

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
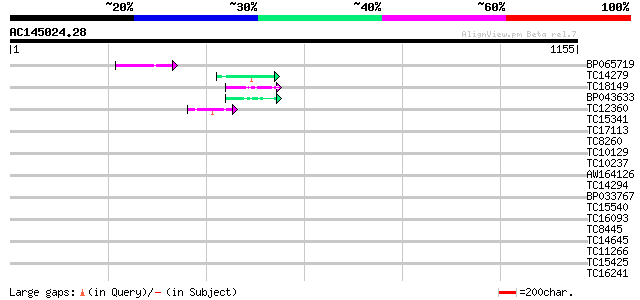
Query= AC145024.28 - phase: 0
(1155 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065719 51 9e-07
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 44 2e-04
TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structura... 43 3e-04
BP043633 43 3e-04
TC12360 similar to UP|O22196 (O22196) At2g40820 protein, partial... 43 3e-04
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 40 0.002
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 40 0.002
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 40 0.002
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 40 0.002
TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precu... 40 0.003
AW164126 39 0.006
TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 37 0.014
BP033767 36 0.031
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 36 0.031
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 36 0.040
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 36 0.040
TC14645 similar to UP|Q41122 (Q41122) Proline-rich protein precu... 36 0.040
TC11266 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pro... 36 0.040
TC15425 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall... 36 0.040
TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%) 36 0.040
>BP065719
Length = 567
Score = 51.2 bits (121), Expect = 9e-07
Identities = 40/132 (30%), Positives = 61/132 (45%), Gaps = 4/132 (3%)
Frame = +3
Query: 215 VPDVNIPPKFKMPVFEKYQGDT--CPQNHLTMYISKMIAYKNNVPLLIHCFQDSLTGPAH 272
V + +P +K+P F K+ GD+ H+ Y + N L + F SLT A
Sbjct: 147 VLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAF 326
Query: 273 TWFMGL--KGVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKA 330
TWF L + V T+ QL F +Q+ + S K+L S+ +K ES +Y RF
Sbjct: 327 TWFTTLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLK 503
Query: 331 AQIRPPLDEREL 342
++ + E EL
Sbjct: 504 SRCFTHVSEHEL 539
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 43.5 bits (101), Expect = 2e-04
Identities = 39/141 (27%), Positives = 49/141 (34%), Gaps = 13/141 (9%)
Frame = +1
Query: 421 AHGGPQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQP 479
+H P P Y P P +P P Y +SP P P+P P Y+ + P
Sbjct: 157 SHSPPPPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPP 321
Query: 480 YY---PQQPYQQRP-----YNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLP----SL 527
YY P P P +PP P P PY+ + TY P L
Sbjct: 322 YYYKSPPPPTSSPPPPYHYVSPPPPSPSPPPPYHYASPPPPSPSPAPTYIYKSPPPPVKL 501
Query: 528 LAQNLVQTIPPPRIPDPLPRW 548
T PPP P P P +
Sbjct: 502 PPPPYHYTSPPPPSPSPAPTY 564
Score = 39.3 bits (90), Expect = 0.004
Identities = 37/142 (26%), Positives = 50/142 (35%), Gaps = 18/142 (12%)
Frame = +1
Query: 425 PQPVYSNHP-----FVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQ 478
P+P Y P + + P +P P Y QSP P +P P Y+
