

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
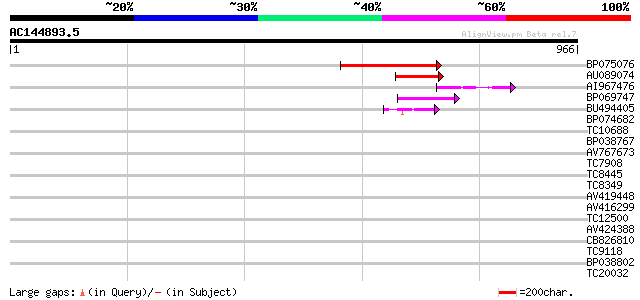
BP075076 250 9e-67
AU089074 122 3e-28
AI967476 55 4e-08
BP069747 46 2e-05
BU494405 45 4e-05
BP074682 38 0.007
TC10688 similar to UP|AAR10812 (AAR10812) Superoxide dismutase, ... 36 0.033
BP038767 35 0.074
AV767673 33 0.28
TC7908 27 0.35
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 32 0.37
TC8349 32 0.63
AV419448 31 0.82
AV416299 31 0.82
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 31 1.1
AV424388 31 1.1
CB826810 30 1.4
TC9118 similar to UP|Q84JE4 (Q84JE4) Squamosa promoter binding l... 30 1.4
BP038802 30 1.4
TC20032 30 1.8
>BP075076
Length = 527
Score = 250 bits (638), Expect = 9e-67
Identities = 121/172 (70%), Positives = 134/172 (77%)
Frame = -1
Query: 564 AHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNSEFLLANDK 623
AHSE S GV R TRPLH LS P+ +N++ S+ +EKEKR PKAN FYHNSEFLLA DK
Sbjct: 527 AHSELASAGVPRPITRPLHHLSLPLLENNQGVSDNLEKEKRTPKANHFYHNSEFLLAKDK 348
Query: 624 FPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTPVDVDG 683
FPPAESNKKSKL+WKKQG G+ G GL+MGSKFFKSCSSLLEKLM+H WVFNTPVDV+G
Sbjct: 347 FPPAESNKKSKLHWKKQGAGEFGHGLRMGSKFFKSCSSLLEKLMKHSLGWVFNTPVDVEG 168
Query: 684 LGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKG 735
LGLHDYFTIIT+PMDLGTVKTRLNKN F AMTYNP+G
Sbjct: 167 LGLHDYFTIITHPMDLGTVKTRLNKNLVHIT*GICRGCETHFSYAMTYNPEG 12
>AU089074
Length = 244
Score = 122 bits (306), Expect = 3e-28
Identities = 53/81 (65%), Positives = 66/81 (81%)
Frame = +1
Query: 658 SCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEF 717
SC +L+KLM+H++ W+FN PVD G+GLHDY+ II PMDLGTVK L+KN Y +P+EF
Sbjct: 1 SCVQVLQKLMKHKHGWIFNVPVDAVGMGLHDYYEIIKKPMDLGTVKANLSKNVYATPEEF 180
Query: 718 AEDVRLTFHNAMTYNPKGQDV 738
A DVRLTF+NA+TYNPKG DV
Sbjct: 181 ASDVRLTFNNALTYNPKGHDV 243
>AI967476
Length = 397
Score = 55.5 bits (132), Expect = 4e-08
Identities = 42/135 (31%), Positives = 60/135 (44%)
Frame = +3
Query: 727 NAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPP 786
NAMTYNP+G DVH MA+ LSK FE RW IE R+ + S V P
Sbjct: 36 NAMTYNPRGNDVHIMADTLSKYFEVRWKSIEKKLPRKDAQRLP--TKPATSDDVETIRPV 209
Query: 787 PPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHS 846
PP R+++ S +P +P+ P K+ M+ EK L L S
Sbjct: 210 PPSKKRKLI------------------SLSP---QPEVMPPAKQVMSDQEKLCLRRELES 326
Query: 847 LPVEKLDAVVQMMRK 861
L E+ ++ +++
Sbjct: 327 LLEEEPVHIIDFLKE 371
>BP069747
Length = 436
Score = 46.2 bits (108), Expect = 2e-05
Identities = 33/107 (30%), Positives = 52/107 (47%), Gaps = 3/107 (2%)
Frame = -1
Query: 662 LLEKLMEHQYAWVFNTPVD-VDGLGLHDYFTIITNPMDLGTVKTRLN-KNW-YKSPKEFA 718
