

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145330.19 - phase: 0 /pseudo
(469 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
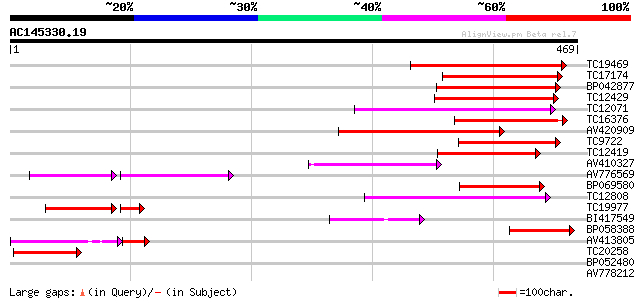
Score E
Sequences producing significant alignments: (bits) Value
TC19469 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, par... 198 1e-51
TC17174 weakly similar to UP|Q940N9 (Q940N9) AT4g21910/T8O5_120,... 179 7e-46
BP042877 129 1e-30
TC12429 similar to UP|Q9FKQ1 (Q9FKQ1) Emb|CAB89401.1 (AT5g65380/... 124 3e-29
TC12071 weakly similar to GB|AAP31968.1|30387605|BT006624 At2g04... 124 4e-29
TC16376 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (17%) 116 7e-27
AV420909 113 6e-26
TC9722 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (16%) 106 9e-24
TC12419 89 2e-18
AV410327 88 3e-18
AV776569 50 1e-13
BP069580 70 1e-12
TC12808 weakly similar to GB|CAA16564.1|2827556|ATT12H17 predict... 64 4e-11
TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%) 43 5e-09
BI417549 50 8e-07
BP058388 47 5e-06
AV413805 35 7e-06
TC20258 similar to UP|Q9FR32 (Q9FR32) Ripening regulated protein... 46 1e-05
BP052480 39 0.001
AV778212 34 0.045
>TC19469 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (26%)
Length = 501
Score = 198 bits (504), Expect = 1e-51
Identities = 91/129 (70%), Positives = 112/129 (86%)
Frame = +1
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
LF RERLAY+FT + EVA AVG+LSPLLSISILLNSV PVLSGV++GAGW S VAYVNIG
Sbjct: 1 LFLRERLAYVFTLDSEVAEAVGDLSPLLSISILLNSVPPVLSGVSVGAGWPSVVAYVNIG 180
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+IGIPVG+VLGN+IH Q KG+W+GMLFGT +QT++L+ IT KT+W+ QV +AR RVN
Sbjct: 181 CYYLIGIPVGVVLGNLIHLQEKGVWIGMLFGTFVQTVMLITITLKTDWENQVVIARNRVN 360
Query: 452 RWSKVESTD 460
+W+ E+T+
Sbjct: 361 KWAVKENTE 387
>TC17174 weakly similar to UP|Q940N9 (Q940N9) AT4g21910/T8O5_120, partial
(18%)
Length = 723
Score = 179 bits (455), Expect = 7e-46
Identities = 81/99 (81%), Positives = 95/99 (95%)
Frame = +1
Query: 359 LSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMG 418
L++SILL SV PVLSGVA+GAGW+STVAYVN+GCYYIIG+PVGIVLGN+ HWQVKGIW+G
Sbjct: 1 LAVSILLYSVPPVLSGVALGAGWRSTVAYVNLGCYYIIGLPVGIVLGNVFHWQVKGIWIG 180
Query: 419 MLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWSKVE 457
MLFGTL+QTIVL+IITY+T+W+EQVT+AR RVNRWSKVE
Sbjct: 181 MLFGTLVQTIVLVIITYRTDWEEQVTIARNRVNRWSKVE 297
>BP042877
Length = 489
Score = 129 bits (323), Expect = 1e-30
Identities = 55/102 (53%), Positives = 81/102 (78%)
Frame = -1
Query: 354 ELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVK 413
+L PLL++SILL+ +QPVLSGVA+G GW + VAYVN+GCYY +GIP+G VLG + +
Sbjct: 486 DLCPLLALSILLHGIQPVLSGVAVGCGWPAFVAYVNVGCYYGVGIPLGSVLGFYFQFGAQ 307
Query: 414 GIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWSK 455
GIW+GML GT + TI+L+ +T++T+WD++V A +R+N+W +
Sbjct: 306 GIWLGMLGGTTMPTIILMWVTFRTDWDQEVEEAAQRLNQWEE 181
>TC12429 similar to UP|Q9FKQ1 (Q9FKQ1) Emb|CAB89401.1 (AT5g65380/MNA5_11),
partial (22%)
Length = 523
Score = 124 bits (312), Expect = 3e-29
Identities = 54/104 (51%), Positives = 80/104 (76%), Gaps = 1/104 (0%)
Frame = +1
