

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.10 + phase: 0
(235 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
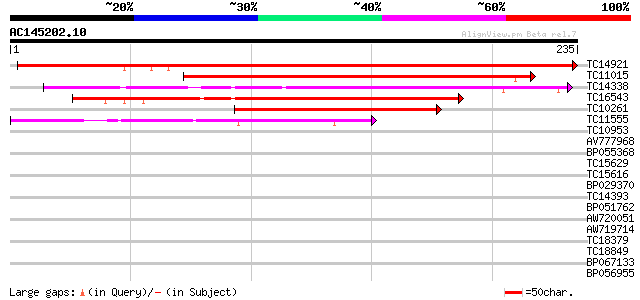
Score E
Sequences producing significant alignments: (bits) Value
TC14921 similar to UP|Q8L949 (Q8L949) Plastid protein, partial (... 318 4e-88
TC11015 similar to PIR|T52623|T52623 DAG protein homolog [import... 230 1e-61
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 146 3e-36
TC16543 129 3e-31
TC10261 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloropl... 107 2e-24
TC11555 similar to UP|O49429 (O49429) DAG-like protein, partial ... 60 3e-10
TC10953 similar to UP|Q40540 (Q40540) Pit2 protein, partial (84%) 37 0.002
AV777968 35 0.009
BP055368 35 0.011
TC15629 similar to UP|Q40540 (Q40540) Pit2 protein, partial (21%) 35 0.015
TC15616 similar to UP|DAG_ANTMA (Q38732) DAG protein, chloroplas... 34 0.019
BP029370 32 0.095
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 32 0.12
BP051762 31 0.21
AW720051 30 0.47
AW719714 29 0.62
TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycop... 29 0.62
TC18849 29 0.62
BP067133 29 0.62
BP056955 29 0.81
>TC14921 similar to UP|Q8L949 (Q8L949) Plastid protein, partial (73%)
Length = 1030
Score = 318 bits (816), Expect = 4e-88
Identities = 162/238 (68%), Positives = 183/238 (76%), Gaps = 6/238 (2%)
Frame = +3
Query: 4 RAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPR-FIIPMSQTIFQ-SLHF 61
RA+ASS R T ++T+T TTT P SPR + +SQ F + F
Sbjct: 51 RALASSCTRHSTLLTMRLFSTTTTTTTTAISLCKSPILSQSPRRSVYALSQVFFTPTTRF 230
Query: 62 GGI----HRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATK 117
GGI +RAG YSPL + SFSDRPP EMAPLFPGCDYNHWLI++DKPGGEGATK
Sbjct: 231 GGIRCRVNRAGDSAYSPLNSSGSNSFSDRPPTEMAPLFPGCDYNHWLIVMDKPGGEGATK 410
Query: 118 QQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPD 177
Q MIDCYV+TLA+VLGSEEEAKKKIYNVSCERYFGFGCE+DEETSNK EG+ GVLFVLPD
Sbjct: 411 QDMIDCYVQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKFEGMNGVLFVLPD 590
Query: 178 SYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRDNQR 235
SYVDPE++DYGAELFVNGEIVQR PERQRRVEPQ QR RPRY+DRT+YVRRR+N +
Sbjct: 591 SYVDPEYKDYGAELFVNGEIVQRPPERQRRVEPQQQRHQDRPRYNDRTRYVRRRENSQ 764
>TC11015 similar to PIR|T52623|T52623 DAG protein homolog [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(65%)
Length = 924
Score = 230 bits (587), Expect = 1e-61
Identities = 107/147 (72%), Positives = 127/147 (85%), Gaps = 1/147 (0%)
Frame = +1
Query: 73 SPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVL 132
S L GS +S S++P AEM PLFPGCDYNHWLII+DKPGGEGATK+QMIDCY+KTLA+V+
Sbjct: 196 SELTTGSPSSISEKPAAEMPPLFPGCDYNHWLIIMDKPGGEGATKKQMIDCYIKTLAKVV 375
Query: 133 GSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELF 192
GSEEEAKK+IY++SCE+Y GFGCE+DEETS+KL+G G++FVLPDSYVDPE+QDYG ELF
