

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
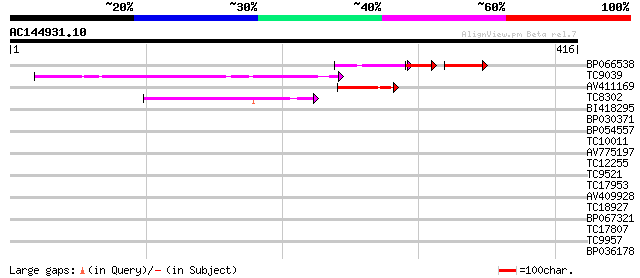
Score E
Sequences producing significant alignments: (bits) Value
BP066538 42 4e-13
TC9039 50 7e-07
AV411169 43 8e-05
TC8302 42 1e-04
BI418295 37 0.006
BP030371 36 0.013
BP054557 35 0.023
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 34 0.051
AV775197 33 0.067
TC12255 33 0.087
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 33 0.11
TC17953 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17... 31 0.33
AV409928 30 0.96
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 28 3.7
BP067321 27 4.8
TC17807 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromo... 27 4.8
TC9957 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor N... 27 4.8
BP036178 27 8.1
>BP066538
Length = 396
Score = 42.4 bits (98), Expect(3) = 4e-13
Identities = 23/31 (74%), Positives = 23/31 (74%)
Frame = -3
Query: 320 VKVKGKGVVAVETLLGIKYILDVLFVPEINQ 350
V VK K VVAVET G KYI DVL VPEINQ
Sbjct: 145 VDVKAK*VVAVETSSGTKYIPDVLLVPEINQ 53
Score = 38.1 bits (87), Expect(3) = 4e-13
Identities = 22/58 (37%), Positives = 34/58 (57%), Gaps = 1/58 (1%)
Frame = -2
Query: 239 GHVEKVCKN-KTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMT 295
GH+EK ++ K +Q E Q V QE+ E +C+ ++ E WLI+S CT+H+T
Sbjct: 389 GHIEKN*ESQKLSQFEAQ---VGDQQEEVEY*LATTCFATNNNCENWLIESRCTHHIT 225
Score = 29.6 bits (65), Expect(3) = 4e-13
Identities = 12/23 (52%), Positives = 17/23 (73%)
Frame = -1
Query: 291 TNHMTNNVSFFKELDESFYSKVV 313
++HMT + FKELD+ F SKV+
Sbjct: 234 SHHMTYDEKLFKELDQPFVSKVI 166
>TC9039
Length = 1218
Score = 50.1 bits (118), Expect = 7e-07
Identities = 53/228 (23%), Positives = 103/228 (44%), Gaps = 1/228 (0%)
Frame = +2
Query: 19 NPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEY 78
NP L + T Q +++ +L II + ++F ++ EI K A + L E
Sbjct: 248 NPLLLRKSSTSEQRKDYEKWDRSNRMSLMIIKRGI-PEVFRGTISEEI-KGAKDFLAEIE 421
Query: 79 QGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKIL 138
+ ++ K + L + ++K + +RE+ +S + ++++ L +LSD ++ +L
Sbjct: 422 KRFAKSDKAETSTLLQNLISMKYQGKGNIREYIMGMSNIASKLKALKLELSDDLLIHLVL 601
Query: 139 VCLPEMF-EAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKN 197
+ LP F + KIS + +S + EL++ E+R L+ E F++++K K
Sbjct: 602 LSLPAQFSQFKISYNCPKEKWS---LNELISFCVQEEER--LKQERKESAHFVSTSKDKG 766
Query: 198 QSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVC 245
+ K+ K K D C++ CN GH++K C
Sbjct: 767 KRKKTVEPKNEAADAPAPKKQKEDDTCYF--------CNVSGHMKKKC 886
>AV411169
Length = 424
Score = 43.1 bits (100), Expect = 8e-05
Identities = 23/45 (51%), Positives = 30/45 (66%)
Frame = +3
Query: 241 VEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWL 285
VEK C NK NQQ QQA+V E+ + DE++LF + Y A S TW+
Sbjct: 267 VEKACNNKENQQ*QQAKVAEYEEHDEKELF-VNNYNAAES-VTWI 395
>TC8302
Length = 494
Score = 42.4 bits (98), Expect = 1e-04
Identities = 27/131 (20%), Positives = 59/131 (44%), Gaps = 3/131 (2%)
Frame = +3
Query: 99 LKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNF 158
L+M E+ ++ + D ++ + Q+ + + E +L LP+ ++ + +++ N
Sbjct: 18 LRMSESTSMSDHIDHMNTLFAQLSASNFTIGENERAELLLESLPDSYDQLVINIKNNNIV 197