+L+++M + W F P+D D L P+ LG V KNW Y P++F
Sbjct: 433 ILKRMMIGRDGWAFKHPLDPFDSESLLMEKNKKKKPI-LGFVDIESKLKNWLYSEPEQFV 257
Query: 719 EDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMR 765
+D+R F +A+ Y P +VH + +LS FE W ++ + E R
Sbjct: 256 DDMRNVFSHALMY-PSRSEVHIIGRRLSDNFEHNWTTLKKKWASEDR 119
>BU494405
Length = 563
Score = 45.4 bits (106), Expect = 4e-05
Identities = 29/101 (28%), Positives = 49/101 (47%), Gaps = 6/101 (5%)
Frame = +1
Query: 638 KKQGGGDMGXGLQMGSKFFKSCSSLLEKLME------HQYAWVFNTPVDVDGLGLHDYFT 691
++ GGG++G +++LE +++ H +++F PV DYF
Sbjct: 169 RRGGGGEVGL------------ANILEGIVDVLVKDRHDLSYLFQKPVSKKEAP--DYFD 306
Query: 692 IITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYN 732
II PMDL ++ R+ YKS ++F D+ +NA YN
Sbjct: 307 IIERPMDLSRIRERVRNMEYKSREDFRHDIWQITYNAHKYN 429
>BP074682
Length = 497
Score = 38.1 bits (87), Expect = 0.007
Identities = 22/40 (55%), Positives = 26/40 (65%), Gaps = 3/40 (7%)
Frame = +2
Query: 930 QMKEMFPHPCLCKGEVKQII---RVDQVVQAVILGLLQVV 966
Q KEMFP PCLC + K I+ +V QV QAV L L QV+
Sbjct: 83 QKKEMFPPPCLCMEKFKWIMGARQVVQVAQAVTLVLHQVI 202
>TC10688 similar to UP|AAR10812 (AAR10812) Superoxide dismutase, partial
(91%)
Length = 822
Score = 35.8 bits (81), Expect = 0.033
Identities = 27/97 (27%), Positives = 38/97 (38%)
Frame = +2
Query: 753 WAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTP 812
W S ++ + D+ +P P S P P P P +P +P
Sbjct: 53 WRPTRSSHSLLTHHSSDHPSPVPPSNSPPTSEPSHP---------PNPSPSSPPPRKPSP 205
Query: 813 SSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPV 849
SSR P P K + P K T D++ + SL SL V
Sbjct: 206 SSRAPPPSKASSPSPKK---TTDQQL*MLESLVSLQV 307
>BP038767
Length = 561
Score = 34.7 bits (78), Expect = 0.074
Identities = 21/59 (35%), Positives = 29/59 (48%), Gaps = 1/59 (1%)
Frame = +2
Query: 773 PSPLSRRVPAFTPPPPLDMRRILDRQEPFART-PRSMNNTPSSRTPAPKKPKAKDPNKR 830
P LS +P PPL + + Q P R+ P + NN S +P PKKP + P+ R
Sbjct: 191 PLSLSSTMPGLVKTPPLRIT-MPQTQTPTRRSEPAATNNKTPSPSPLPKKPPSPSPSTR 364
>AV767673
Length = 497
Score = 32.7 bits (73), Expect = 0.28
Identities = 19/83 (22%), Positives = 35/83 (41%)
Frame = -1
Query: 272 AQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRP 331
A P + + S+ + +A+ +S+ S STQK +P++ S P
Sbjct: 446 AAPRAADPSSSTSSSSLNQHQKEIAEELSTSQVPKATSSLASASTQKSNLTPELTNQSAP 267
Query: 332 EDASLTQPEVNSRLEDRSFPQPE 354
+A Q E +S + + P P+
Sbjct: 266 TEAEPMQIEASSSANENANPPPK 198
>TC7908
Length = 1257
Score = 26.9 bits (58), Expect(2) = 0.35
Identities = 26/88 (29%), Positives = 38/88 (42%), Gaps = 8/88 (9%)
Frame = +3
Query: 354 ELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPV--------ANSTSEDLNLA 405
EL S TSQ Q+NS+ E+ S P L +R S PV T+ A