Sbjct: 43 PKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPSPSPPP 222
Query: 479 PYY-----PQQPYQQRPY---NPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQ 530
PYY P P PY +PP P P PY ++ P Y + P +
Sbjct: 223 PYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHYVSPPPPSP 402
Query: 531 N----LVQTIPPPRIPDPLPRW 548
+ PPP P P P +
Sbjct: 403 SPPPPYHYASPPPPSPSPAPTY 468
Score = 37.0 bits (84), Expect = 0.018
Identities = 28/103 (27%), Positives = 33/103 (31%), Gaps = 3/103 (2%)
Frame = +1
Query: 450 YQSPRPQGPAP---YYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQ 506
Y+SP P P P YY P Y + P PY + PP P P Y
Sbjct: 13 YKSPPPPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKS 192
Query: 507 QNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWY 549
P P Y + PPP P P P +Y
Sbjct: 193 PPPPSPSPPPPYYYKS--------------PPPPSPSPPPPYY 279
Score = 33.5 bits (75), Expect = 0.20
Identities = 25/97 (25%), Positives = 33/97 (33%), Gaps = 3/97 (3%)
Frame = +1
Query: 425 PQPVYSNHPFVANITPQMTAPQNP---NYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY 481
P P S P ++P +P P +Y SP P P+P P Y +
Sbjct: 340 PPPTSSPPPPYHYVSPPPPSPSPPPPYHYASPPPPSPSP------APTYIYKSPPPPVKL 501
Query: 482 PQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPM 518
P PY PP P P Y + P P+
Sbjct: 502 PPPPYHYTSPPPPSPSPAPTYIYKSPPPPTKSPPPPV 612
>TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structural protein
pp78/81, partial (5%)
Length = 710
Score = 43.1 bits (100), Expect = 3e-04
Identities = 41/120 (34%), Positives = 52/120 (43%), Gaps = 6/120 (5%)
Frame = +3
Query: 440 PQMTAPQNPNYQSPRPQGPA--PYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQ 497
P P N Q+ P P PY+PP Q Q QQP P QP+ Q P PP QQ
Sbjct: 162 PPSFPPPNQFSQTQIPAVPQRDPYFPPPVQSQETPNQQYQQPLAP-QPHPQ-PGAPPHQQ 335
Query: 498 ----PRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDL 553
P PQ Y+Q Q P+P L S L ++ + PP +P + Y P+L
Sbjct: 336 YQQTPHPQ--YSQPAPPQHQPPIPSINPPQLQSSLGHHVEE---PPYVPS---QTYPPNL 491
Score = 40.8 bits (94), Expect = 0.001
Identities = 38/123 (30%), Positives = 47/123 (37%), Gaps = 25/123 (20%)
Frame = +3
Query: 425 PQPVYSNH--------PFVANITPQMTAPQNPNYQ-SPRPQ----GPAPYYPPLYQPLYN 471
P PV S P PQ AP + YQ +P PQ P + PP+ P N
Sbjct: 237 PPPVQSQETPNQQYQQPLAPQPHPQPGAPPHQQYQQTPHPQYSQPAPPQHQPPI--PSIN 410
Query: 472 LQQLS--------QQPYYPQQPYQQRPYNPPQQQPR-PQAPYNQQ---NQKQQFDPLPMT 519
QL + PY P Q Y PP Q P P AP +QQ ++P P
Sbjct: 411 PPQLQSSLGHHVEEPPYVPSQTYPPNLRQPPSQPPSGPPAPPSQQFYGTAPHAYEPPPSR 590
Query: 520 YGA 522
G+
Sbjct: 591 SGS 599
>BP043633
Length = 542
Score = 43.1 bits (100), Expect = 3e-04
Identities = 36/117 (30%), Positives = 43/117 (35%), Gaps = 2/117 (1%)
Frame = +1
Query: 440 PQMTAPQNPNYQSPRPQGPAPYYP--PLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQ 497
P P +PN+ PQ P P+ P P Y P + SQ P P Q P PP
Sbjct: 256 PPPPPPPDPNHHHHFPQIPPPHPPQAPWYSPQFQYHHPSQT---PSPPPQWGP--PPPPP 420