Sbjct: 376 GSEEEAKKRIYSISCEKYLGFGCEIDEETSDKLDGKAGIMFVLPDSYVDPEYQDYGGELF 555
Query: 193 VNGEIVQRSPERQRRV-EPQAQRGDSR 218
VNG IVQR PERQ+R + Q R SR
Sbjct: 556 VNGAIVQRPPERQKRFGQKQTTRKTSR 636
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 146 bits (369), Expect = 3e-36
Identities = 88/226 (38%), Positives = 132/226 (57%), Gaps = 7/226 (3%)
Frame = +2
Query: 15 FSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGGIHRAGYCNYSP 74
F+ + STST+T+T + + L + L++ IP + +F S P
Sbjct: 86 FTRSLSTSTTTTTPSLSALSFLRRYRPLSAVS--IPTCRHLFPSARSLSTRATTSSLNDP 259
Query: 75 LVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGS 134
N ++S+RPP E L GCD+ HWL++++KP G+ T+ +ID Y+KTLAQV+GS
Sbjct: 260 -----NPNWSNRPPKETI-LLDGCDFEHWLVVMEKPEGD-PTRDDIIDGYIKTLAQVVGS 418
Query: 135 EEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVN 194
E+EA+ KIY+VS + YF FG + EE S KL+ +P V +VLPDSY++ +DYG E F+N
Sbjct: 419 EQEARMKIYSVSTKHYFAFGALVSEELSYKLKELPKVRWVLPDSYLNVREKDYGGEPFIN 598
Query: 195 GEIVQRSPE------RQRRVEPQAQRGDSRPRYHDRTK-YVRRRDN 233
GE V P+ R + R + RPR +DR++ + RRR+N
Sbjct: 599 GEAVPYDPKYHEEWVRNNARANERNRRNDRPRNNDRSRNFDRRREN 736
>TC16543
Length = 575
Score = 129 bits (325), Expect = 3e-31
Identities = 75/170 (44%), Positives = 107/170 (62%), Gaps = 8/170 (4%)
Frame = +1
Query: 27 TYTTTVAGALSK-PSSLTSPR-----FIIPMSQT--IFQSLHFGGIHRAGYCNYSPLVPG 78
T +T++ ALS PSS ++P F + QT + ++ F ++ YSPL
Sbjct: 70 TLASTLSRALSSSPSSFSTPSRCRCIFALAAKQTLPVPDTVKFSVRTKSSGSGYSPLNDP 249
Query: 79 SNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEA 138
S ++S+RPP E L GCDY HWLI+++ P ++Q+M++ YVKTL Q++GSEEEA
Sbjct: 250 S-PNWSNRPPKETI-LLDGCDYEHWLIVMEFPENPKPSEQEMVNAYVKTLTQIVGSEEEA 423
Query: 139 KKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYG 188
KKIY+VS Y GFG + EE S K++ PGVL+VLPDSY+D ++DYG
Sbjct: 424 MKKIYSVSTHTYTGFGALISEELSYKVKEXPGVLWVLPDSYLDVPNKDYG 573
>TC10261 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (41%)
Length = 510
Score = 107 bits (266), Expect = 2e-24
Identities = 48/86 (55%), Positives = 63/86 (72%)
Frame = +2
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
+ PGCDYNHWLI+++ P +++QMI+ Y+ TL+ VLGS EEAKK +Y S Y GF
Sbjct: 251 MLPGCDYNHWLIVMEFPKDPAPSREQMIETYLFTLSTVLGSMEEAKKNMYAFSTTTYTGF 430
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSY 179
C +DE TS K +G+PGVL+VLPDSY
Sbjct: 431 QCTVDEATSEKFKGLPGVLWVLPDSY 508
>TC11555 similar to UP|O49429 (O49429) DAG-like protein, partial (14%)
Length = 467
Score = 60.5 bits (145), Expect = 3e-10
Identities = 51/156 (32%), Positives = 78/156 (49%), Gaps = 4/156 (2%)
Frame = +3
Query: 1 MTTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLH 60
++ A+ASS RL + T+ + +++ T SL++P +S +QS +
Sbjct: 33 LSLTAMASSSLIRLRRSLTALAGFHRSFSAT---------SLSAP-IRYGVSSPCWQSPN 182
Query: 61 FGGIHRAGYCNYSPLVPGSNTSFSDRPPAEMAP---LFPGCDYNHWLIIIDKPGGEGATK 117
