

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145329.13 - phase: 2 /pseudo
(528 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
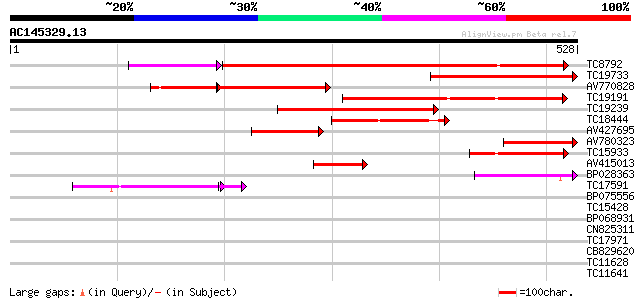
Score E
Sequences producing significant alignments: (bits) Value
TC8792 similar to GB|AAM00218.1|19879972|AF362990 beta-D-xylosid... 439 e-134
TC19733 weakly similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11,... 248 2e-66
AV770828 167 2e-59
TC19191 similar to UP|BAC98299 (BAC98299) LEXYL2 protein (Fragme... 199 7e-52
TC19239 homologue to UP|Q9ZU04 (Q9ZU04) Beta-glucosidase (Fragm... 189 1e-48
TC18444 similar to UP|Q8W011 (Q8W011) Beta-D-xylosidase, partial... 123 8e-29
AV427695 120 4e-28
AV780323 112 1e-25
TC15933 weakly similar to UP|BAC98299 (BAC98299) LEXYL2 protein ... 88 3e-18
AV415013 69 1e-12
BP028363 63 1e-10
TC17591 similar to UP|Q9FLG1 (Q9FLG1) Beta-xylosidase, partial (... 42 2e-05
BP075556 38 0.004
TC15428 36 0.014
BP068931 36 0.018
CN825311 32 0.20
TC17971 32 0.26
CB829620 31 0.44
TC11628 similar to PIR|A86350|A86350 F8K7.10 protein - Arabidops... 31 0.57
TC11641 similar to GB|BAC45220.1|27261107|AP005292 heat shock tr... 30 0.75
>TC8792 similar to GB|AAM00218.1|19879972|AF362990 beta-D-xylosidase
{Prunus persica;} , partial (90%)
Length = 1636
Score = 439 bits (1129), Expect(2) = e-134
Identities = 207/323 (64%), Positives = 255/323 (78%), Gaps = 1/323 (0%)
Frame = +3
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RY KTIHQ GC +V+CR + FG A AR +DAT++V+GLDQSIEAE DR
Sbjct: 351 PLQGIARYVKTIHQAGCRDVSCRGTELFGNAEIVARQSDATVVVVGLDQSIEAEFRDRVG 530
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPGHQQ+LVS+VA A++GP +LV+MSGGPVD+ FAK+DPK++ ILW GYPGQAGG AI
Sbjct: 531 LLLPGHQQELVSRVARAARGPVVLVIMSGGPVDVQFAKDDPKISAILWVGYPGQAGGTAI 710
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVYPFGH 377
AD++FG +PGG+LP TWYPQ Y+ L MTNM MRP+ GYPGRTYRFYKGPVV+PFGH
Sbjct: 711 ADVIFGRTNPGGRLPNTWYPQSYVAKLPMTNMDMRPNPASGYPGRTYRFYKGPVVFPFGH 890
Query: 378 GLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
GL+Y+ F H L+ AP VSVP+ + N+ +S+KA++V+HA+CG L + HVDVKN G
Sbjct: 891 GLSYSRFTHTLAVAPEQVSVPIASLQAFRNSTMSSKAVKVSHAKCGALKMGFHVDVKNEG 1070
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
S DGTHTLL+FS PP G W K LV+F K +VPA +KQRV V +HVCK LSVVD++G
Sbjct: 1071SMDGTHTLLIFSTPPPG--KWSASKQLVSFHKTYVPAGSKQRVNVGVHVCKHLSVVDENG 1244
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQ 520
IRRIPMG+H LHIGDVKH++S+Q
Sbjct: 1245IRRIPMGDHELHIGDVKHTISVQ 1313
Score = 57.0 bits (136), Expect(2) = e-134
Identities = 32/87 (36%), Positives = 47/87 (53%)
Frame = +2
Query: 111 CFGEYIEGANEIRDV*WRAISSSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHWSYFA 170
C G+ +EI V RAI S+ ++ P RC+ R P*S * + +++K A
Sbjct: 65 CLGQPYYCPDEIGHVRRRAIGPSIRELGPERCLHPGP*RTCP*SS*AGHSSARKQRKCPA 244