Query: 498 PRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDLH 554
P P PY+ +QF P P PPPR P P + P H
Sbjct: 421 PPPSNPYSY--HPKQFPPPP-------------------PPPRPPHPPHPQFPPHSH 528
>TC12360 similar to UP|O22196 (O22196) At2g40820 protein, partial (11%)
Length = 1030
Score = 42.7 bits (99), Expect = 3e-04
Identities = 35/114 (30%), Positives = 48/114 (41%), Gaps = 12/114 (10%)
Frame = +3
Query: 363 SQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSSGNK-KFGNGYP----------- 410
S K + V+ +R+ E K S GG +S GNK +FG YP
Sbjct: 429 SNKPSGFVDVSIRVFE--DKEEVDSHPGEGGGILLSDHGNKGRFGQAYPHQMAPASFHGP 602
Query: 411 KRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPP 464
++ AQ +H P P ++P+V P A P+YQ PR P P PP
Sbjct: 603 EKQAQINTPYSHPVPFPANYSNPYVGG--PSYPAAAGPSYQPPRTHPPPPPPPP 758
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 40.4 bits (93), Expect = 0.002
Identities = 27/66 (40%), Positives = 29/66 (43%), Gaps = 7/66 (10%)
Frame = +1
Query: 439 TPQM-----TAPQN-PNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ-PYQQRPY 491
TPQM APQ N P P YP QP QQ QQ +PQQ P Q +P
Sbjct: 757 TPQMQDMSRVAPQPCTNXSGXSPPPPVQQYPQYQQPQQQQQQQQQQQQWPQQLPQQVQPT 936
Query: 492 NPPQQQ 497
PP Q
Sbjct: 937 QPPSMQ 954
Score = 35.4 bits (80), Expect = 0.052
Identities = 28/100 (28%), Positives = 38/100 (38%), Gaps = 1/100 (1%)
Frame = +1
Query: 427 PVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPY 486
P+ N VA + P + Y++P+ Q + P QP N S P Q P
Sbjct: 682 PLPGNTQTVAQLPQNQYLPPDQQYRTPQMQDMSRVAP---QPCTNXSGXSPPPPVQQYPQ 852
Query: 487 QQRPYNPPQQQPRPQA-PYNQQNQKQQFDPLPMTYGALLP 525
Q+P QQQ + Q P Q Q P M + P
Sbjct: 853 YQQPQQQQQQQQQQQQWPQQLPQQVQPTQPPSMQSPQIRP 972
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 40.0 bits (92), Expect = 0.002
Identities = 39/129 (30%), Positives = 50/129 (38%), Gaps = 2/129 (1%)
Frame = +1
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLY--NLQQLSQQPYYP 482
P P + P + N NP SP P P P+ PP PL+ Q S P P
Sbjct: 445 PNP-FQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSPPPLFPNPFQPPSPPPLIP 621
Query: 483 QQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP 542
P+Q P +PP P P P + P P ++P L T PPP P
Sbjct: 622 -NPFQPPP-SPPPFIPNPFQPPPSKQPPSPLFPFP---PIVIPGL-------TPPPP--P 759
Query: 543 DPLPRWYRP 551
P P ++ P
Sbjct: 760 PPPPPFFPP 786
Score = 33.9 bits (76), Expect = 0.15
Identities = 20/55 (36%), Positives = 23/55 (41%)
Frame = +1
Query: 490 PYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDP 544
P+ PP P P P N F P P+ P L N Q PPP +PDP
Sbjct: 394 PFFPPPLIPNPFQPPLVPNP---FQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDP 549
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 40.0 bits (92), Expect = 0.002
Identities = 43/191 (22%), Positives = 69/191 (35%), Gaps = 11/191 (5%)
Frame = +3
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P + +NH ++ P P+ +Y SP P +P PP Y+ + + PY
Sbjct: 84 PSEISANHYAYSSPPPP---PKPYSYHSPPPPVHSP--PPPYEKPHPVYHSPPPPYEKPH 248
