

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.2 - phase: 0
(355 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
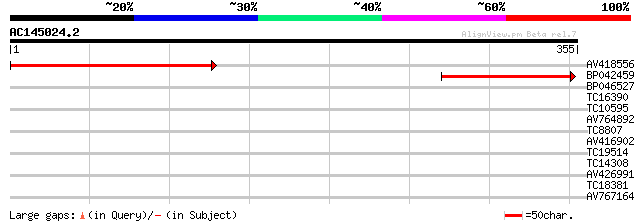
Score E
Sequences producing significant alignments: (bits) Value
AV418556 224 2e-59
BP042459 71 3e-13
BP046527 39 0.001
TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-asso... 36 0.011
TC10595 similar to UP|AAPC_PENCL (Q40784) Possible apospory-asso... 35 0.025
AV764892 27 4.0
TC8807 similar to UP|O82730 (O82730) Monogalactosyldiacylglycero... 27 5.2
AV416902 27 6.8
TC19514 weakly similar to UP|SCWB_YEAST (P53189) Probable family... 27 6.8
TC14308 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial ... 27 6.8
AV426991 26 8.9
TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, parti... 26 8.9
AV767164 26 8.9
>AV418556
Length = 415
Score = 224 bits (571), Expect = 2e-59
Identities = 115/131 (87%), Positives = 125/131 (94%), Gaps = 2/131 (1%)
Frame = +3
Query: 1 MASFLSLSLPKLNLIKASSAA-TTTTPLSTPEALNDKFGRKGIKFLES-NSIPIVELTVR 58
MASFLSLSLPKLNLIKASSAA T+TT L++PEALN+KFGRKGIKFLES N+IPIVELTVR
Sbjct: 18 MASFLSLSLPKLNLIKASSAANTSTTTLTSPEALNEKFGRKGIKFLESDNNIPIVELTVR 197
Query: 59 NGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPG 118
NGSSLSLR+PDAHVTSYKPKVFWKDDG EE+LYT+PA E+GLYKAKGGIGLVLNEVLQPG
Sbjct: 198 NGSSLSLRIPDAHVTSYKPKVFWKDDGFEEVLYTVPATESGLYKAKGGIGLVLNEVLQPG 377
Query: 119 AKELLPSTLEW 129
AK +LPSTLEW
Sbjct: 378 AKGVLPSTLEW 410
>BP042459
Length = 529
Score = 70.9 bits (172), Expect = 3e-13
Identities = 28/84 (33%), Positives = 51/84 (60%)
Frame = -1
Query: 271 KERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYFVCTGPAS 330
+++ Y P + +ID+GR + R GFE+ Y+ SPGS + Y +D ++C G A+
Sbjct: 439 RDKLSLVYTNAPRSFTVIDRGRRNSVVIGRNGFEETYLYSPGSRVESYSKDAYICVGQAA 260
Query: 331 ILEPVTVNPGEEFRGAQVIEHDNL 354
+L+P+ + P E ++G Q +++ NL
Sbjct: 259 VLQPIVLGPEEVWKGGQYLQNPNL 188
>BP046527
Length = 521
Score = 38.9 bits (89), Expect = 0.001
Identities = 25/76 (32%), Positives = 42/76 (54%), Gaps = 8/76 (10%)
Frame = -1
Query: 278 YNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG-----SMSDKYGRD---YFVCTGPA 329
Y + P+K I+D ++ + + + G D V +P SM+D +G D + VC A
Sbjct: 482 YLSTPAKISILDHEKKRSFVLRKDGLPDAVVWNPWDKKAKSMAD-FGDDEYKHMVCVEAA 306
Query: 330 SILEPVTVNPGEEFRG 345
+I +P+T+ PGEE+ G
Sbjct: 305 AIEKPITLKPGEEWNG 258
>TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-associated
protein C {Arabidopsis thaliana;}, partial (34%)
Length = 532
Score = 35.8 bits (81), Expect = 0.011
Identities = 18/66 (27%), Positives = 34/66 (51%), Gaps = 7/66 (10%)
Frame = +3
Query: 287 IIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRD-------YFVCTGPASILEPVTVNP 339
++D R+ + + + G D+ V +P K D + +C A++ +P+T+ P
Sbjct: 84 VLDHERKRTFMIRKEGLPDVVVWNPWEKKSKSVVDLGDEEYKHMLCVDGAAVGKPITLKP 263
Query: 340 GEEFRG 345
GEE+RG
Sbjct: 264 GEEWRG 281
>TC10595 similar to UP|AAPC_PENCL (Q40784) Possible apospory-associated
protein C, partial (28%)
Length = 726
Score = 34.7 bits (78), Expect = 0.025
Identities = 20/66 (30%), Positives = 36/66 (54%), Gaps = 7/66 (10%)
Frame = +3
Query: 287 IIDQGREIFYRVIRMGFEDIYVSSPGSMSDK----YGRD---YFVCTGPASILEPVTVNP 339
IID ++ + +++ G D V +P K +G D + +C A+I +P+T+ P
Sbjct: 18 IIDHEKKRTFVLLKDGLPDAVVWNPWDKKAKALADFGDDEYKHMLCVEAAAIEKPITLKP 197
