaps = 6/67 (8%)
Frame = +3
Query: 312 VRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQTGEILFQPRLAG 365
VR +KEKL Y+A DYE EL S S+E +G+ T+ ERF+ E+LFQP + G
Sbjct: 150 VRDMKEKLAYIALDYEQELETSKTSSAVEKSYELPDGQVITIGAERFRCPEVLFQPSMIG 329
Query: 366 VRAMSLH 372
+ + +H
Sbjct: 330 MESPGIH 350
>TC9368 UP|Q41211 (Q41211) Actin (Fragment), partial (89%)
Length = 592
Score = 52.4 bits (124), Expect = 2e-07
Identities = 24/61 (39%), Positives = 37/61 (60%)
Frame = +2
Query: 392 DWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSV 451
D + +VLSGGS PG+A+R+ KE+ +L P M I+V+ PP + W G ++ S+
Sbjct: 56 DLYGNIVLSGGSTMFPGIADRMSKEITALAPSSMK--IKVVAPPERKYSVWIGGSILASL 229
Query: 452 S 452
S
Sbjct: 230 S 232
>TC14249 homologue to UP|Q7XZJ2 (Q7XZJ2) Actin, partial (63%)
Length = 916
Score = 51.2 bits (121), Expect = 3e-07
Identities = 42/142 (29%), Positives = 69/142 (48%), Gaps = 4/142 (2%)
Frame = +2
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + NVPA+ QA
Sbjct: 476 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NVPAMYVAIQA 625
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G V+ VPI G + + + L LT L + + +
Sbjct: 626 VLSLYASGRTTGIVLDSGDGVSHTVPIYEGYALPH-AILRLDLAGRDLTDSLMKILTERG 802
Query: 304 LYFES---LYTVRTLKEKLCYV 322
F + VR +KEKL YV
Sbjct: 803 YMFTTSAEREIVRDIKEKLAYV 868
>BI419411
Length = 408
Score = 48.5 bits (114), Expect = 2e-06
Identities = 22/61 (36%), Positives = 36/61 (58%)
Frame = +2
Query: 392 DWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSV 451
D + VLSGG+ G+AER++KE+ +L P M I+++ PP + W G ++ S+
Sbjct: 11 DLYGNTVLSGGTTMFEGIAERMQKEITALAPASMK--IKIVAPPERKYSVWIGGSILASL 184
Query: 452 S 452
S
Sbjct: 185 S 187
>TC12003 homologue to UP|AAQ55798 (AAQ55798) Actin, partial (50%)
Length = 612
Score = 45.8 bits (107), Expect = 1e-05
Identities = 44/172 (25%), Positives = 74/172 (42%), Gaps = 36/172 (20%)
Frame = +1
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 61 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHQGVMVGMGQKDAYVGDEAQSKRGILT 240
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 241 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 399
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNG 273
+ + N PA+ QA L+LYA+ +T+GI ++ G V+ VPI G
Sbjct: 400 TQIMFETF---NTPAMYVAIQAVLSLYASGRTTGIVLDSGDGVSHTVPIYEG 546
>BI419994
Length = 549
Score = 44.7 bits (104), Expect = 3e-05
Identities = 29/91 (31%), Positives = 44/91 (47%), Gaps = 1/91 (1%)
Frame = +2
Query: 33 HHHHAPSPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCKSWRSLVSDNELW-AHFLQTH 91
H + L LF LP +++L+I L + + VCKSWR L+SD + +HF
Sbjct: 5 HPQTSTESLSLFSHLPDELILQILLRLPVRSLLRFKTVCKSWRYLISDPQFTKSHFNLAA 184
Query: 92 QPIHFHSILFSETNLTSGYPLPIFDTQTTPH 122
P H I + TN G+ + DT ++ H
Sbjct: 185 SPT--HRIFLNNTN---GF*IESLDTDSSLH 262
>AV423520
Length = 468
Score = 43.9 bits (102), Expect = 6e-05
Identities = 39/127 (30%), Positives = 59/127 (45%), Gaps = 12/127 (9%)
Frame = +3
Query: 13 SIKRSTNTTNS----SSSSRSHQHHHH-HAPSPLGLFDSLPPDILLKITRLLGPKHAAKL 67
S+K+ T +T + S S S+ H H P PL +LP +++++I L K +
Sbjct: 57 SVKQFTTSTEALTPPSFPSSSNPHGDSLHPPPPL---PTLPFELVVEILCRLPVKSLLQF 227
Query: 68 CLVCKSWRSLVSDNELWA--HFLQTHQPIHFHSILFSETN---LTSGYPLPIFD--TQTT 120
VCKSW SL+S + +A H + + H ++F TN TS L +FD
Sbjct: 228 RCVCKSWNSLISGDPKFARKHLRCSPKDFTRHHLIFGSTNEFLFTSSPLLSVFDDAAAIA 407
Query: 121 PHVSFKH 127
P F H
Sbjct: 408 PSTQFDH 428
>TC18273 similar to AAQ06240 (AAQ06240) Tubby-like protein TULP1, partial
(46%)
Length = 1016
Score = 39.7 bits (91), Expect = 0.001
Identities = 28/92 (30%), Positives = 42/92 (45%), Gaps = 13/92 (14%)
Frame = +2
Query: 2 LLRKLCDSASSSIKRSTNTT---NSSSSSRSHQHHHHHAPSPL--GLFDSLPPDILLKIT 56
++R + D S +RS + +S S SRS H H P + + SLPP++L +
Sbjct: 320 IVRDVRDGFGSLSRRSFDVILPGHSRSKSRSSVHELHDQPLVIQSSRWASLPPELLRDVI 499
Query: 57 RLL--------GPKHAAKLCLVCKSWRSLVSD 80
+ L G KH VCKSWR + +
Sbjct: 500 KRLETNESAWPGRKHVVACAAVCKSWREMCKE 595
>BP078288
Length = 428
Score = 38.5 bits (88), Expect = 0.002
Identities = 24/102 (23%), Positives = 45/102 (43%), Gaps = 9/102 (8%)
Frame = -3
Query: 359 FQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKELH 418
F P L V + + + C++ +++ ++++ +VLSGG+ PGL RLEKE+
Sbjct: 426 FSPHLVDVESPGVAELVFDCIN---KSDIDTRAEFYNHIVLSGGTTMYPGLPSRLEKEIR 256
Query: 419 SLLPPYMSNG---------IRVIPPPHGADTAWFGAKLIGSV 451
L + G R+ PP + G ++ +
Sbjct: 255 RLYFERVCKGNQAQLAKFKCRIEDPPRRKHMVFLGGAVLAEI 130
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,390,101
Number of Sequences: 28460
Number of extensions: 165053
Number of successful extensions: 3224
Number of sequences better than 10.0: 183
Number of HSP's better than 10.0 without gapping: 2168
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2828
length of query: 457
length of database: 4,897,600
effective HSP length: 93
effective length of query: 364
effective length of database: 2,250,820
effective search space: 819298480
effective search space used: 819298480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145470.4