Query: 485 PYQQRPYNPPQQQP-RPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPP--RI 541
P P PP +P + +P ++ P P+ + P + PPP
Sbjct: 249 PVYHSPPPPPPHKPYKYPSPPPPPHKYPHPHPHPVYHSPPPPPPKKHYKYSSPPPPVHTY 428
Query: 542 PDPLPRWYR--------PDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTFTDPDAV 593
P P P ++ P H +YH P E+ + ++ T P V
Sbjct: 429 PHPHPVYHSPPPPVHTYPHPHPVYHSPPPPPPHEKPYKYASPPPPPVH------TYPHPV 590
Query: 594 AQNNPLPTHGP 604
+ P P H P
Sbjct: 591 YHSPPPPVHSP 623
Score = 30.8 bits (68), Expect = 1.3
Identities = 33/116 (28%), Positives = 50/116 (42%)
Frame = +2
Query: 44 STPVVVVTAAPEGPPQPSPTTTTTIGLTQPLITDFSSGNMATMGNSGFRPLGPQGPFATP 103
STP V + P P P +TTTT Q LI+ +S ++ + S PL P T
Sbjct: 308 STPTTQVPSPPPSPSLPLSSTTTT*EALQVLISSTTSSHLPSPSPS--LPLSP-----TT 466
Query: 104 QYSMPSGYPWGMPIATNGVFGPGATEMPFAQGQQTSAHFQMSQPIPQATMTQAGPT 159
+PS P +P ++ A ++ A TS+H S +P + + PT
Sbjct: 467 SSHLPSPTP-SLPFSSTTTTT*EAIQICIA-STTTSSHLP-SPSLPFTSSSSPFPT 625
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 40.0 bits (92), Expect = 0.002
Identities = 37/123 (30%), Positives = 47/123 (38%), Gaps = 4/123 (3%)
Frame = +1
Query: 427 PVYSNHPFVANITPQMTAPQN--PNYQSPRPQGPAPYYPPLYQPL--YNLQQLSQQPYYP 482
P N P I P+ +P PN SP P+ P+P P Y P SQ PY P
Sbjct: 49 PSPGNQPPHTPIPPKTPSPSYSPPNVPSP-PKAPSPNNHPPYTPTPPKTPSPTSQPPYIP 225
Query: 483 QQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP 542
P P + P P P + + +Q P T PS +Q T PP P
Sbjct: 226 LPPKTPSPISQPPHVPTPPSTPSPISQPPYTPTPPKT-----PSPTSQP-PHTPTPPNTP 387
Query: 543 DPL 545
P+
Sbjct: 388 SPI 396
Score = 39.3 bits (90), Expect = 0.004
Identities = 36/129 (27%), Positives = 47/129 (35%), Gaps = 11/129 (8%)
Frame = +1
Query: 427 PVYSNHPFVANITPQMTAPQN-PNYQSPRPQGPAPYYPPLYQPL--YNLQQLSQQPYYPQ 483
P +NHP P+ +P + P Y P+ P+P P + P +SQ PY P
Sbjct: 145 PSPNNHPPYTPTPPKTPSPTSQPPYIPLPPKTPSPISQPPHVPTPPSTPSPISQPPYTPT 324
Query: 484 QPYQQRPYNPPQQQPRP--------QAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQT 535
P P + P P P Q PY K P P +PS + T
Sbjct: 325 PPKTPSPTSQPPHTPTPPNTPSPISQPPYTPTPPK---TPSPTNQPPHIPS-PPNSSSPT 492
Query: 536 IPPPRIPDP 544
PP P P
Sbjct: 493 SQPPNTPSP 519
Score = 33.9 bits (76), Expect = 0.15
Identities = 26/97 (26%), Positives = 34/97 (34%), Gaps = 2/97 (2%)
Frame = +1
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPL--YNLQQLSQQPYYP 482
P P+ S P V + P Y P+ P+P P + P +SQ PY P
Sbjct: 241 PSPI-SQPPHVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPPYTP 417
Query: 483 QQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMT 519
P P N P P P + +Q P T
Sbjct: 418 TPPKTPSPTNQPPHIPSPPNSSSPTSQPPNTPSPPKT 528
>TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75)
(NGM-75), partial (49%)
Length = 803
Score = 39.7 bits (91), Expect = 0.003
Identities = 38/139 (27%), Positives = 51/139 (36%), Gaps = 6/139 (4%)
Frame = +1
