

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145218.1 - phase: 0
(63 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
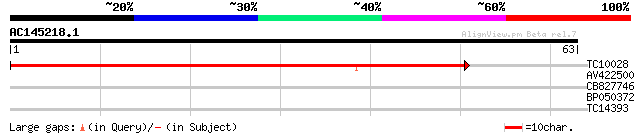
TC10028 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-lik... 55 3e-09
AV422500 33 0.014
CB827746 26 1.3
BP050372 24 6.6
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 23 8.6
>TC10028 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-like protein,
partial (16%)
Length = 874
Score = 54.7 bits (130), Expect = 3e-09
Identities = 28/52 (53%), Positives = 37/52 (70%), Gaps = 1/52 (1%)
Frame = +1
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARD-EKIFDLRDIMRSSS 51
MAS+EL SV +RSYA+E++NEEI TRL P M + A + DL+DI+R S
Sbjct: 472 MASMELGSVAIRSYAMELVNEEIQTRLTPLMAATAASANNVLDLQDILRRFS 627
>AV422500
Length = 446
Score = 32.7 bits (73), Expect = 0.014
Identities = 14/47 (29%), Positives = 27/47 (56%)
Frame = -3
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
+AS T +VR + + EE+ L+P M A +E++ DL++++
Sbjct: 408 LASHVFTPRSVRQFVTTTLEEEVRENLLPAMEVLAEEERVVDLQELL 268
>CB827746
Length = 550
Score = 26.2 bits (56), Expect = 1.3
Identities = 18/54 (33%), Positives = 26/54 (47%)
Frame = +3
Query: 3 SLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMRSSSKYMSK 56
S+ TS TVRS +L ++ I +PF SFA L I+ S S + +
Sbjct: 96 SILRTSSTVRSGSLHLLPPRISHTPLPFPLSFASKLGFIGLGLILLSVSNQLDR 257
>BP050372
Length = 332
Score = 23.9 bits (50), Expect = 6.6
Identities = 8/22 (36%), Positives = 15/22 (67%)
Frame = -3
Query: 26 RLIPFMFSFARDEKIFDLRDIM 47
+++PFMF + IFDL +++
Sbjct: 273 KILPFMFPYTSQLHIFDLFNVL 208
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 23.5 bits (49), Expect = 8.6
Identities = 8/15 (53%), Positives = 12/15 (79%)
Frame = -1
Query: 26 RLIPFMFSFARDEKI 40
RL+PF+F F R+ +I
Sbjct: 75 RLLPFLFGFEREREI 31
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.135 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 697,996
Number of Sequences: 28460
Number of extensions: 5169
Number of successful extensions: 42
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 42
length of query: 63
length of database: 4,897,600
effective HSP length: 39
effective length of query: 24
effective length of database: 3,787,660
effective search space: 90903840
effective search space used: 90903840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC145218.1


