

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.5 + phase: 0
(112 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
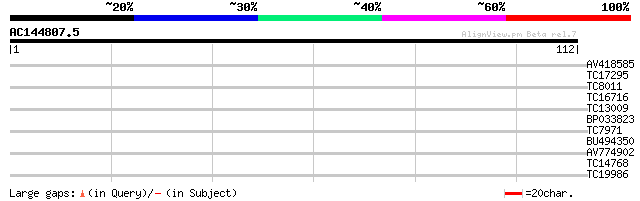
Score E
Sequences producing significant alignments: (bits) Value
AV418585 27 0.38
TC17295 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, parti... 26 0.84
TC8011 similar to UP|TPIC_FRAAN (Q9M4S8) Triosephosphate isomera... 26 0.84
TC16716 similar to GB|AAP37657.1|30725270|BT008298 At5g05630 {Ar... 25 1.4
TC13009 similar to GB|AAL09709.1|15982737|AY057475 AT4g23750/F9D... 24 3.2
BP033823 24 4.2
TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported... 23 5.4
BU494350 23 7.1
AV774902 23 7.1
TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 23 9.3
TC19986 23 9.3
>AV418585
Length = 414
Score = 27.3 bits (59), Expect = 0.38
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -2
Query: 49 EKDSLTKTNKSIRNTLWDWK 68
E+D++ KTNK + LW W+
Sbjct: 278 EEDTVMKTNKQVYRCLWYWR 219
>TC17295 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, partial (28%)
Length = 566
Score = 26.2 bits (56), Expect = 0.84
Identities = 16/38 (42%), Positives = 20/38 (52%), Gaps = 3/38 (7%)
Frame = +3
Query: 24 GLGAVFAILGARSYNQ---QKIIEALEAEKDSLTKTNK 58
GLG+ AILG Y+Q K + +E EK S K K
Sbjct: 297 GLGSAIAILGTFLYSQATSAKTAKKIEGEKSS*RKRRK 410
>TC8011 similar to UP|TPIC_FRAAN (Q9M4S8) Triosephosphate isomerase,
chloroplast precursor (TIM) , partial (89%)
Length = 1305
Score = 26.2 bits (56), Expect = 0.84
Identities = 14/31 (45%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Frame = +2
Query: 81 PVPLARLYEIYGEAAP-PQQSAPGNLFFSLL 110
P+P AR + P P QS+P NLFF L
Sbjct: 137 PIPQARFSQFSIHPFPLPFQSSPSNLFFQTL 229
>TC16716 similar to GB|AAP37657.1|30725270|BT008298 At5g05630 {Arabidopsis
thaliana;}, partial (28%)
Length = 580
Score = 25.4 bits (54), Expect = 1.4
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = -3
Query: 43 IEALEAEKDSLTKTNKSIRNTLWDWKQQ 70
I + E + +N+S+RNT ++W+QQ
Sbjct: 368 ITTIFREHGTHLSSNQSLRNTPYEWEQQ 285
>TC13009 similar to GB|AAL09709.1|15982737|AY057475 AT4g23750/F9D16_220
{Arabidopsis thaliana;}, partial (5%)
Length = 642
Score = 24.3 bits (51), Expect = 3.2
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -3
Query: 40 QKIIEALEAEKDSLTKTNKSIRNTLWDWKQ 69
Q IIEA+ + ++ L K + + + +W+WKQ
Sbjct: 235 QHIIEAVVSVQNIL-KLDVKLHHIIWNWKQ 149
>BP033823
Length = 529
Score = 23.9 bits (50), Expect = 4.2
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = -3
Query: 86 RLYEIYGEAAPPQQSAPGNLFFSLLLI 112
RLY +G PP P LF LL++
Sbjct: 473 RLYRFFGRLFPPSFE*PFQLFRVLLIL 393
>TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(98%)
Length = 1662
Score = 23.5 bits (49), Expect = 5.4
Identities = 12/41 (29%), Positives = 21/41 (50%), Gaps = 7/41 (17%)
Frame = +3
Query: 61 RNTLWD-------WKQQLYAEAASDSAPVPLARLYEIYGEA 94
+ +LWD ++ + YAEA + P+A +YG+A
Sbjct: 1110 KTSLWDLLESTLTYQHRTYAEAVKKAIAKPIAS*GGLYGDA 1232
>BU494350
Length = 537
Score = 23.1 bits (48), Expect = 7.1
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = +3
Query: 71 LYAEAASDSAPVPLA 85
L +E+ASDS PVP A
Sbjct: 459 LKSESASDSTPVPAA 503
>AV774902
Length = 594
Score = 23.1 bits (48), Expect = 7.1
Identities = 7/10 (70%), Positives = 8/10 (80%)
Frame = +3
Query: 64 LWDWKQQLYA 73
LWDW QLY+
Sbjct: 453 LWDWASQLYS 482
>TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (21%)
Length = 1036
Score = 22.7 bits (47), Expect = 9.3
Identities = 13/32 (40%), Positives = 16/32 (49%), Gaps = 3/32 (9%)
Frame = +1
Query: 59 SIRNTLWDWKQQL---YAEAASDSAPVPLARL 87
SIRN W W Q L + + S P+ L RL
Sbjct: 169 SIRNEAWIWSQWLGHPLSCSIGKSRPILLCRL 264
>TC19986
Length = 531
Score = 22.7 bits (47), Expect = 9.3
Identities = 9/25 (36%), Positives = 14/25 (56%)
Frame = -2
Query: 60 IRNTLWDWKQQLYAEAASDSAPVPL 84
+RN W WK+ L E+ S + + L
Sbjct: 98 VRNNSWVWKRLLRKESFSKAEGIKL 24
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.131 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,605,589
Number of Sequences: 28460
Number of extensions: 14808
Number of successful extensions: 87
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 87
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 87
length of query: 112
length of database: 4,897,600
effective HSP length: 88
effective length of query: 24
effective length of database: 2,393,120
effective search space: 57434880
effective search space used: 57434880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 47 (22.7 bits)
Medicago: description of AC144807.5