Query: 171 SLPTTPPNCGCHWAQFRCHSYHDWKLC 197
+P + P+C HW FRC+ Y+D KLC
Sbjct: 245 IIPYSTPHCRSHWT*FRCYCYNDRKLC 325
>TC19733 weakly similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, partial
(15%)
Length = 598
Score = 248 bits (633), Expect = 2e-66
Identities = 120/136 (88%), Positives = 130/136 (95%)
Frame = +2
Query: 393 TVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPP 452
TVVSVPV GHRH N+TNISNKAIRVTHARCGKLSIALHV+VKNVGSRDGTHTLLVFSAPP
Sbjct: 2 TVVSVPVDGHRHANSTNISNKAIRVTHARCGKLSIALHVNVKNVGSRDGTHTLLVFSAPP 181
Query: 453 NGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGD 512
+GG HW QK LVAFEKVHVPAK++QRV++NIHVCKLLSVVD+SGIRRIPMGEHSLHIGD
Sbjct: 182 DGGAHWAVQKQLVAFEKVHVPAKSQQRVQINIHVCKLLSVVDRSGIRRIPMGEHSLHIGD 361
Query: 513 VKHSVSLQAEALGIIK 528
VKHSVSLQA+A+GIIK
Sbjct: 362 VKHSVSLQADAIGIIK 409
>AV770828
Length = 527
Score = 167 bits (423), Expect(2) = 2e-59
Identities = 84/105 (80%), Positives = 93/105 (88%)
Frame = +1
Query: 194 WKLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAET 253
++ P +GIG+YAKTIH+ GC+NVA DDKQFG AL+AAR ADAT+LV+GLDQSIEAET
Sbjct: 211 YRYTSP*EGIGKYAKTIHELGCANVAGTDDKQFGSALNAARQADATVLVMGLDQSIEAET 390
Query: 254 VDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKND 298
VDR LLLPG QQDLVSKVAAASKGPTILVLMSGGPVDITFAKND
Sbjct: 391 VDRAGLLLPGRQQDLVSKVAAASKGPTILVLMSGGPVDITFAKND 525
Score = 79.3 bits (194), Expect(2) = 2e-59
Identities = 39/66 (59%), Positives = 48/66 (72%)
Frame = +3
Query: 132 SSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHWSYFASLPTTPPNCGCHWAQFRCHSY 191
SS+W RCV +S RASP*SC*TR+CA ++ WS FASL +TP + G HWAQF CH +
Sbjct: 6 SSIWHWSK-RCVHPTSPRASP*SC*TRDCAPKEQWSLFASLHSTPSHPGYHWAQF*CHCH 182
Query: 192 HDWKLC 197
+D KLC
Sbjct: 183 NDRKLC 200
>TC19191 similar to UP|BAC98299 (BAC98299) LEXYL2 protein (Fragment),
partial (31%)
Length = 770
Score = 199 bits (507), Expect = 7e-52
Identities = 97/210 (46%), Positives = 137/210 (65%), Gaps = 1/210 (0%)
Frame = +2
Query: 311 GQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKG 369
G+AGGAAIAD++FG +P G+LP+TWYPQ Y+ + MTNM MRP GYPGRTYRFYKG
Sbjct: 2 GEAGGAAIADVIFGFHNPSGRLPMTWYPQSYVDKVPMTNMNMRPDPTTGYPGRTYRFYKG 181
Query: 370 PVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIAL 429
V+ FG GL+Y+ H+L AP +VS+P+ ++ + K++ V C L +
Sbjct: 182 ETVFSFGDGLSYSTIEHKLVKAPQLVSIPLAEDHVCHS--LKCKSLDVAGEHCENLEFEI 355
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
H+ +KN G HT+ +FS+PP H P K LV FEKVH+ T+ V+ + VCK
Sbjct: 356 HLSIKNKGRMSSDHTVFLFSSPP--AVHNAPHKHLVGFEKVHLEGNTEALVKFKVDVCKD 529
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
LSVVD+ G R++ +G+H LH+GD+KH++ +
Sbjct: 530 LSVVDELGNRKVALGQHLLHVGDLKHALGV 619
>TC19239 homologue to UP|Q9ZU04 (Q9ZU04) Beta-glucosidase (Fragment) ,
partial (69%)
Length = 453
Score = 189 bits (480), Expect = 1e-48