Sbjct: 483 ELMSLATKSTSQSQNNSVQENQPSFAPSL-IRTSADSSAPVIPGVNIIPCTGTNSIPEHA 659
Query: 406 QPLSQSVSDDLHINQQAEPSNLNVRQED 433
Q S+ + DL I ++A +++D
Sbjct: 660 QVSSRPIVCDLPIARKASLHRFLEKRKD 743
Score = 23.9 bits (50), Expect(2) = 0.35
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +1
Query: 438 SPIHRQGAISDNLHSHQQVEPSNP 461
SP H G + D+L S + PSNP
Sbjct: 808 SPCHGLGLVPDDLKSKFKHWPSNP 879
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 32.3 bits (72), Expect = 0.37
Identities = 35/126 (27%), Positives = 49/126 (38%)
Frame = +1
Query: 129 QPHASFQLADGNSPQQKVENQDSAHPQASLRIGDGNSPQQQLVEPNLAQPQASLRAGDGN 188
QPH Q PQ Q HPQ L+ + PQ Q ++ + QPQ+ +
Sbjct: 49 QPHLQHQ------PQHHPHFQP--HPQPQLQ----SQPQPQHIQQSQPQPQSQPQHIQQP 192
Query: 189 SPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLV 248
PQ QL+ +QPQ + H Q Y PP + G N + L
Sbjct: 193 QPQPQLQHIQQSQPQPQFQN-----QHFQYPGTY-PPPPWAATPGYPNYQNHLSSTNALS 354
Query: 249 QPQASS 254
PQA++
Sbjct: 355 TPQANT 372
>TC8349
Length = 749
Score = 31.6 bits (70), Expect = 0.63
Identities = 19/61 (31%), Positives = 26/61 (42%)
Frame = +2
Query: 538 QGYVGGYGNSNVVLGGGITNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASE 597
Q G +GN N+ GG+ GG R + +V S P P SF NS ++
Sbjct: 137 QSLGGSHGNGNMAKNGGVGFGGQTLRVAGDTSNVSGSNGPV-PSRSNSFKAASNSDSSAA 313
Query: 598 G 598
G
Sbjct: 314 G 316
>AV419448
Length = 382
Score = 31.2 bits (69), Expect = 0.82
Identities = 17/57 (29%), Positives = 23/57 (39%)
Frame = +2
Query: 772 APSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPN 828
AP P S P PPL L + +P ++N +S P P P K P+
Sbjct: 143 APPPSSPTATPSAPTPPLSAALSLASTATSSPSPSPISNPSASLPPTPPPPPTKTPS 313
>AV416299
Length = 403
Score = 31.2 bits (69), Expect = 0.82
Identities = 19/64 (29%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +3
Query: 855 VVQMMRKKNL*LNQQDDEIEVDFDAVD-AEILWELDRFVLNYKKSLSKNKRKAEQARERA 913
V+ RK NL + + +++ D D D EI E +++LN + LS K++ + + +
Sbjct: 42 VIDFGRKSNL--SSKVEKVHEDVDIQDLGEISSEAQQYILNLQSRLSSMKKELHEVKRKN 215
Query: 914 EALQ 917
ALQ
Sbjct: 216 AALQ 227
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 30.8 bits (68), Expect = 1.1
Identities = 20/56 (35%), Positives = 25/56 (43%), Gaps = 1/56 (1%)
Frame = +1
Query: 773 PSPLSRRVPAFTPPPPLDMRRILDRQEPFARTP-RSMNNTPSSRTPAPKKPKAKDP 827
PSP + A PPPPL M I P TP S++ P + T +P P P
Sbjct: 505 PSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDAPPPTPTLSPPPPPPPTP 672
>AV424388
Length = 305
Score = 30.8 bits (68), Expect = 1.1
Identities = 16/44 (36%), Positives = 23/44 (51%)
Frame = +2
Query: 272 AQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNS 315
AQP + R DDGD +Q+Q+ L + SR +N P N+