Query: 159 SEITVPELVNALQASEQRR---SLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCK 215
+ + + L L + +R R + + L +G+++S K K C HC
Sbjct: 198 NHLPLMMLSEVLLEEDSQR*NKEDR*DSSKPMEALTMTRGRSKSRKKINLK----CYHCG 365
Query: 216 KDTHLDKFCWY 226
+ HL K CW+
Sbjct: 366 QR*HLKKDCWF 398
>BI418295
Length = 523
Score = 37.0 bits (84), Expect = 0.006
Identities = 24/105 (22%), Positives = 55/105 (51%)
Frame = +2
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++N E + + W ++++ ++ R + K LR E + + K ++++ E+ RI ++
Sbjct: 215 VINCERSSQVWREILDFFENQCRAQSAK---LRSELKNIT-KGSKSITEYLRRIKTIVNT 382
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPE 165
+ +GE +S + +E I LPE + A + + + + S I+ E
Sbjct: 383 LVSIGEPVSFRDHLEAIFDGLPEDYSALSTIIFTHASSSTISEVE 517
>BP030371
Length = 516
Score = 35.8 bits (81), Expect = 0.013
Identities = 33/159 (20%), Positives = 62/159 (38%), Gaps = 16/159 (10%)
Frame = -2
Query: 106 TVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPE 165
TV E+ I + R +G+ +S + E IL P +E+ ++ + + F +++ E
Sbjct: 512 TVAEYLM*IKSITDPPRSIGDPVSAKEQAEIILEGFPTKYESLVNLIHMSDKFQALSLDE 333
Query: 166 LVNALQASEQR----RSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLD 221
+ + L+ E R R E V N+G + + + P P + D
Sbjct: 332 IESLLRDQEARIECYRKFLTEI*VNLT*TLENQGSSSA------NTYAPAPVHSAEASHD 171
Query: 222 KFCWYRP------------GVKCRACNQLGHVEKVCKNK 248
+ + GV+C+ C GH VC ++
Sbjct: 170 GYSNFNGKRCGGKG*GRGYGVQCQVCLPTGHTASVCYHR 54
>BP054557
Length = 538
Score = 35.0 bits (79), Expect = 0.023
Identities = 27/93 (29%), Positives = 45/93 (48%), Gaps = 1/93 (1%)
Frame = -2
Query: 285 LIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYIL-DVL 343
L+D+G T+H+ +N+ F+ L + + NGQ K +A K+ L VL
Sbjct: 537 LLDTGATHHVCHNLDVFQALRQVHPVLLSMPNGQ----KEIAHMADSVYFSEKFFLAXVL 370
Query: 344 FVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFD 376
+VP N +L+SV +++ N + IFD
Sbjct: 369 YVPSFNFNLISVSALVKS------LNCRLVIFD 289
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 33.9 bits (76), Expect = 0.051
Identities = 31/118 (26%), Positives = 51/118 (42%), Gaps = 10/118 (8%)
Frame = +2
Query: 148 KISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASN--KGKNQSFKSFGE 205
++ ++ NF+ + E + A + R +L + VEG+ + KG + + +
Sbjct: 176 RVRGVDMKHNFAFV---EFSDPRDADDARYNLDGRD-VEGSRIVVEFAKGGPRGNREYLG 343
Query: 206 KKFPP----CPHCKKDTHLDKFC----WYRPGVKCRACNQLGHVEKVCKNKTNQQEQQ 255
+ PP C +C D H + C W KC C GHVE+ CKN + E Q
Sbjct: 344 RGPPPGSGRCFNCGLDGHWARDCKAGDWKN---KCYRCGDRGHVERNCKNSPKKNEWQ 508
>AV775197
Length = 439
Score = 33.5 bits (75), Expect = 0.067
Identities = 21/59 (35%), Positives = 29/59 (48%), Gaps = 5/59 (8%)
Frame = +1
Query: 240 HVEKVCKNKTNQQEQQARV-----VEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNH 293
H+ KV + ++ QARV ++ HQE E+Q CY S +TWL S C H
Sbjct: 256 HIHKVARVVVSE*VNQARVEQDHHIQ*HQEQEDQ----DCYYHQSVTQTWL*SSVCFPH 420
>TC12255
Length = 558
Score = 33.1 bits (74), Expect = 0.087
Identities = 16/56 (28%), Positives = 30/56 (53%), Gaps = 1/56 (1%)
Frame = +3
Query: 272 ASCYLACSSKETWLIDSGCTNHMTNNVSFFKE-LDESFYSKVVFGNGQHVKVKGKG 326
+S + + W DSG T+H+TNN F + + + +++ GNGQ + ++ G
Sbjct: 15 SSTQFTAAPQVNWCPDSGATHHVTNNPGIFVDNVPLATQDQLLVGNGQGLPIQSIG 182