G R+ + + L+ +S + P E+ P LF GCDYNHWL ++D P
Sbjct: 183 VGFQSRSFMSSPTSLL-SIRSSQRNIPDGEIGPDEILFEGCDYNHWLFVMDFPRDNKPPP 359
Query: 118 QQMIDCYVKTLAQVLG-SEEEAKKKIYNVSCERYFG 152
+QM+ Y +T A+ L S EEAKKKIY S Y G
Sbjct: 360 EQMVRTYEETCAKGLNISLEEAKKKIYACSTTTYTG 467
>TC10953 similar to UP|Q40540 (Q40540) Pit2 protein, partial (84%)
Length = 591
Score = 37.4 bits (85), Expect = 0.002
Identities = 29/103 (28%), Positives = 45/103 (43%)
Frame = +1
Query: 74 PLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLG 133
PL +NTS S P ++ +KP E +++TL+ VLG
Sbjct: 109 PLDSNANTSSSSAPAVH-------------IVYTEKPQDEEPEAY-----HIRTLSAVLG 234
Query: 134 SEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLP 176
SEE AK+ + GF +L + +++ PGVL V+P
Sbjct: 235 SEETAKEALLYSYKSAASGFSAKLTPDQVDQISKQPGVLQVVP 363
>AV777968
Length = 383
Score = 35.4 bits (80), Expect = 0.009
Identities = 14/22 (63%), Positives = 17/22 (76%)
Frame = -2
Query: 147 CERYFGFGCELDEETSNKLEGI 168
CE F GCE+DEET +KLEG+
Sbjct: 178 CEMSFRLGCEMDEETPDKLEGL 113
>BP055368
Length = 514
Score = 35.0 bits (79), Expect = 0.011
Identities = 19/30 (63%), Positives = 21/30 (69%), Gaps = 1/30 (3%)
Frame = -1
Query: 190 ELFVNGEIVQRSPERQRRV-EPQAQRGDSR 218
ELFVNG IVQR PERQ+R + Q R SR
Sbjct: 352 ELFVNGAIVQRPPERQKRFGQKQTTRKTSR 263
>TC15629 similar to UP|Q40540 (Q40540) Pit2 protein, partial (21%)
Length = 616
Score = 34.7 bits (78), Expect = 0.015
Identities = 20/53 (37%), Positives = 31/53 (57%), Gaps = 1/53 (1%)
Frame = +3
Query: 125 VKTLAQVLGSEEEAKKK-IYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLP 176
++TL+ VLGSEE AK+ +Y+ E F + E + +L PGVL ++P
Sbjct: 255 IRTLSAVLGSEEAAKEAVVYDFEDEINDNFSAVITPENAVQLLKQPGVLHIIP 413
>TC15616 similar to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (13%)
Length = 650
Score = 34.3 bits (77), Expect = 0.019
Identities = 18/53 (33%), Positives = 33/53 (61%)
Frame = +3
Query: 180 VDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRD 232
+D +++DYG + ++NGEI+ P + +P+ + G SR +D +Y R+RD
Sbjct: 3 IDVKNKDYGGDKYINGEII---PSKYPTYQPK-RSGGSR---NDSRRYERKRD 140
>BP029370
Length = 364
Score = 32.0 bits (71), Expect = 0.095
Identities = 17/35 (48%), Positives = 25/35 (70%)
Frame = +2
Query: 15 FSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFII 49
++TTT+T+T T+T TTT+A SK S +T P +I
Sbjct: 170 YTTTTTTTTRTTTTTTTIAS--SKKSQIT*PMPLI 268
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 31.6 bits (70), Expect = 0.12
Identities = 18/46 (39%), Positives = 27/46 (58%)
Frame = +1
Query: 9 SFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQT 54
+F RR S++ S TST+T T + S PS+ +PR ++P S T
Sbjct: 244 AFHRR--SSSASWWTSTATTTASSISPSSPPSAAPTPRTVVPPSST 375
>BP051762
Length = 557
Score = 30.8 bits (68), Expect = 0.21
Identities = 18/50 (36%), Positives = 27/50 (54%)
Frame = -2
Query: 5 AIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQT 54