Query: 352 VGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQ 411
V LS LL+ +ILLNS+QPVLSGVA+G+GWQS VAY+N+GCYY++G+P+G ++G + +
Sbjct: 13 VNNLSLLLAFTILLNSIQPVLSGVAVGSGWQSYVAYINLGCYYLVGVPLGFIMGWVFNQG 192
Query: 412 VKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVTVARKRVNRWS 454
V GIW GM+F GT +QT++L +IT + +WD + A+ + +WS
Sbjct: 193 VMGIWAGMIFGGTALQTLILGVITIRCDWDGEAEKAKLHLTKWS 324
>TC12071 weakly similar to GB|AAP31968.1|30387605|BT006624 At2g04100
{Arabidopsis thaliana;}, partial (32%)
Length = 764
Score = 124 bits (310), Expect = 4e-29
Identities = 66/166 (39%), Positives = 98/166 (58%)
Frame = +3
Query: 286 SLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSN 345
S G AA S RVSNELG G K A+ +I ++ + L R L + F+++
Sbjct: 21 SYGVGAAVSTRVSNELGAGKPKEARDAIFAVIILATLDAIILSSALFCCRHVLGFAFSND 200
Query: 346 KEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLG 405
EV +V ++ PLL +S+ ++S VLSGVA GAGWQ A N+ YY IGIPV ++LG
Sbjct: 201 LEVVHSVAKIVPLLCLSVCVDSFLGVLSGVARGAGWQKIGAVTNLLAYYAIGIPVALLLG 380
Query: 406 NIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
++ KG+W+G+L G+ QT+VL ++T T W++Q +A R +
Sbjct: 381 FVLKLNAKGLWIGILTGSTTQTLVLALLTAFTKWEKQAPLATVRTS 518
>TC16376 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (17%)
Length = 550
Score = 116 bits (291), Expect = 7e-27
Identities = 52/93 (55%), Positives = 72/93 (76%)
Frame = +2
Query: 369 QPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTI 428
QPVLSGVA+G GWQ+ VAYVN+GCYY IGIP G VLG + GIW+GML GT++QTI
Sbjct: 2 QPVLSGVAVGCGWQTFVAYVNVGCYYGIGIPFGAVLGFYFKFGAMGIWLGMLAGTVLQTI 181
Query: 429 VLLIITYKTNWDEQVTVARKRVNRWSKVESTDQ 461
+L+ +T++T+W++++ A KR+N+W EST +
Sbjct: 182 ILVWVTFRTDWNKEIEEATKRLNKWE--ESTTE 274
>AV420909
Length = 421
Score = 113 bits (283), Expect = 6e-26
Identities = 57/137 (41%), Positives = 86/137 (62%)
Frame = +3
Query: 273 SICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFL 332
S+CLN ++ AAAS RVSNELG G+ K AK ++ V V+ + + FF+
Sbjct: 3 SVCLNTTTLHYFVTYAVGAAASTRVSNELGAGNPKTAKGAVRVVVIVGIVEAIIVSAFFI 182
Query: 333 FFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGC 392
FR L Y ++++KEV V +++PLL S++ +S+ LSGVA G G+Q AY N+G
Sbjct: 183 CFRNVLGYAYSNDKEVVDYVADMAPLLCGSVIGDSLIGALSGVARGGGFQQIGAYANLGA 362
Query: 393 YYIIGIPVGIVLGNIIH 409
YY++GIP+G++LG +H
Sbjct: 363 YYVVGIPIGLLLGFHLH 413
>TC9722 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (16%)
Length = 571
Score = 106 bits (264), Expect = 9e-24
Identities = 46/84 (54%), Positives = 66/84 (77%)
Frame = +3
Query: 372 LSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLL 431
LSGVA+G G Q+ VAYVN+GCYY +GIP+G VLG + KGIW+GML GT++QTI+L+
Sbjct: 3 LSGVAVGCGCQTFVAYVNVGCYYGVGIPLGAVLGFYFKFGAKGIWLGMLGGTVMQTIILM 182
Query: 432 IITYKTNWDEQVTVARKRVNRWSK 455
+T++T+W ++V A KR+N+W +
Sbjct: 183 WVTFRTDWIKEVEEASKRLNKWEE 254
>TC12419
Length = 667
Score = 89.0 bits (219), Expect = 2e-18
Identities = 40/85 (47%), Positives = 59/85 (69%)
Frame = +3
Query: 355 LSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKG 414
++PLL+ISILL+S+Q VLSGVA G GWQ AYVN+ +Y+IG+P+ +L Q KG
Sbjct: 15 VAPLLAISILLDSIQGVLSGVARGCGWQHLAAYVNLATFYLIGLPISCLLAFKTSLQYKG 194