Sbjct: 164 AQPNFAQRDDDGDDSQEQI----LVEGRAKSRAAILNRPSSLNA 283
>CB826810
Length = 603
Score = 30.4 bits (67), Expect = 1.4
Identities = 24/93 (25%), Positives = 37/93 (38%), Gaps = 3/93 (3%)
Frame = +2
Query: 735 GQDVHAMAEQLSKIFEDRWAI--IESDYNREMRYGMDYGAPSPLSR-RVPAFTPPPPLDM 791
G+ H IF + +I I D+ RE + + P+P + + P F PP
Sbjct: 266 GKQRHGETNSFIPIFREPSSIEKIFGDFEREQQQILTIRPPTPPEQPKTPPFVPP----- 430
Query: 792 RRILDRQEPFARTPRSMNNTPSSRTPAPKKPKA 824
R + P + P +P S +P PKA
Sbjct: 431 -RAASPRAPSPKAPSPRAASPRSPSPRVTSPKA 526
>TC9118 similar to UP|Q84JE4 (Q84JE4) Squamosa promoter binding
like-protein, partial (16%)
Length = 779
Score = 30.4 bits (67), Expect = 1.4
Identities = 34/179 (18%), Positives = 64/179 (34%), Gaps = 3/179 (1%)
Frame = +2
Query: 346 EDRSFPQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLA 405
E+ + E K ED+ ++ SS + +G + +S+S+D
Sbjct: 20 EEEEEEEEEEEEKREDVRDHKRSRKKHRSHRSSHTSRDRDRDGRKHKR-RHSSSKDKYSY 196
Query: 406 QPLSQSVSDDLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLL 465
+ +SDD H + + E R S R+ ++SD+ H H
Sbjct: 197 RGAKHGISDDEHQHSSHHHKYDSSSDEKHRSSRRRQREDSMSDHEHKH------------ 340
Query: 466 EDDGPSSPIHRHKAVPSTEYRHSENVTE---EPNLEDRIKINLASTSKQEKQEMCLKLE 521
S H+H+ E RH T+ ++E ++ K +K + ++E
Sbjct: 341 ------SRRHKHETSSEDERRHRSRHTKHKSRSHVERESELEEGEVMKSDKSQQASEVE 499
>BP038802
Length = 533
Score = 30.4 bits (67), Expect = 1.4
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 5/78 (6%)
Frame = +2
Query: 767 GMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPS----SRTPAPKKP 822
G+DY + PL P PPPP +++ +P A P +M T + S T + P
Sbjct: 182 GLDYFSHHPLPLTPPPLFPPPPPP*LKLI--TQPTAAAPATMTTTAAMTVLSST*SSLFP 355
Query: 823 -KAKDPNKRDMTYDEKQK 839
K NK +T +K+K
Sbjct: 356 MNQKTVNKTQITPTQKKK 409
>TC20032
Length = 655
Score = 30.0 bits (66), Expect = 1.8
Identities = 20/53 (37%), Positives = 25/53 (46%), Gaps = 3/53 (5%)
Frame = -3
Query: 773 PSPLSRRVPAFTPPP-PLDMRRILDRQEPFARTP--RSMNNTPSSRTPAPKKP 822
PSP S +PPP P RRI R P TP + TP++ +P P P
Sbjct: 311 PSPPSIHPNHSSPPPCPSSTRRISPRTYPPTPTPPGHGSSETPTTPSPPPSLP 153
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,305,897
Number of Sequences: 28460
Number of extensions: 200083
Number of successful extensions: 1331
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 1266
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1324
length of query: 966
length of database: 4,897,600
effective HSP length: 99
effective length of query: 867
effective length of database: 2,080,060
effective search space: 1803412020
effective search space used: 1803412020
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144893.5