Query: 426 QPVYSNHPFVANITPQMTAPQN----PNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY 481
QP + PF P + P + P YQ P + P+ Y PP P Y Q P Y
Sbjct: 19 QPPHEKPPFEK---PPLEKPPHEKPPPEYQPPHKKPPSQYQPP---PEYEPPQEKPPPVY 180
Query: 482 PQQPYQQRP--YNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPP 539
PY + P Y PP ++P P+ Q+ + P PP
Sbjct: 181 -LPPYWKPPPAYPPPYEKPPPEYEPPQEKPPPVYSP-----------------PYERPPT 306
Query: 540 RIPDPLPRWYRPDLHCIYH 558
P P P +Y L +YH
Sbjct: 307 GHPTPYPPFYH--LPPVYH 357
Score = 39.7 bits (91), Expect = 0.003
Identities = 35/120 (29%), Positives = 52/120 (43%), Gaps = 5/120 (4%)
Frame = +1
Query: 448 PNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRP--YNPPQQQPRP-QAPY 504
P Y+ P+ + P Y PP ++P P YP PY++ P Y PPQ++P P +P
Sbjct: 139 PEYEPPQEKPPPVYLPPYWKP---------PPAYP-PPYEKPPPEYEPPQEKPPPVYSPP 288
Query: 505 NQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPL--PRWYRPDLHCIYHQGAP 562
++ P P Y LP + PP P PL P +P ++ H+ P
Sbjct: 289 YERPPTGHPTPYPPFYH--LPPVYH-------PPYEKPPPLYQPPHEKPPIYEPPHEKPP 441
Score = 39.7 bits (91), Expect = 0.003
Identities = 34/127 (26%), Positives = 48/127 (37%), Gaps = 10/127 (7%)
Frame = +1
Query: 410 PKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQN--PNYQSPRPQGPAPYYPPLYQ 467
P+ V + + P P Y P P+ PQ P SP + P +P Y
Sbjct: 154 PQEKPPPVYLPPYWKPPPAYP--PPYEKPPPEYEPPQEKPPPVYSPPYERPPTGHPTPYP 327
Query: 468 PLYNLQQLSQQPYYPQQPYQQRP------YNPPQQQPRPQAPYNQQNQKQQFDP--LPMT 519
P Y+L + PY P Q P Y PP ++P P +++ + P P
Sbjct: 328 PFYHLPPVYHPPYEKPPPLYQPPHEKPPIYEPPHEKPPIYEPPHEKPPLYEPPPGYNPPP 507
Query: 520 YGALLPS 526
YG PS
Sbjct: 508 YGHYPPS 528
>AW164126
Length = 387
Score = 38.5 bits (88), Expect = 0.006
Identities = 25/81 (30%), Positives = 32/81 (38%), Gaps = 1/81 (1%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQ 483
P+P Y P P +P P Y +SP P P P+ P Y+ + PYY
Sbjct: 164 PKPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPVPHPPYYYKSPPPPSPIPHPPYYYS 328
Query: 484 QPYQQRPYNPPQQQPRPQAPY 504
P PP + P P PY
Sbjct: 329 SP------PPPVKSPPPPTPY 373
>TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (65%)
Length = 1192
Score = 37.4 bits (85), Expect = 0.014
Identities = 38/143 (26%), Positives = 49/143 (33%), Gaps = 21/143 (14%)
Frame = +1
Query: 407 NGYPKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLY 466
+GYP + + +G P P Y + P AP + +Y +P P P PP
Sbjct: 64 SGYPNKPSG----YGYGAPPP-YQPYGAAPPSQPYGAAPPSQSYGAPPPPQPYGAAPP-- 222
Query: 467 QPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQP-------RPQAPYNQQNQKQQFD----- 514
QPY P Q PP QP +P APY QK D
Sbjct: 223 ----------SQPYGAPPPSQSYGGPPPPSQPYSASPYGQPSAPYAAPYQKPPKDESHSH 372
Query: 515 ---------PLPMTYGALLPSLL 528
P P YG+ SL+
Sbjct: 373 GGGGGSGYPPPPSAYGSPFASLV 441
>BP033767
Length = 541
Score = 36.2 bits (82), Expect = 0.031
Identities = 20/63 (31%), Positives = 26/63 (40%)
Frame = +3
Query: 431 NHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRP 490
