

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
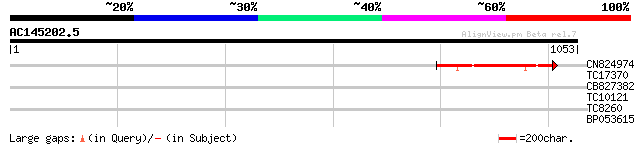
Query= AC145202.5 - phase: 0 /pseudo
(1053 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CN824974 219 2e-57
TC17370 similar to UP|Q8LGE7 (Q8LGE7) NADH:ubiquinone oxidoreduc... 30 2.0
CB827382 29 3.4
TC10121 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (S... 29 4.4
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 28 9.8
BP053615 28 9.8
>CN824974
Length = 723
Score = 219 bits (557), Expect = 2e-57
Identities = 129/240 (53%), Positives = 165/240 (68%), Gaps = 15/240 (6%)
Frame = +3
Query: 793 DGLVERINEQTFRIKELPIRMWTEDYKKFLEKITARG------LIESFRQKGDYEIVNFK 846
+G VE I+EQ+FRI ELPIR WT+DYK+FLE IT LIE FRQ GD +V+ +
Sbjct: 12 EGTVEEIDEQSFRITELPIRKWTQDYKQFLESITDGSPNVKDPLIEDFRQNGDDAMVDIE 191
Query: 847 VNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLK 906
+ +K E+IA I + E L KKF LT TISTSNM+LFDAEGKIKKY PEQILEEF LRL
Sbjct: 192 IRMKLEKIAMIMQ-EGLLKKFKLTTTISTSNMHLFDAEGKIKKYENPEQILEEFFPLRLD 368
Query: 907 YYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEIL--------KK 957
YY RRKQY+++NF +LL L K +FI +V+ + NR +A+LL+E+ KK
Sbjct: 369 YYERRKQYILENFERLLLVLDNKVRFILGVVSGEIVVSNRKKADLLMELQQKGFTPMPKK 548
Query: 958 GKSAEPHVAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKL 1017
GKSAEP VAGA D +S+EQE ++ + ++V +E A GDYEYL S+ TLES++KL
Sbjct: 549 GKSAEPQVAGANDENSEEQE--DENTREEAVRVEEAKRGDYEYLLSMSIGTLTLESVQKL 722
>TC17370 similar to UP|Q8LGE7 (Q8LGE7) NADH:ubiquinone oxidoreductase-like
protein (At5g18800), partial (88%)
Length = 566
Score = 30.0 bits (66), Expect = 2.0
Identities = 22/71 (30%), Positives = 35/71 (48%), Gaps = 4/71 (5%)
Frame = -1
Query: 904 RLKYYVRRKQYLVK----NFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIEILKKGK 959
RL+YY R +QY++ F + + KFI +V+ I+ LL+ IL+K K
Sbjct: 326 RLRYYFR*QQYIISFQRTLFLKCFLLFTTQIKFICMVVHAANVIIHLFRALLV*ILQKAK 147
Query: 960 SAEPHVAGAID 970
+VA I+
Sbjct: 146 DTASNVAAFIE 114
>CB827382
Length = 559
Score = 29.3 bits (64), Expect = 3.4
Identities = 19/35 (54%), Positives = 22/35 (62%)
Frame = +1
Query: 997 DYEYLSSLPFENFTLESLKKLEAELDEKENELKTL 1031
+YE+L LP T ESLKKL+A EKE EL L
Sbjct: 10 NYEHL--LPMCTLTFESLKKLQA---EKEKELDNL 99
>TC10121 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (Sucrose-UDP
glucosyltransferase 2) , partial (41%)
Length = 996
Score = 28.9 bits (63), Expect = 4.4
Identities = 21/51 (41%), Positives = 29/51 (56%), Gaps = 10/51 (19%)
Frame = +2
Query: 829 GLIESFRQKGDY-EIVNFKV-----NVKQ----EEIATIEKDEELRKKFNL 869
GL+ES+ + E+VN + +VK+ EEIA IEK EL K +NL
Sbjct: 704 GLVESYAKNNKLRELVNLVIVAGYIDVKKSRDREEIAEIEKMHELMKNYNL 856
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 27.7 bits (60), Expect = 9.8
Identities = 22/79 (27%), Positives = 39/79 (48%), Gaps = 12/79 (15%)
Frame = -2
Query: 64 WTGWK*TLFLRLMRFMFT*KVFHF*AM-----------KHLIM*SKSFIPRIFTRLI-VM 111
W GW+* + +RLM + + +V H+ + L++ + + R + R++ V
Sbjct: 320 WWGWR*WVLVRLMWWWWWWRVVHWVWLLVWWRWRVVHWVRLLVWWRW*VNRWWWRVVGVR 141
Query: 112 LRLLMDRGFGVSLCDNGKW 130
LR RG GV +C N +W
Sbjct: 140 LRRRWRRGIGVVVC*NFRW 84
>BP053615
Length = 460
Score = 27.7 bits (60), Expect = 9.8
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = -2
Query: 392 KLFKKKEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDG 435
K+ K+EI +M I GL++ K C+RY L + +++ G
Sbjct: 342 KISFKQEISEVMHIHGLIKLKWSFICICMRYALLYTLLREEDFG 211
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,521,352
Number of Sequences: 28460
Number of extensions: 211723
Number of successful extensions: 918
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 906
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 914
length of query: 1053
length of database: 4,897,600
effective HSP length: 100
effective length of query: 953
effective length of database: 2,051,600
effective search space: 1955174800
effective search space used: 1955174800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145202.5


