

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
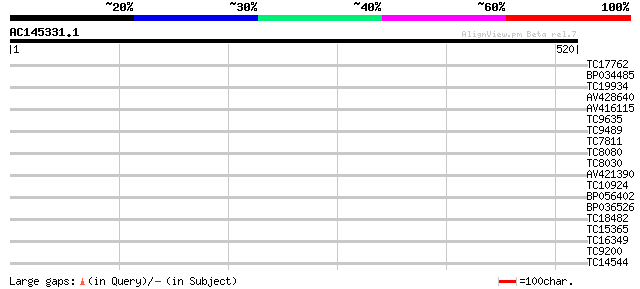
Query= AC145331.1 - phase: 0
(520 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17762 34 0.066
BP034485 33 0.15
TC19934 30 0.73
AV428640 30 1.3
AV416115 29 1.6
TC9635 28 2.8
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 28 3.6
TC7811 homologue to PIR|T04368|T04368 plasma membrane intrinsic ... 28 4.8
TC8080 27 6.2
TC8030 27 6.2
AV421390 27 6.2
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 27 8.1
BP056402 27 8.1
BP036526 27 8.1
TC18482 27 8.1
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 27 8.1
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 27 8.1
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 27 8.1
TC14544 27 8.1
>TC17762
Length = 463
Score = 33.9 bits (76), Expect = 0.066
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = -3
Query: 36 WWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKK 74
WW+R+ K T RW ++R+R WV+ K+E++
Sbjct: 185 WWRRVRVKETWRWQHARERKRLAVVAAWPWVAAAKKERR 69
>BP034485
Length = 485
Score = 32.7 bits (73), Expect = 0.15
Identities = 14/45 (31%), Positives = 26/45 (57%), Gaps = 2/45 (4%)
Frame = +2
Query: 11 WEKKRENERRRILMDEVEKQQNDKDWWKR--LNNKITERWTNSRK 53
WE+++E RRR D ++Q+ +++ W+R + ERW R+
Sbjct: 128 WEEEQEGRRRRRR*DSKQEQEQEQEQWRRKQQRERRKERWRRRRE 262
>TC19934
Length = 384
Score = 30.4 bits (67), Expect = 0.73
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +3
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEW 169
E H WK+ +K L+ SW+ N P T+ K W
Sbjct: 165 ELFHPAWKRHHCKKVLKKIFQSWLTN*PLCTQHKTW 272
>AV428640
Length = 368
Score = 29.6 bits (65), Expect = 1.3
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +2
Query: 75 LDSLVGRRPTAPPAPKGLYIYGNV 98
L S + +RP PP PKGL I GN+
Sbjct: 95 LASRIRKRPPYPPGPKGLPIIGNM 166
>AV416115
Length = 430
Score = 29.3 bits (64), Expect = 1.6
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +1
Query: 118 KHRRRYHFHEAMLRINEHMHKTWKKQMEEK 147
+HR R+H H N H + WKK+ EE+
Sbjct: 52 RHRNRHHHH------NHHERRRWKKRPEEE 123
>TC9635
Length = 585
Score = 28.5 bits (62), Expect = 2.8
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +3
Query: 367 NYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHS 402
NYH VF +P MR+R + I + ++Y HHS
Sbjct: 42 NYHRVFKGIMPSPGMRVRLSNKLQIQGMPKVYWHHS 149
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 28.1 bits (61), Expect = 3.6
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +1
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSAT 113
PKG+ +YG G+GKT+L ++T
Sbjct: 484 PKGVILYGEPGTGKTLLAKAVANST 558
>TC7811 homologue to PIR|T04368|T04368 plasma membrane intrinsic protein
BPW2 - barley {Hordeum vulgare;} , partial (15%)
Length = 464
Score = 27.7 bits (60), Expect = 4.8
Identities = 14/43 (32%), Positives = 19/43 (43%)
Frame = +1
Query: 363 AVAENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLC 405
A+A YH V I +P S + T + E +N CLC
Sbjct: 46 ALAAIYHVVVIRALPFQSQ*LESNGSLIRTTVWENHNPSHCLC 174
>TC8080
Length = 1432
Score = 27.3 bits (59), Expect = 6.2
Identities = 17/58 (29%), Positives = 30/58 (51%)
Frame = -1
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILP 187
L+I+ H+H Q EE+ + + WI+ PF + +L A + K++ + H LP
Sbjct: 580 LQISRHLH-----QEEEETI---L*DWILENPFSCRQIYFLTARKTVKQKEFLVHQLP 431
>TC8030
Length = 991
