

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.14 - phase: 0 /pseudo
(122 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
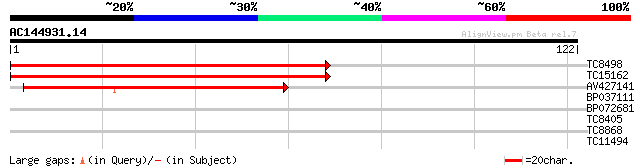
Score E
Sequences producing significant alignments: (bits) Value
TC8498 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16... 137 4e-34
TC15162 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S1... 131 3e-32
AV427141 82 2e-17
BP037111 26 2.2
BP072681 25 4.8
TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase is... 24 6.3
TC8868 similar to UP|AAR06294 (AAR06294) Adenylosuccinate syntha... 24 6.3
TC11494 similar to UP|Q7X9B2 (Q7X9B2) 9/13 hydroperoxide lyase, ... 24 8.3
>TC8498 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 693
Score = 137 bits (346), Expect = 4e-34
Identities = 64/69 (92%), Positives = 69/69 (99%)
Frame = +1
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKKNAVAVT+CKRGRGLIKINGCPIEL++PEILRFKA+EPILLLGRHRFAGV
Sbjct: 34 LEQVQCFGRKKNAVAVTYCKRGRGLIKINGCPIELIQPEILRFKAFEPILLLGRHRFAGV 213
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 214 DMRIRVKGG 240
>TC15162 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 808
Score = 131 bits (330), Expect = 3e-32
Identities = 63/69 (91%), Positives = 67/69 (96%)
Frame = +1
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
MEQVQCFGRKKNAVAVT+CKRGRGLIKING PIEL+EPEILRFKA+EPILLLG+ RFAGV
Sbjct: 64 MEQVQCFGRKKNAVAVTYCKRGRGLIKINGSPIELIEPEILRFKAFEPILLLGKSRFAGV 243
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 244 DMRIRVKGG 270
>AV427141
Length = 286
Score = 82.4 bits (202), Expect = 2e-17
Identities = 41/58 (70%), Positives = 48/58 (82%), Gaps = 1/58 (1%)
Frame = +1
Query: 4 VQCFGRKKNAVAVTHCKR-GRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
VQ FGRKK AVAV K G+GL+K+NGCPIEL+EPEILR K +EP+LLLG+ RFAGV
Sbjct: 112 VQVFGRKKTAVAVALIKGDGKGLMKVNGCPIELLEPEILRVKVFEPLLLLGQERFAGV 285
>BP037111
Length = 563
Score = 25.8 bits (55), Expect = 2.2
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = -2
Query: 108 GSGGGKRYGRTRGGV 122
G GGG+R GR GG+
Sbjct: 385 GGGGGRRVGRRNGGI 341
>BP072681
Length = 461
Score = 24.6 bits (52), Expect = 4.8
Identities = 13/35 (37%), Positives = 17/35 (48%)
Frame = +2
Query: 38 PEILRFKAYEPILLLGRHRFAGVDMRIRVKGGVTL 72
P L Y PILLL F ++R+K G +L
Sbjct: 167 PHNLHTHHYTPILLLSYASFYSPTFKLRLKEGASL 271
>TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase isolog,
25331-24357 (At1g11090), partial (18%)
Length = 552
Score = 24.3 bits (51), Expect = 6.3
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = -2
Query: 108 GSGGGKRYGRTRGG 121
GS GG+R GR RGG
Sbjct: 344 GSRGGRRSGRGRGG 303
>TC8868 similar to UP|AAR06294 (AAR06294) Adenylosuccinate synthase ,
partial (62%)
Length = 1273
Score = 24.3 bits (51), Expect = 6.3
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 4/37 (10%)
Frame = -3
Query: 20 KRGRGLIKINGCPIEL----VEPEILRFKAYEPILLL 52
+RGR ++ N CP+ L EP+ K EP L++
Sbjct: 584 RRGRPVVVPNSCPVNLRRSPPEPKFSVGKGPEPTLVV 474
>TC11494 similar to UP|Q7X9B2 (Q7X9B2) 9/13 hydroperoxide lyase, partial
(27%)
Length = 541
Score = 23.9 bits (50), Expect = 8.3
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = -1
Query: 109 SGGGKRYGRTRGG 121
S GG+R+GR RGG
Sbjct: 220 SRGGRRWGRRRGG 182
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.147 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,474,228
Number of Sequences: 28460
Number of extensions: 14119
Number of successful extensions: 73
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 72
length of query: 122
length of database: 4,897,600
effective HSP length: 79
effective length of query: 43
effective length of database: 2,649,260
effective search space: 113918180
effective search space used: 113918180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 49 (23.5 bits)
Medicago: description of AC144931.14