Query: 340 GEEFRG 345
GEE++G
Sbjct: 198 GEEWKG 215
>AV764892
Length = 505
Score = 27.3 bits (59), Expect = 4.0
Identities = 18/60 (30%), Positives = 26/60 (43%)
Frame = -1
Query: 267 AAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYFVCT 326
AA P R F++ PSK+ I R R + F I + SMS R+ +C+
Sbjct: 283 AAQPINRAP*FHSIHPSKHSSIASKRGKIQRDREISFLCIIL*GSSSMSSPISRESVICS 104
>TC8807 similar to UP|O82730 (O82730) Monogalactosyldiacylglycerol synthase
, partial (65%)
Length = 1634
Score = 26.9 bits (58), Expect = 5.2
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +1
Query: 192 FKRRGGAAIKGLQTCSYCSHPPLDSPFQIL 221
+ +RGG G+Q Y PPLD+ + I+
Sbjct: 118 YAKRGGGRADGVQARYYNQCPPLDAAYSIV 207
>AV416902
Length = 404
Score = 26.6 bits (57), Expect = 6.8
Identities = 19/50 (38%), Positives = 25/50 (50%)
Frame = +3
Query: 47 SNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPAN 96
S+ IP+VEL R L L D HVT P + ++ LL T+P N
Sbjct: 18 SHLIPLVELARR----LVLHHHDLHVTLLIPTLVPLTASMKALLNTLPPN 155
>TC19514 weakly similar to UP|SCWB_YEAST (P53189) Probable family 17
glucosidase SCW11 precursor (Soluble cell wall protein
11) , partial (5%)
Length = 518
Score = 26.6 bits (57), Expect = 6.8
Identities = 16/44 (36%), Positives = 27/44 (61%), Gaps = 1/44 (2%)
Frame = -2
Query: 5 LSLSLPKLNLIKASSAATTTTPLSTPEAL-NDKFGRKGIKFLES 47
LSLS+ + + I + TTTT ++ + L ND F + IKF+++
Sbjct: 145 LSLSMIQTSQISHAGTTTTTTKTNSIQFLTND*FPQTNIKFMKN 14
>TC14308 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial (96%)
Length = 764
Score = 26.6 bits (57), Expect = 6.8
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = -1
Query: 185 AILSHFRFKRRGGAAIKGLQTCSYCSH 211
++LSH R GG ++GL C C H
Sbjct: 152 SLLSHRRLLLLGGLLLRGLLVCLGCHH 72
>AV426991
Length = 429
Score = 26.2 bits (56), Expect = 8.9
Identities = 18/47 (38%), Positives = 26/47 (55%)
Frame = +2
Query: 5 LSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIP 51
LSLS L+ + ++A TTTP S P + + +K I F E+ S P
Sbjct: 221 LSLSSLSLSNVDEATATGTTTP*SLPMLFSSECSKKTI-FCETVSDP 358
>TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, partial (53%)
Length = 758
Score = 26.2 bits (56), Expect = 8.9
Identities = 21/84 (25%), Positives = 42/84 (50%), Gaps = 4/84 (4%)
Frame = +1
Query: 146 LISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTN----AILSHFRFKRRGGAAIK 201
L +T RF ++I + + VS A ++ ++K+ + + N +I S+ +F+ +
Sbjct: 25 LTATPRFISTSFIXAFH-VSNARSLQSQSKNNQTLEFNNNINQSINSNLQFQFQFQ---- 189
Query: 202 GLQTCSYCSHPPLDSPFQILTPSE 225
CS+C H P+ + L+ SE
Sbjct: 190 -FHRCSFCFHDPIQQNVEPLSRSE 258
>AV767164
Length = 263
Score = 26.2 bits (56), Expect = 8.9
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = -2
Query: 345 GAQVIEHDNLS 355
GAQVIEHDNLS
Sbjct: 259 GAQVIEHDNLS 227
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,830,287
Number of Sequences: 28460
Number of extensions: 76682
Number of successful extensions: 399
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 397
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 397
length of query: 355
length of database: 4,897,600
effective HSP length: 91
effective length of query: 264
effective length of database: 2,307,740
effective search space: 609243360
effective search space used: 609243360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC145024.2