Identities = 86/151 (56%), Positives = 118/151 (77%), Gaps = 1/151 (0%)
Frame = +1
Query: 250 EAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGY 309
+AE++DR +++LPG QQ LVS+VA AS+GP ILV+MSGG +D++FAK + K+ ILW GY
Sbjct: 1 QAESLDRVNIVLPGQQQLLVSEVANASRGPVILVIMSGGGMDVSFAKANDKITSILWVGY 180
Query: 310 PGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYK 368
PG+AGGAAIAD++FG +P G+LP+TWYP+ Y++N+ MTNM MR GYPGRTYRFYK
Sbjct: 181 PGEAGGAAIADVIFGFYNPSGRLPMTWYPESYVENVPMTNMNMRADPSTGYPGRTYRFYK 360
Query: 369 GPVVYPFGHGLTYTHFVHELSSAPTVVSVPV 399
G V+ FG G+ Y++ H++ AP +V VP+
Sbjct: 361 GETVFSFGDGIGYSNVEHKIVKAPQLVFVPL 453
>TC18444 similar to UP|Q8W011 (Q8W011) Beta-D-xylosidase, partial (12%)
Length = 387
Score = 123 bits (308), Expect = 8e-29
Identities = 56/111 (50%), Positives = 78/111 (69%), Gaps = 1/111 (0%)
Frame = +2
Query: 300 KVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIG 358
K+ GILWAGYPG+ GG A+A ++FG +PGG+LP+TWYP++Y+K + MT+M MR G
Sbjct: 2 KIGGILWAGYPGELGGIALAQVIFGDHNPGGRLPITWYPKDYIK-VPMTDMRMRADPATG 178
Query: 359 YPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTN 409
YPGRTYRFYKGP +Y FG+GL+Y+ + +E S H + H N ++
Sbjct: 179 YPGRTYRFYKGPKLYEFGYGLSYSKYSYEFISV-------THNNLHFNQSS 310
>AV427695
Length = 202
Score = 120 bits (302), Expect = 4e-28
Identities = 60/67 (89%), Positives = 64/67 (94%)
Frame = +2
Query: 226 FGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLM 285
FGPALDAARHADAT+LV+GLDQSIEAET DR LLLPGHQQDLVSKVAAAS+GPTILVLM
Sbjct: 2 FGPALDAARHADATVLVMGLDQSIEAETRDRVGLLLPGHQQDLVSKVAAASRGPTILVLM 181
Query: 286 SGGPVDI 292
SGGP+DI
Sbjct: 182 SGGPLDI 202
>AV780323
Length = 544
Score = 112 bits (280), Expect = 1e-25
Identities = 52/68 (76%), Positives = 62/68 (90%)
Frame = -3
Query: 461 QKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
+K LVAFEKVH+PAK ++RVRVNIHVCKLLSVVD+SGIRRIPMGEH IGD++HS+SLQ
Sbjct: 542 RKQLVAFEKVHIPAKAQRRVRVNIHVCKLLSVVDRSGIRRIPMGEHRFQIGDIEHSLSLQ 363
Query: 521 AEALGIIK 528
A+ LG+IK
Sbjct: 362 AKTLGVIK 339
>TC15933 weakly similar to UP|BAC98299 (BAC98299) LEXYL2 protein (Fragment),
partial (14%)
Length = 453
Score = 88.2 bits (217), Expect = 3e-18
Identities = 41/92 (44%), Positives = 64/92 (69%)
Frame = +1
Query: 429 LHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCK 488
+H+ VKN+G +HT+L+F PP H PQK L+ EKVH+ K++ +VR + VCK
Sbjct: 1 IHLRVKNMGKMTSSHTVLLFFTPP--AVHNAPQKHLLGIEKVHLAGKSEAQVRFMVDVCK 174
Query: 489 LLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
LSVVD+ RR+P+G+H LH+G++K+ +S++
Sbjct: 175 DLSVVDELAHRRVPLGQHLLHVGNLKYPLSVR 270
>AV415013
Length = 154
Score = 69.3 bits (168), Expect = 1e-12
Identities = 28/50 (56%), Positives = 38/50 (76%)
Frame = +2
Query: 284 LMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLP 333
+M+ GP+DI+F K+ + GILW GYPGQAGG AIA ++FG +PGG+ P