Query: 415 IWMGMLFGTLIQTIVLLIITYKTNW 439
+W+G++ G + Q LL++T + W
Sbjct: 195 LWIGLICGMVCQAGTLLLLTTRAKW 269
>AV410327
Length = 424
Score = 87.8 bits (216), Expect = 3e-18
Identities = 51/110 (46%), Positives = 66/110 (59%)
Frame = +2
Query: 248 WFLIFSIQRPLACCQTFPFSRRNVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAK 307
W+ F I LA P + +SIC I+GW MIS+GF AAASVRVSNELG + K
Sbjct: 101 WY--FQILVLLAGLLPHPELALDSLSICTTISGWVFMISVGFNAAASVRVSNELGARNPK 274
Query: 308 AAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSP 357
+A FS+VV L S + L L R+ ++Y+FT +EVAAAV +L P
Sbjct: 275 SASFSVVVVTLISFIMSVIAALVVLALRDIISYVFTGGEEVAAAVSDLCP 424
>AV776569
Length = 633
Score = 50.1 bits (118), Expect(2) = 1e-13
Identities = 22/72 (30%), Positives = 43/72 (59%)
Frame = +2
Query: 17 ENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVF 76
+ +++ LSL +R E+K +W +A PAI T + + + FVGH+G +LAA ++
Sbjct: 17 DEDEKPLSLVRRFGIESKKLWKIAGPAILTMLCQYSLGAFTLTFVGHVGELDLAAVSVEN 196
Query: 77 TVLIRFANGILM 88
+ + F+ G+++
Sbjct: 197SCIAGFSFGVML 232
Score = 42.7 bits (99), Expect(2) = 1e-13
Identities = 26/94 (27%), Positives = 44/94 (46%)
Frame = +1
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 151
VG G+C+ + LWTSI C + G + +KIM I C C+CL +
Sbjct: 229 VGYGECLGDSLWTSIRCGSEQDVGGVHAKIMGHLIYYCLDFGPCICLVTSHLEGHWTNY* 408
Query: 152 HIRSCRKHFSLVNSYYVCFYCLIHLPNIPSITKQ 185
++RSC K + + VC + + ++T++
Sbjct: 409 NLRSCWKICFVDVTTVVCICFQLSNAEVLTVTEE 510
>BP069580
Length = 483
Score = 69.7 bits (169), Expect = 1e-12
Identities = 27/70 (38%), Positives = 48/70 (68%)
Frame = -1
Query: 373 SGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLI 432
SG A G G Q A++N+G YY++GIP+ +VL ++H KG+W+G++ ++Q L+I
Sbjct: 264 SGTARGCGLQKIGAFINLGSYYLVGIPLAMVLAFVLHIGGKGLWLGIICALIVQVFSLMI 85
Query: 433 ITYKTNWDEQ 442
IT + +W+++
Sbjct: 84 ITLRIDWEKE 55
>TC12808 weakly similar to GB|CAA16564.1|2827556|ATT12H17 predicted protein
{Arabidopsis thaliana;}, partial (32%)
Length = 624
Score = 64.3 bits (155), Expect = 4e-11
Identities = 39/154 (25%), Positives = 76/154 (49%)
Frame = +1
Query: 294 SVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVG 353
S RVSNELG A A S V++ L +G L + R +F+ ++ + V
Sbjct: 4 STRVSNELGANQAGRAYRSACVSLALGLILGCIGSLVMVAARGIWGPLFSHDRGIINGVK 183
Query: 354 ELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVK 413
+ L+++ + N V G+ G Y N+G +Y + +P+G+V ++ +
Sbjct: 184 KTMLLMALVEVFNFPLAVCGGIVRGTARPWLGMYANLGGFYFLALPLGVVFAFKLNLGLF 363
Query: 414 GIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVAR 447
G+++G++ G + +L+I + NW+E+ + A+
Sbjct: 364 GLFIGLITGIVTCLSLLVIFIARINWEEEASKAQ 465
>TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%)
Length = 452
Score = 43.1 bits (100), Expect(2) = 5e-09
Identities = 21/59 (35%), Positives = 36/59 (60%)
Frame = +3
Query: 30 WNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
W E KL++ +A PA+ + + + +Q F GH+G+ ELAA +L T + FA G+++
Sbjct: 216 WIELKLLFYLAVPAVIVYLINYVMSMSTQIFSGHLGNLELAAASLGNTGIQVFAYGLML 392
Score = 33.9 bits (76), Expect(2) = 5e-09
Identities = 11/20 (55%), Positives = 16/20 (80%)
Frame = +2
Query: 92 VGNGDCVVNPLWTSIWCKRI 111
V +G+C N +WTSIWCK++