Score = 27.3 bits (59), Expect = 6.2
Identities = 17/53 (32%), Positives = 22/53 (41%)
Frame = -1
Query: 15 RENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVS 67
RENER R WW R+ K T RW ++R+R WV+
Sbjct: 247 RENERGR--------GDGGCPWWWRVPVKDTWRWQHARERKRLAVVAAWPWVT 113
>AV421390
Length = 409
Score = 27.3 bits (59), Expect = 6.2
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +2
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
V YL+ K L G+ PKG+ + G G+GKT+L
Sbjct: 119 VEYLRNPAKFTRLGGK------LPKGILLTGAPGTGKTLL 220
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 26.9 bits (58), Expect = 8.1
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +2
Query: 89 PKGLYIYGNVGSGKTML 105
PKG+ +YG G+GKT+L
Sbjct: 560 PKGVLLYGPPGTGKTLL 610
>BP056402
Length = 475
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/39 (28%), Positives = 21/39 (53%)
Frame = +3
Query: 41 NNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLDSLV 79
+N+ T RW + K +N++PG Y E+ +D ++
Sbjct: 189 SNEATSRWRPTNKTTDNLNPGT---ACYSAAEESVDKII 296
>BP036526
Length = 531
Score = 26.9 bits (58), Expect = 8.1
Identities = 24/96 (25%), Positives = 42/96 (43%), Gaps = 7/96 (7%)
Frame = +2
Query: 208 CFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKD------LNEA-NMVPEFFQN 260
CF E++ + A SG+ + + S T IV + P NE ++ P+ +
Sbjct: 47 CFAEVKQLSFSAKTLYSGLHALVPSLSTAIVDKNLGFPVFSAIDALFNEGLSLPPQKARG 226
Query: 261 LLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLW 296
LLS L K++ +E D RF + ++ + W
Sbjct: 227 LLSGLLPRLVKLIQDTENDVLRFETPATMDKDRFFW 334
>TC18482
Length = 530
Score = 26.9 bits (58), Expect = 8.1
Identities = 15/53 (28%), Positives = 29/53 (54%), Gaps = 4/53 (7%)
Frame = -3
Query: 262 LSNLEEHCEKVLVGSEIDYRRF----IAQRSENRVNYLWPIERETINKFEKKW 310
+S+++E E + +E ++ R + ++ E RV LW + I +FEK+W
Sbjct: 207 VSHVQEELEGGMAETETEWWRIEEVGL*RKGEERVG-LW*VSGSEIGRFEKRW 52
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 26.9 bits (58), Expect = 8.1
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +3
Query: 89 PKGLYIYGNVGSGKTML 105
PKG+ +YG G+GKT+L
Sbjct: 429 PKGVLLYGPPGTGKTLL 479
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 26.9 bits (58), Expect = 8.1
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +3
Query: 89 PKGLYIYGNVGSGKTML 105
P+G+ +YG G+GKTML
Sbjct: 600 PRGVLLYGPPGTGKTML 650
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/45 (24%), Positives = 23/45 (50%)
Frame = +2
Query: 11 WEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRP 55
W +K++ +R+ + + +K++ K WKR + T + S P
Sbjct: 230 WRRKKKRRKRKRRVKKKKKKKKKKRKWKRKRRRKTMNRSRSFSNP 364
>TC14544
Length = 1160
Score = 26.9 bits (58), Expect = 8.1
Identities = 9/22 (40%), Positives = 12/22 (53%)
Frame = +2
Query: 44 ITERWTNSRKRPENVDPGVGKW 65
I + WT S E +DP G+W
Sbjct: 173 IADNWTRSAHWAETLDPATGRW 238
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,925,601
Number of Sequences: 28460
Number of extensions: 117788
Number of successful extensions: 751
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 745
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 751
length of query: 520
length of database: 4,897,600
effective HSP length: 94
effective length of query: 426
effective length of database: 2,222,360
effective search space: 946725360
effective search space used: 946725360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145331.1