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 32.7 bits (73), Expect = 0.11
Identities = 17/51 (33%), Positives = 23/51 (44%), Gaps = 4/51 (7%)
Frame = +1
Query: 211 CPHCKKDTHLDKFC----WYRPGVKCRACNQLGHVEKVCKNKTNQQEQQAR 257
C +C D H + C W KC C + GH+EK CKN + + R
Sbjct: 391 CFNCGIDGHWARDCKAGDWKN---KCYRCGERGHIEKNCKNSPKKLSRHPR 534
>TC17953 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17A9_9
{Arabidopsis thaliana;}, partial (6%)
Length = 527
Score = 31.2 bits (69), Expect = 0.33
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +3
Query: 230 VKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQED 265
++C C +LGH +K CKN EQ + +H D
Sbjct: 117 LQCSKCKELGHNKKTCKNSL-PLEQDT*AISNHSRD 221
>AV409928
Length = 414
Score = 29.6 bits (65), Expect = 0.96
Identities = 19/49 (38%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Frame = +2
Query: 92 LRREFEALKMKETETVREFSDRISKVIT--QIRLLGEDLSDQRVVEKIL 138
LR E + E E R +R+ K+ T IRL+GE L + V EKI+
Sbjct: 26 LREELRQMTAPEQEMERRDKERLLKLRTLGNIRLIGELLKQKMVPEKIV 172
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 27.7 bits (60), Expect = 3.7
Identities = 13/37 (35%), Positives = 15/37 (40%)
Frame = -2
Query: 211 CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKN 247
C C K H C P + C C + GH K C N
Sbjct: 329 CHRCSKKGHFANRC---PDLVCWNCQKTGHSGKDCTN 228
>BP067321
Length = 513
Score = 27.3 bits (59), Expect = 4.8
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = +3
Query: 15 EKGNNPPPLPNNPTLAQIRN 34
+K NPPPLP NP L N
Sbjct: 150 KKSQNPPPLPPNPGLKHHHN 209
>TC17807 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromosome 5, TAC
clone:K17O22 (AT5g45100/K17O22_9), partial (26%)
Length = 844
Score = 27.3 bits (59), Expect = 4.8
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 5/58 (8%)
Frame = -3
Query: 231 KCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQED-----EEQLFKASCYLACSSKET 283
K + ++ ++V NQQ + RV+ HH E EE + S LAC +ET
Sbjct: 431 KNKLSSESSFFDQVLYQFQNQQSEIDRVLAHHTEKVRMEMEEHKMRQSRMLACVIQET 258
>TC9957 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor Nt-SubD48,
partial (16%)
Length = 653
Score = 27.3 bits (59), Expect = 4.8
Identities = 14/45 (31%), Positives = 24/45 (53%), Gaps = 5/45 (11%)
Frame = -2
Query: 351 SLLSVGQMMEKNYSL-----HFKNMKCTIFDPYGSKLMIVEMRGK 390
S+ S G ++N+ L + +N + D YGSK+ I+ RG+
Sbjct: 343 SMASSGSTQKRNHPLELGNCNIRNSSAIVLDFYGSKVGIMRCRGR 209
>BP036178
Length = 561
Score = 26.6 bits (57), Expect = 8.1
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = -2
Query: 209 PPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVC 245
PP KK ++ F + V+C C+ LGH +K C
Sbjct: 428 PPGRPKKKVLRVENFKRPKRIVQCGRCHMLGHSQKKC 318
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,330,741
Number of Sequences: 28460
Number of extensions: 102063
Number of successful extensions: 603
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 599
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 416
length of database: 4,897,600
effective HSP length: 93
effective length of query: 323
effective length of database: 2,250,820
effective search space: 727014860
effective search space used: 727014860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144931.10