Sbjct: 389 VRHGECSGNTMWTSIWCKKV 448
>BI417549
Length = 404
Score = 50.1 bits (118), Expect = 8e-07
Identities = 34/81 (41%), Positives = 48/81 (58%), Gaps = 2/81 (2%)
Frame = +1
Query: 265 PFSRRNVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLA-I 323
P +++SICL I+ I GF AAS RV+NELG G+ KAA+ +VV++ SLA I
Sbjct: 166 PELETSILSICLTISTLHFTIPYGFGVAASTRVANELGAGNPKAAR--VVVSIAMSLAFI 339
Query: 324 GSFLFLFFLF-FRERLAYIFT 343
+ + LF FR L Y ++
Sbjct: 340 EAVIVSTALFGFRHILGYAYS 402
>BP058388
Length = 352
Score = 47.4 bits (111), Expect = 5e-06
Identities = 19/55 (34%), Positives = 37/55 (66%), Gaps = 1/55 (1%)
Frame = -1
Query: 414 GIWMGMLFGTLIQTIVLLIITYKTNWDE-QVTVARKRVNRWSKVESTDQETKTKL 467
G+W+GMLFGT +QT+++ IT + + V + R RVN+W+ +E+ + +++ +
Sbjct: 322 GVWIGMLFGTFVQTVIVTTITTEN*XGKITVVITRNRVNKWAVMENIEPNSRSDI 158
>AV413805
Length = 418
Score = 34.7 bits (78), Expect(2) = 7e-06
Identities = 26/94 (27%), Positives = 48/94 (50%), Gaps = 1/94 (1%)
Frame = +3
Query: 1 MERDLKQNLLLKKPEEENEQEELSLGKRV-WNETKLMWVVAAPAIFTRFSTFGIQIISQA 59
M +DL LL K ++++E ++ + + E K + +AAP + S + +Q++S
Sbjct: 81 MAQDLAAPLLTKSGDKDDEVGVVTSSESTFYQELKKVSFMAAPMVAVTVSQYLLQVVSLM 260
Query: 60 FVGHIGSRELAAFALVFTVLIRFANGILMKIVVG 93
VGH EL +F+ V + I FA +++G
Sbjct: 261 MVGHFS--ELVSFSGV-AIAISFAEVTGFSVLLG 353
Score = 31.6 bits (70), Expect(2) = 7e-06
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +2
Query: 94 NGDCVVNPLWTSIWCKRIWNDG 115
N C+ N +W + WC+R+W G
Sbjct: 353 NVRCIGNFMWPNFWCRRVWKAG 418
>TC20258 similar to UP|Q9FR32 (Q9FR32) Ripening regulated protein DDTFR18,
partial (8%)
Length = 303
Score = 46.2 bits (108), Expect = 1e-05
Identities = 19/56 (33%), Positives = 37/56 (65%)
Frame = +2
Query: 4 DLKQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQA 59
++K LL P + ++E L +++W E+K +W +A PAIF+R +++ + +I+QA
Sbjct: 134 EIKAPLLEVHPLPDAAEQEQDLKRKIWVESKKLWHIAGPAIFSRIASYSMLVITQA 301
>BP052480
Length = 435
Score = 39.3 bits (90), Expect = 0.001
Identities = 14/36 (38%), Positives = 27/36 (74%)
Frame = -2
Query: 418 GMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRW 453
G++ G ++ T+VLL+I Y+T+W+++V +RV +W
Sbjct: 434 GIICGIVLPTLVLLLILYQTHWNQEVEDTSERVRKW 327
>AV778212
Length = 557
Score = 34.3 bits (77), Expect = 0.045
Identities = 19/59 (32%), Positives = 31/59 (52%)
Frame = +1
Query: 30 WNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
W E L++ +A PAI + ++AF GH+G+ LAA L + FA G+++
Sbjct: 145 WIELNLLFPLAGPAILVYLINNFMSTATRAFAGHLGNLALAAANLGNDGIQLFAYGLML 321
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.329 0.141 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,869,292
Number of Sequences: 28460
Number of extensions: 216001
Number of successful extensions: 1884
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 1850
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1877
length of query: 469
length of database: 4,897,600
effective HSP length: 94
effective length of query: 375
effective length of database: 2,222,360
effective search space: 833385000
effective search space used: 833385000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145330.19