Sbjct: 2 IMAAGPIDISFTKSIRNIGGILWVGYPGQAGGDAIAQVIFGEYNPGGRSP 151
>BP028363
Length = 508
Score = 62.8 bits (151), Expect = 1e-10
Identities = 35/101 (34%), Positives = 54/101 (52%), Gaps = 6/101 (5%)
Frame = -3
Query: 434 KNVGSRDGTHTLLVFSAPPNGGN-HWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSV 492
KN G +G+H +LVF PP P K L+ FE+ HV + V + I +C+LLS
Sbjct: 506 KNTGPYNGSHVVLVFWEPPTSELVTGAPIKQLIGFERAHVMVGMTEFVTLEIDICQLLSN 327
Query: 493 VDKSGIRRIPMGEHSLHIG-----DVKHSVSLQAEALGIIK 528
VD G R++ +G+H+L +G V+H + + G+ K
Sbjct: 326 VDSDGKRKLIIGKHTLLVGSSSETQVRHQIDISLSGNGMEK 204
>TC17591 similar to UP|Q9FLG1 (Q9FLG1) Beta-xylosidase, partial (24%)
Length = 582
Score = 41.6 bits (96), Expect(2) = 2e-05
Identities = 45/156 (28%), Positives = 65/156 (40%), Gaps = 13/156 (8%)
Frame = +2
Query: 59 WVHCI*L*LCGSALQ*PTLYINAGRSCCRCHQSSA------------HAGCGEKRFVN*S 106
W++ *L*L SA Q L+ ++ RSC + H S GC + F *S
Sbjct: 38 WIYSF*L*LG*SAFQRSALHQDS*RSCSQIHSSRVGFGLWKLSWQIYRRGCQARAF-G*S 214
Query: 107 RCE*CFGEYIEGANEIRDV*WRAISSSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHW 166
C+ + + +*W + ++LW C+ P SC R+C + K
Sbjct: 215 IH*QCYLQQFRHIDATWFL*W*SKHATLWGPWSKGCMHIREPGTRPGSCKARDCVA*K*S 394
Query: 167 SYFASLPTTPPNCGCHWAQFRCHSYHDWKLCR-PLQ 201
AS G + Q +C+ HDWKL R PLQ
Sbjct: 395 RITASKCQIH*IIGSNRPQCKCY*GHDWKLRRHPLQ 502
Score = 23.5 bits (49), Expect(2) = 2e-05
Identities = 10/26 (38%), Positives = 13/26 (49%)
Frame = +3
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVAC 220
K PLQG+ T + GC +V C
Sbjct: 501 KYISPLQGLTALVPTSYVPGCPDVHC 578
>BP075556
Length = 469
Score = 38.1 bits (87), Expect = 0.004
Identities = 25/82 (30%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Frame = -3
Query: 434 KNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRV-RVNIHVCKLLSV 492
+N G HTL +FS+PP P +L + + K + + + + VCK LSV
Sbjct: 443 QNQGRMSSDHTLFLFSSPPPVPTP--PSSTLTGI*ESSLGRKYRTPLAKFKVDVCKDLSV 270
Query: 493 VDKSGIRRIPMGEHSLHIGDVK 514
VD+ G ++ +G+H G V+
Sbjct: 269 VDELGHSQVALGQHLASCG*VE 204
>TC15428
Length = 629
Score = 36.2 bits (82), Expect = 0.014
Identities = 22/58 (37%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Frame = +1
Query: 422 CGKLSIALHVDVKNVGSRDGTHTLLVFSAP--PNGGNHWVPQKSLVAFEKVHVPAKTK 477
C +S+++ + VKN GS G H +L+F P GN P K LV F V + A+ K
Sbjct: 133 CQSMSVSVTLGVKNDGSMAGKHPVLLFMRPGKQRNGN---PLKPLVGFPSVKLDARGK 297
>BP068931
Length = 471
Score = 35.8 bits (81), Expect = 0.018
Identities = 31/100 (31%), Positives = 47/100 (47%)
Frame = -1
Query: 281 ILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQE 340
++VL++G PV I PK+ ++ A PG G +AD+LFG GKL TW+ +
Sbjct: 414 VVVLITGRPVVIQ--PYLPKIDALVAAWLPGTEG-QGVADLLFGDYKFTGKLARTWFKR- 247
Query: 341 YLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHGLT 380
+ P +G + Y ++PFG GLT
Sbjct: 246 ---------VDQLPMNVG--DKHY-----DPLFPFGFGLT 175
>CN825311
Length = 522