+ PF P + P NP SP P P P YP P Q + P++P P P
Sbjct: 147 HQPFTQGSLPP-SQPPNPPPPSPSPSPPNPKYPFSSTPTTTTTQNASTPFFPTYPSPPPP 323
Query: 491 YNP 493
+P
Sbjct: 324 PSP 332
Score = 34.3 bits (77), Expect = 0.12
Identities = 33/105 (31%), Positives = 40/105 (37%), Gaps = 6/105 (5%)
Frame = +1
Query: 446 QNPNYQSPRPQGPAPYYPP-LYQPLYNLQQ-LSQQPYYPQQPYQQRPYNPPQQQ---PRP 500
QN ++ P PQ P P PP L P +L L P P P P +PP P P
Sbjct: 232 QNTHFPPP-PQPPPPKMPPHLSSPHTHLHHHLLLPPPSPPSP----PTSPPSSSLKPPNP 396
Query: 501 QAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPP-RIPDP 544
P + P P PS A +T PP R+ DP
Sbjct: 397 NPPTSSSASPLPPSPAPSPSSLSPPSSTADAAAKTTPPTRRLSDP 531
Score = 28.1 bits (61), Expect = 8.3
Identities = 21/76 (27%), Positives = 29/76 (37%)
Frame = +3
Query: 467 QPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPS 526
+P+ N +++ QP+ +P NPP P P P N K F P T S
Sbjct: 120 EPIIN-RRILHQPFTQGSLPPSQPPNPPPPSPSPSPP----NPKYPFSSTPTTTTTQNAS 284
Query: 527 LLAQNLVQTIPPPRIP 542
+ PPP P
Sbjct: 285 TPFFPTYPSPPPPPSP 332
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 36.2 bits (82), Expect = 0.031
Identities = 26/71 (36%), Positives = 29/71 (40%)
Frame = +2
Query: 472 LQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQN 531
L QL QQ QQP Q QQQ PQ Q +Q QQ LP L Q
Sbjct: 8 LPQLQQQQQLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQ---------QQLPQLQQQQ 160
Query: 532 LVQTIPPPRIP 542
L Q P ++P
Sbjct: 161 LSQLQPQQQLP 193
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 35.8 bits (81), Expect = 0.040
Identities = 23/54 (42%), Positives = 25/54 (45%)
Frame = +1
Query: 473 QQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPS 526
Q QQP QQ QQ+P PQQQ + Q QQ Q QQ L M PS
Sbjct: 133 QHQFQQPIQQQQQQQQQPIQQPQQQMQQQ----QQEQHQQSGDLKMRSATSSPS 282
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 35.8 bits (81), Expect = 0.040
Identities = 30/86 (34%), Positives = 38/86 (43%), Gaps = 5/86 (5%)
Frame = +1
Query: 445 PQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQ-QPYYPQQPY---QQRPYNPPQQQPRP 500
PQ YQ P + P +QP Q SQ QP + QQ Q +P + Q QP+P
Sbjct: 25 PQPLQYQPQPHLQHQPQHHPHFQPHPQPQLQSQPQPQHIQQSQPQPQSQPQHIQQPQPQP 204
Query: 501 QAPYNQQNQKQ-QFDPLPMTYGALLP 525
Q + QQ+Q Q QF Y P
Sbjct: 205 QLQHIQQSQPQPQFQNQHFQYPGTYP 282
Score = 33.1 bits (74), Expect = 0.26
Identities = 33/104 (31%), Positives = 42/104 (39%), Gaps = 2/104 (1%)
Frame = +1
Query: 419 MVAHGGPQPV-YSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQ 477
M H PQP+ Y P + + PQ+ + P PQ P + Q
Sbjct: 7 MQLHNEPQPLQYQPQPHLQH------QPQHHPHFQPHPQPQLQSQPQPQHIQQSQPQPQS 168
Query: 478 QPYYPQQPYQQRPYNPPQQ-QPRPQAPYNQQNQKQQFDPLPMTY 520
QP + QQP Q QQ QP+PQ QNQ Q+ P TY
Sbjct: 169 QPQHIQQPQPQPQLQHIQQSQPQPQF----QNQHFQY---PGTY 279
Score = 31.6 bits (70), Expect = 0.75
Identities = 24/78 (30%), Positives = 33/78 (41%), Gaps = 11/78 (14%)
Frame = +1
Query: 474 QLSQQP----YYPQQPYQQRPYNPPQQQPRPQAPYNQQNQK---QQFDPLPMTYGALL-- 524
