

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145219.5 + phase: 0
(130 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
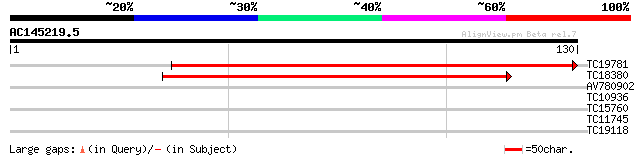
TC19781 similar to UP|AAQ73526 (AAQ73526) Early flowering 4, par... 150 7e-38
TC18380 81 7e-17
AV780902 26 2.5
TC10936 weakly similar to UP|Q9C887 (Q9C887) Myosin-like protein... 25 3.3
TC15760 similar to UP|Q9LW29 (Q9LW29) Transport inhibitor respon... 25 3.3
TC11745 25 3.3
TC19118 weakly similar to UP|Q84YE8 (Q84YE8) Expressed protein-l... 25 5.6
>TC19781 similar to UP|AAQ73526 (AAQ73526) Early flowering 4, partial (42%)
Length = 543
Score = 150 bits (379), Expect = 7e-38
Identities = 72/93 (77%), Positives = 81/93 (86%)
Frame = +1
Query: 38 DVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELN 97
+ +D DGD EAW T+NKSFRQVQSVLDRNR +IQQVNE QQSRMPDNMVKNV LIQELN
Sbjct: 19 EFDDTHDGDXEAWATMNKSFRQVQSVLDRNRLLIQQVNEXQQSRMPDNMVKNVSLIQELN 198
Query: 98 GNISKVASLYSDLNSDFTNICHQQQRSSHNRGK 130
GNISKV SLYSDLNS+F+N+C QQQR ++ RGK
Sbjct: 199 GNISKVVSLYSDLNSNFSNVCQQQQRFNNGRGK 297
>TC18380
Length = 664
Score = 80.9 bits (198), Expect = 7e-17
Identities = 34/80 (42%), Positives = 54/80 (67%)
Frame = +1
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+ ++ + D Q KS Q Q +L++NR +I ++N+N +S+MPDN+ +NVGLI+E
Sbjct: 88 FGELGNTSQVDSRVMQVFQKSLLQAQDILNQNRVLINEINQNHESKMPDNLSRNVGLIKE 267
Query: 96 LNGNISKVASLYSDLNSDFT 115
LN NI +V LY+D++S FT
Sbjct: 268 LNSNIRRVVDLYADISSSFT 327
>AV780902
Length = 506
Score = 25.8 bits (55), Expect = 2.5
Identities = 10/28 (35%), Positives = 15/28 (52%)
Frame = +1
Query: 20 KNQRRRRPSATPTNNDYSDVEDAEDGDP 47
+ ++R R S P DY ++D GDP
Sbjct: 313 QGKKRTRTSLAPCKRDYCLIDDRSCGDP 396
>TC10936 weakly similar to UP|Q9C887 (Q9C887) Myosin-like protein, partial
(14%)
Length = 586
Score = 25.4 bits (54), Expect = 3.3
Identities = 12/33 (36%), Positives = 15/33 (45%)
Frame = -2
Query: 67 NRAIIQQVNENQQSRMPDNMVKNVGLIQELNGN 99
N+ I Q N SR+PDNM +V N
Sbjct: 474 NKTYICQFRTNNISRIPDNMAMHVACTHRRTSN 376
>TC15760 similar to UP|Q9LW29 (Q9LW29) Transport inhibitor response-like
protein, partial (17%)
Length = 664
Score = 25.4 bits (54), Expect = 3.3
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -3
Query: 2 STLYSPSSAMEDTNHRSK 19
S +Y PS +ME TNH S+
Sbjct: 542 SNMYKPSKSMEITNHYSQ 489
>TC11745
Length = 440
Score = 25.4 bits (54), Expect = 3.3
Identities = 11/23 (47%), Positives = 13/23 (55%)
Frame = +2
Query: 7 PSSAMEDTNHRSKKNQRRRRPSA 29
P D N R++KN RRRP A
Sbjct: 245 PQRTDRDRNRRARKNVARRRPVA 313
>TC19118 weakly similar to UP|Q84YE8 (Q84YE8) Expressed protein-like
protein, partial (11%)
Length = 932
Score = 24.6 bits (52), Expect = 5.6
Identities = 11/42 (26%), Positives = 18/42 (42%)
Frame = +3
Query: 7 PSSAMEDTNHRSKKNQRRRRPSATPTNNDYSDVEDAEDGDPE 48
P+ A E+T ++ R TP+ N +DG+ E
Sbjct: 732 PTIAAEETRDEDGDDETNREEPQTPSKNICKSSNSLDDGEEE 857
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.123 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,826,055
Number of Sequences: 28460
Number of extensions: 17512
Number of successful extensions: 113
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 112
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 113
length of query: 130
length of database: 4,897,600
effective HSP length: 80
effective length of query: 50
effective length of database: 2,620,800
effective search space: 131040000
effective search space used: 131040000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC145219.5


