

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
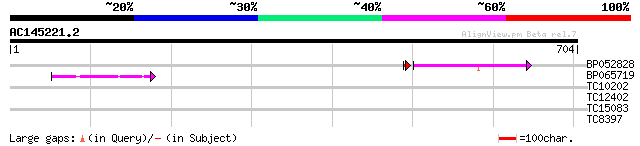
Query= AC145221.2 - phase: 0
(704 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP052828 106 1e-23
BP065719 44 7e-05
TC10202 similar to UP|AX28_SOYBN (P13089) Auxin-induced protein ... 28 3.8
TC12402 similar to UP|AAS46231 (AAS46231) Methionine sulfoxide r... 28 6.5
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 28 6.5
TC8397 similar to UP|Q9RV45 (Q9RV45) S-layer-like array-related ... 27 8.5
>BP052828
Length = 523
Score = 106 bits (264), Expect(2) = 1e-23
Identities = 54/149 (36%), Positives = 88/149 (58%), Gaps = 3/149 (2%)
Frame = +2
Query: 502 PVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDV 561
P IG ++DLG+ IN++P S+ + ++ T + +QLA++S ARP G+ EDV
Sbjct: 71 PCTIGDNKYENCMLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDV 250
Query: 562 LVKVDKFVFPVDFVVMDIE---EDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEIN 618
LV+V++ +FP DF ++D+E + P+ILGR FM A+ ID D+G M + D
Sbjct: 251 LVQVNELIFPADFYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAK 430
Query: 619 FNLHSAMKHSKDKGACFKVDAMDEVIMET 647
FN+ AMKH + + F + + E++ +T
Sbjct: 431 FNIDDAMKHPLEDHSVFHIGVVSELVDDT 517
Score = 21.2 bits (43), Expect(2) = 1e-23
Identities = 7/9 (77%), Positives = 9/9 (99%)
Frame = +1
Query: 489 LPKKEKDPG 497
LP+K+KDPG
Sbjct: 34 LPQKQKDPG 60
>BP065719
Length = 567
Score = 44.3 bits (103), Expect = 7e-05
Identities = 32/129 (24%), Positives = 58/129 (44%)
Frame = +3
Query: 53 FAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKAREWYLDQSPDVLKD 112
F+G E +H++ + A L +E+ + ++ F SL A W+ +P +
Sbjct: 198 FSGDSGESTVEHIARYQIEAGDLAINENLK----MKYFPSSLTKNAFTWFTTLAPRSVHT 365
Query: 113 WNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSLLRKCPKHGFDDATQ 172
W LE+ F E+FF + K +A + ES+ D R++ L +C H +
Sbjct: 366 WAQLERIFHEQFFRGECKVSXKD-LASVKRKPAESIDDYLNRFRMLKSRCFTH-VSEHEL 539
Query: 173 IHMFRGGLQ 181
+ + GGL+
Sbjct: 540 VVLXAGGLE 566
>TC10202 similar to UP|AX28_SOYBN (P13089) Auxin-induced protein AUX28,
partial (60%)
Length = 664
Score = 28.5 bits (62), Expect = 3.8
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = +3
Query: 266 KEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQ 296
KEQ + +R +Q C++C +H PPQ
Sbjct: 456 KEQRRSREREEQQP*CKLCESEHGWSTLPPQ 548
>TC12402 similar to UP|AAS46231 (AAS46231) Methionine sulfoxide reductase A,
partial (95%)
Length = 756
Score = 27.7 bits (60), Expect = 6.5
Identities = 19/54 (35%), Positives = 26/54 (47%)
Frame = +3
Query: 224 GTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQ 277
GTQ +SGI A LAQ +Q+E K + E+ K + E+ HQQ
Sbjct: 351 GTQYRSGIYYYNETQARLAQESKEAKQLELKGKVVTEILP-AKRFYRAEEYHQQ 509
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 27.7 bits (60), Expect = 6.5
Identities = 19/62 (30%), Positives = 30/62 (47%), Gaps = 6/62 (9%)
Frame = +1
Query: 373 DTLNQFMQLTMANQ------QNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLE 426
DT + ++ +AN Q N + + SDV K ++ + K KLPY KA + L
Sbjct: 952 DTTSTALEWVLANLDKYPRVQKNIVDEISDVMKGREDKEVKEEDLVKLPYLKAVILEGLR 1131
Query: 427 KH 428
+H
Sbjct: 1132 RH 1137
>TC8397 similar to UP|Q9RV45 (Q9RV45) S-layer-like array-related protein,
partial (5%)
Length = 1357
Score = 27.3 bits (59), Expect = 8.5
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 7/51 (13%)
Frame = -2
Query: 322 NQQRFPNNNNFQRG----NNFRQPWKSNAG---PSNQQPFNNNFQQYPPQQ 365
NQ++FP+N ++QR QPW ++ G PS + +F + P Q
Sbjct: 798 NQEQFPDNMDWQRK*EPLQLTVQPWLASQG*QHPSPEGSLQYHFASFSPHQ 646
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,796,391
Number of Sequences: 28460
Number of extensions: 139086
Number of successful extensions: 721
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 716
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 719
length of query: 704
length of database: 4,897,600
effective HSP length: 97
effective length of query: 607
effective length of database: 2,136,980
effective search space: 1297146860
effective search space used: 1297146860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC145221.2