QL +P Y PQ Q +P + P QP PQ Q Q QQ P P + +
Sbjct: 10 QLHNEPQPLQYQPQPHLQHQPQHHPHFQPHPQPQLQSQPQPQHIQQSQPQPQSQPQHIQQ 189
Query: 525 --PSLLAQNLVQTIPPPR 540
P Q++ Q+ P P+
Sbjct: 190 PQPQPQLQHIQQSQPQPQ 243
>TC14645 similar to UP|Q41122 (Q41122) Proline-rich protein precursor,
partial (52%)
Length = 1060
Score = 35.8 bits (81), Expect = 0.040
Identities = 23/83 (27%), Positives = 33/83 (39%), Gaps = 7/83 (8%)
Frame = +2
Query: 425 PQPVYSNHPFV-------ANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQ 477
P P Y NHP + I P AP + ++ P PAP PP++ P +
Sbjct: 200 PPPNYHNHPVAPVHPPTHSPIHPPAHAPHHHHHHHPPAPAPAPIKPPVHPP-------TS 358
Query: 478 QPYYPQQPYQQRPYNPPQQQPRP 500
P P + ++PP P P
Sbjct: 359 VPVKPPTGHHHH-HHPPSPAPAP 424
>TC11266 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 354
Score = 35.8 bits (81), Expect = 0.040
Identities = 21/68 (30%), Positives = 29/68 (41%), Gaps = 2/68 (2%)
Frame = +1
Query: 433 PFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPY--YPQQPYQQRP 490
P + N T P PN P P GP+P++P + P+ S P +P P P
Sbjct: 4 PLIPNPLQPPTPPLIPNPLQPPPSGPSPFFP--FPPVPGFTPPSPPPVSPFPFLPLFPPP 177
Query: 491 YNPPQQQP 498
+PP P
Sbjct: 178 GSPPASSP 201
>TC15425 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (6%)
Length = 667
Score = 35.8 bits (81), Expect = 0.040
Identities = 38/124 (30%), Positives = 43/124 (34%), Gaps = 2/124 (1%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
PQ Y ++ + PQ P YQS Q P P Y + P Y Q P Y
Sbjct: 2 PQTPYYQPDQRSDTQTNPSRPQTPYYQSAE-QPPVPAYGH-HPPPYGHQP---PPPYQIP 166
Query: 485 PYQQRPYNPPQ--QQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP 542
P PY PPQ QQP Y Q P G + Q PP IP
Sbjct: 167 PTSAAPYPPPQVHQQPPASHEYGQ-------PAYPGWRGPYYNAHAQQPASAPRPPYTIP 325
Query: 543 DPLP 546
P P
Sbjct: 326 SPYP 337
>TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%)
Length = 799
Score = 35.8 bits (81), Expect = 0.040
Identities = 31/107 (28%), Positives = 40/107 (36%), Gaps = 5/107 (4%)
Frame = +1
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLY-----NLQQLSQQP 479
P P S P++A TP T +P Q P Q PA P QP Y + QP
Sbjct: 43 PSPT-SQPPYIA--TPPNT--HSPTSQPPNTQTPANTPSPSSQPPYLSTPPKSNSPTSQP 207
Query: 480 YYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPS 526
Q P P + P P +PY N + + AL P+
Sbjct: 208 PVAQTPRAVTPISSPSSSPTSNSPYPSTNPPSLAPSISTSPPALTPA 348
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,542,969
Number of Sequences: 28460
Number of extensions: 309809
Number of successful extensions: 3049
Number of sequences better than 10.0: 158
Number of HSP's better than 10.0 without gapping: 2504
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2874
length of query: 1155
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1055
effective length of database: 2,051,600
effective search space: 2164438000
effective search space used: 2164438000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145024.28


