

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
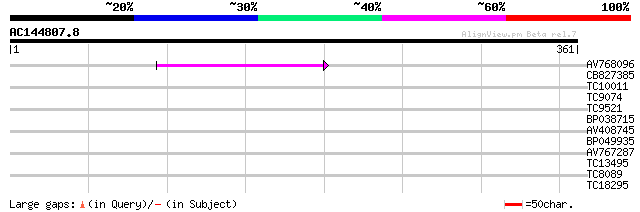
Query= AC144807.8 - phase: 0
(361 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV768096 47 4e-06
CB827385 31 0.28
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 28 2.4
TC9074 similar to UP|WR53_ARATH (Q9SUP6) Probable WRKY transcrip... 28 3.1
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 28 3.1
BP038715 27 5.3
AV408745 27 7.0
BP049935 27 7.0
AV767287 27 7.0
TC13495 similar to UP|Q9SYH5 (Q9SYH5) F15I1.27, partial (7%) 27 7.0
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 27 7.0
TC18295 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30... 26 9.1
>AV768096
Length = 358
Score = 47.4 bits (111), Expect = 4e-06
Identities = 24/110 (21%), Positives = 46/110 (41%)
Frame = +3
Query: 94 KAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIV 153
K +++ +++ + W N + +V + F F + RI+ PW + L +
Sbjct: 12 KVLNKLAVKQTIVRAWGNLQDLQVGDLNLNTFLFTFSDANLATRILAEGPWNIMGNLLSL 191
Query: 154 HPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASE 203
W I + F P W Q GLP + + K+ +++G +L E
Sbjct: 192 QFWDPQISMAEVHFNLVPFWTQFHGLPMEFMSLTNVSKLAAAVGNLLEIE 341
>CB827385
Length = 529
Score = 31.2 bits (69), Expect = 0.28
Identities = 16/44 (36%), Positives = 25/44 (56%)
Frame = -3
Query: 6 PNNPYPTQNSTQINNKLMKNYLCSLPLMKMHHQNAITSQDPAIE 49
P +P PT+NST + + SLPL + HH +A+ + P +E
Sbjct: 521 PPSP-PTENSTSPEPHRVPSLFLSLPLPR*HHSSALQTLHPPLE 393
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 28.1 bits (61), Expect = 2.4
Identities = 16/44 (36%), Positives = 20/44 (45%), Gaps = 1/44 (2%)
Frame = +2
Query: 225 PIKAGIYIGSAKDGAHWI-DFRYENLPQFCFACGLIGHTEANCK 267
P +G DG HW D + + C+ CG GH E NCK
Sbjct: 353 PPGSGRCFNCGLDG-HWARDCKAGDWKNKCYRCGDRGHVERNCK 481
Score = 26.6 bits (57), Expect = 7.0
Identities = 10/18 (55%), Positives = 11/18 (60%)
Frame = +2
Query: 253 CFACGLIGHTEANCKLHD 270
CF CGL GH +CK D
Sbjct: 371 CFNCGLDGHWARDCKAGD 424
>TC9074 similar to UP|WR53_ARATH (Q9SUP6) Probable WRKY transcription
factor 53 (WRKY DNA-binding protein 53), partial (15%)
Length = 1044
Score = 27.7 bits (60), Expect = 3.1
Identities = 16/73 (21%), Positives = 29/73 (38%)
Frame = +3
Query: 242 IDFRYENLPQFCFACGLIGHTEANCKLHDTRTNAESRNKNVLGPWLRYNHFGRRIMDYKD 301
+D++ + P FCF+ IG + + + ES + + P + D+
Sbjct: 318 LDYKEDTFPSFCFSSPSIGSEQEDNNIFSENNFIESFSPPYISP--ATSELVLHTSDFDV 491
Query: 302 RKFSSNPTKCKNY 314
+ S PT NY
Sbjct: 492 TEIISTPTSVTNY 530
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 27.7 bits (60), Expect = 3.1
Identities = 12/29 (41%), Positives = 15/29 (51%), Gaps = 1/29 (3%)
Frame = +1
Query: 240 HWI-DFRYENLPQFCFACGLIGHTEANCK 267
HW D + + C+ CG GH E NCK
Sbjct: 415 HWARDCKAGDWKNKCYRCGERGHIEKNCK 501
>BP038715
Length = 530
Score = 26.9 bits (58), Expect = 5.3
Identities = 12/43 (27%), Positives = 19/43 (43%)
Frame = -3
Query: 82 CQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKL 124
C+ + G + + + N C P G RV HIGD++
Sbjct: 159 CEEFVCGVVTTLGVVGEADSTPTGVNWCCLPSGLRVSHIGDEI 31
>AV408745
Length = 292
Score = 26.6 bits (57), Expect = 7.0
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = -1
Query: 150 WLIVHPWRRDIDARSLEFR 168
WL+VH W +D+ S+ FR
Sbjct: 100 WLLVHSWLKDLTKYSIFFR 44
>BP049935
Length = 438
Score = 26.6 bits (57), Expect = 7.0
Identities = 10/31 (32%), Positives = 15/31 (48%)
Frame = -2
Query: 236 KDGAHWIDFRYENLPQFCFACGLIGHTEANC 266
KDG +F + L F CG++ H +C
Sbjct: 359 KDGLWHCEFMFNGLSAVLFVCGMLLHQTQSC 267
>AV767287
Length = 547
Score = 26.6 bits (57), Expect = 7.0
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = -3
Query: 25 NYLCSLPLMKMHHQNAITSQDPAIENDPTEETVTE 59
N+LCS +K H+Q + S+D P ET++E
Sbjct: 485 NFLCSTLELKKHYQQLVRSKDQVW--SPKLETISE 387
>TC13495 similar to UP|Q9SYH5 (Q9SYH5) F15I1.27, partial (7%)
Length = 548
Score = 26.6 bits (57), Expect = 7.0
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = -2
Query: 316 HFMPPIPPKMLEEMAKMSMLNHIKDHPQNH 345
HF P +PP +M ++++H+KD H
Sbjct: 484 HFSPGLPPGAQ*DMLLNTLVDHVKDSHHYH 395
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 26.6 bits (57), Expect = 7.0
Identities = 16/58 (27%), Positives = 25/58 (42%)
Frame = +3
Query: 289 YNHFGRRIMDYKDRKFSSNPTKCKNYGHFMPPIPPKMLEEMAKMSMLNHIKDHPQNHN 346
+NH RR R P H +PP PP++L + + +H + P+ HN
Sbjct: 327 HNHRLRRQQRRHGRPPLKTPINHHRQEHPLPPRPPRLLRR--QEAPPDHQRHRPRRHN 494
>TC18295 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (29%)
Length = 561
Score = 26.2 bits (56), Expect = 9.1
Identities = 12/37 (32%), Positives = 17/37 (45%), Gaps = 5/37 (13%)
Frame = -2
Query: 97 HRNSIQNALSNIWCNPK-----GFRVEHIGDKLFHFF 128
H S+ L N WC PK ++VE + + H F
Sbjct: 146 HNKSLTKPLPNFWCLPKFLLLFYWKVEEVLNTFLHIF 36
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,526,465
Number of Sequences: 28460
Number of extensions: 117174
Number of successful extensions: 680
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 670
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 680
length of query: 361
length of database: 4,897,600
effective HSP length: 91
effective length of query: 270
effective length of database: 2,307,740
effective search space: 623089800
effective search space used: 623089800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144807.8