+I + F FS ST + ++T + GAL ++LTS RF P+S T
Sbjct: 331 SIITGFNLDHFSNIPSTVSPSATDDFRMEGALVSSNNLTSKRFSYPVSTT 182
>AW720051
Length = 564
Score = 29.6 bits (65), Expect = 0.47
Identities = 22/61 (36%), Positives = 31/61 (50%), Gaps = 9/61 (14%)
Frame = +2
Query: 16 STTTSTSTSTSTYTTTVAGALSKP---SSLTSPRFIIPMSQ------TIFQSLHFGGIHR 66
S TT+T+TS S + A + P SS TSP +P +Q T+F+ LH HR
Sbjct: 173 SATTTTTTSLSRAISAKAAVGTGPKAASSATSPSAAVPGNQSAPSTKTLFRLLHHHHHHR 352
Query: 67 A 67
+
Sbjct: 353 S 355
>AW719714
Length = 586
Score = 29.3 bits (64), Expect = 0.62
Identities = 13/24 (54%), Positives = 19/24 (79%)
Frame = +3
Query: 8 SSFKRRLFSTTTSTSTSTSTYTTT 31
SSFKR F+TTT++S S+++TT
Sbjct: 189 SSFKRDSFTTTTTSSRRRSSFSTT 260
>TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycoprotein
DZ-HRGP precursor, partial (8%)
Length = 584
Score = 29.3 bits (64), Expect = 0.62
Identities = 13/33 (39%), Positives = 24/33 (72%)
Frame = +3
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSPRFI 48
+TTT+T+++++T TTT+ G+ + S TS +I
Sbjct: 252 ATTTTTTSTSTTRTTTILGSTTTSSLWTSSFYI 350
>TC18849
Length = 770
Score = 29.3 bits (64), Expect = 0.62
Identities = 17/39 (43%), Positives = 21/39 (53%), Gaps = 4/39 (10%)
Frame = +1
Query: 11 KRRLFSTTT----STSTSTSTYTTTVAGALSKPSSLTSP 45
+RR F T ++STS S T G LS P S+TSP
Sbjct: 91 RRRRFQKTNGS*HTSSTSPSASVTCFHGTLSSPPSITSP 207
>BP067133
Length = 507
Score = 29.3 bits (64), Expect = 0.62
Identities = 29/93 (31%), Positives = 41/93 (43%), Gaps = 7/93 (7%)
Frame = +1
Query: 7 ASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPM-------SQTIFQSL 59
ASS S+ +S++TSTST TT A S P++ S P S+T S+
Sbjct: 235 ASSASTTTASSASSSATSTSTSPTTTTTATSSPTASPSSPSPTPASSPPSAPSRTSPPSI 414
Query: 60 HFGGIHRAGYCNYSPLVPGSNTSFSDRPPAEMA 92
F + S L+ G +TS S A +A
Sbjct: 415 PFSNTWTP---SASSLISGPSTSTSHYSAARIA 504
>BP056955
Length = 581
Score = 28.9 bits (63), Expect = 0.81
Identities = 17/33 (51%), Positives = 23/33 (69%)
Frame = +1
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSPRFI 48
ST+TSTSTSTST T+T S +S ++ RF+
Sbjct: 181 STSTSTSTSTSTSTST-----SISTSTSTTRFV 264
Score = 27.3 bits (59), Expect = 2.4
Identities = 14/23 (60%), Positives = 16/23 (68%)
Frame = +1
Query: 16 STTTSTSTSTSTYTTTVAGALSK 38
ST+TSTS STST TT G S+
Sbjct: 211 STSTSTSISTSTSTTRFVGRPSR 279
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,965,891
Number of Sequences: 28460
Number of extensions: 57359
Number of successful extensions: 527
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 473
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 492
length of query: 235
length of database: 4,897,600
effective HSP length: 87
effective length of query: 148
effective length of database: 2,421,580
effective search space: 358393840
effective search space used: 358393840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC145202.10