Score = 32.3 bits (72), Expect = 0.20
Identities = 29/86 (33%), Positives = 37/86 (42%)
Frame = +1
Query: 1 CAGE*ARHRRHI*CTIQDVCEGRKSS*CHVFLQSSKWGPYMC*S*PLKENSSWSMGS*WV 60
C G A H+ +C RKS VFLQ +W +C* L +G * V
Sbjct: 184 CCGISAGFGGHLSTPFSRLCSTRKSKLLDVFLQ*GQWCSCLC**GSLGSG*K-QLGV*RV 360
Query: 61 HCI*L*LCGSALQ*PTLYINAGRSCC 86
H +*L* CG + * + R CC
Sbjct: 361 HYL*L*CCGYGV-*VSGVCKISRGCC 435
>TC17971
Length = 551
Score = 32.0 bits (71), Expect = 0.26
Identities = 29/86 (33%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Frame = +1
Query: 194 WKLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGP---ALDAARHADATILVIGLDQSIE 250
W C PL+G A+ +H Q CS V D + GP L A IL G + E
Sbjct: 229 WLSCLPLKGDLIEARVVHDQLCSMVERSDRELLGPNNQYLSKIVAVFAEILCAGNSLATE 408
Query: 251 AETVDRTSLLLPGHQQDLVSKVAAAS 276
+TV R LL QQ L A++
Sbjct: 409 -QTVSRMINLLRQLQQTLPPSTLAST 483
>CB829620
Length = 548
Score = 31.2 bits (69), Expect = 0.44
Identities = 18/61 (29%), Positives = 29/61 (47%)
Frame = -3
Query: 366 FYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKL 425
FYK +V+ G++Y HF+H S P + + + NI +R ++ R GK
Sbjct: 375 FYKSTLVHLNKRGISY*HFIHYTSQRPPITFHAISFLQQNFRGNI----VRCSNRRIGKC 208
Query: 426 S 426
S
Sbjct: 207 S 205
>TC11628 similar to PIR|A86350|A86350 F8K7.10 protein - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (7%)
Length = 548
Score = 30.8 bits (68), Expect = 0.57
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Frame = -3
Query: 171 SLPTTPPNCGCHWAQF---RCHSYHDWKLCRPLQGIGRYAKTIHQQGCSNV 218
S+P + P CG +A+F C YH L L+ +G Y + + +GC NV
Sbjct: 342 SIPLSKPQCGV*YARFCSNFCICYHF*VLSCRLRALG-YERYLKNKGCKNV 193
>TC11641 similar to GB|BAC45220.1|27261107|AP005292 heat shock transcription
factor-like protein {Oryza sativa (japonica
cultivar-group);}, partial (7%)
Length = 420
Score = 30.4 bits (67), Expect = 0.75
Identities = 23/86 (26%), Positives = 34/86 (38%)
Frame = +1
Query: 332 LPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSA 391
+P W P +Y + M + + P YP P P+ + Y H H+ +
Sbjct: 208 VPQQWVPMQY-PAMVMPHHMLPPPPQHYP---------PPPPPY---VPYHHHHHQFAP- 345
Query: 392 PTVVSVPVHGHRHGNNTNISNKAIRV 417
P H HG+N NI NK + V
Sbjct: 346 ------PHQQHHHGSNNNIENKTVWV 405
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.337 0.146 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,873,953
Number of Sequences: 28460
Number of extensions: 183294
Number of successful extensions: 1402
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1375
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1390
length of query: 528
length of database: 4,897,600
effective HSP length: 94
effective length of query: 434
effective length of database: 2,222,360
effective search space: 964504240
effective search space used: 964504240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145329.13


