

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
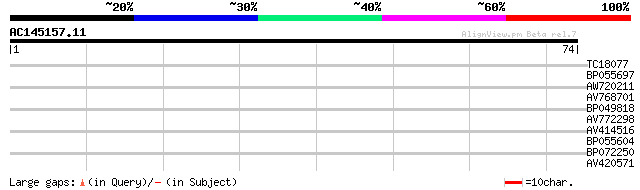
Query= AC145157.11 + phase: 0
(74 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18077 similar to UP|Q8LBH4 (Q8LBH4) Ids4-like protein, partial... 26 1.6
BP055697 25 2.7
AW720211 25 2.7
AV768701 25 3.5
BP049818 25 3.5
AV772298 24 6.1
AV414516 24 6.1
BP055604 24 6.1
BP072250 23 7.9
AV420571 23 7.9
>TC18077 similar to UP|Q8LBH4 (Q8LBH4) Ids4-like protein, partial (51%)
Length = 995
Score = 25.8 bits (55), Expect = 1.6
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -2
Query: 48 PRDGFNLTLKINLSKVPANQGISG 71
P +NL LK++ VP NQ +SG
Sbjct: 289 PSPPYNLLLKVDCHLVPTNQVLSG 218
>BP055697
Length = 191
Score = 25.0 bits (53), Expect = 2.7
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +2
Query: 55 TLKINLSKVPANQGISGFLV 74
T K+NLS +PA GIS L+
Sbjct: 14 TRKLNLSPLPARYGISSMLL 73
>AW720211
Length = 465
Score = 25.0 bits (53), Expect = 2.7
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +1
Query: 2 KNPNIFLLSVSLPTPSSETIFVCGL 26
+NPN+F S P S+++F+ G+
Sbjct: 112 QNPNLFTTHFSNPNSPSKSLFILGV 186
>AV768701
Length = 292
Score = 24.6 bits (52), Expect = 3.5
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 7 FLLSVSLPTPSSETIFVCGLPFGAIEAIKAAY 38
FLL +++ T IFVC LP ++ +K Y
Sbjct: 104 FLLLINVTT---NVIFVCALPTQIVKVLKLKY 18
>BP049818
Length = 604
Score = 24.6 bits (52), Expect = 3.5
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = -3
Query: 1 MKNPNIFLLSVSLPTP 16
+KNPN F++S PTP
Sbjct: 548 IKNPNTFVVSSKYPTP 501
>AV772298
Length = 528
Score = 23.9 bits (50), Expect = 6.1
Identities = 11/18 (61%), Positives = 14/18 (77%)
Frame = -3
Query: 6 IFLLSVSLPTPSSETIFV 23
+ L+ VSLPT + ETIFV
Sbjct: 289 LILILVSLPTYTLETIFV 236
>AV414516
Length = 424
Score = 23.9 bits (50), Expect = 6.1
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +1
Query: 5 NIFLLSVSLPTPSSET 20
NIF+ VSLP PSS +
Sbjct: 1 NIFIFFVSLPPPSSSS 48
>BP055604
Length = 464
Score = 23.9 bits (50), Expect = 6.1
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = -2
Query: 3 NPNIFLLSVSLPTPSSE 19
+PN F LS+ +P+P E
Sbjct: 208 SPNAFCLSIQMPSPQPE 158
>BP072250
Length = 444
Score = 23.5 bits (49), Expect = 7.9
Identities = 13/39 (33%), Positives = 18/39 (45%)
Frame = -2
Query: 27 PFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINLSKVPA 65
PF E +K Y + P+D LTL + LS P+
Sbjct: 410 PFRMHEFLKGQYPCALPNAPQPKDSLILTLLLTLSIEPS 294
>AV420571
Length = 304
Score = 23.5 bits (49), Expect = 7.9
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 38 YGSLVQILDPPRDGFNL 54
YGS+V+IL+PP NL
Sbjct: 112 YGSVVRILNPPLTTTNL 162
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.141 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,315,178
Number of Sequences: 28460
Number of extensions: 14959
Number of successful extensions: 76
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 76
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 76
length of query: 74
length of database: 4,897,600
effective HSP length: 50
effective length of query: 24
effective length of database: 3,474,600
effective search space: 83390400
effective search space used: 83390400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 48 (23.1 bits)
Medicago: description of AC145157.11


