

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
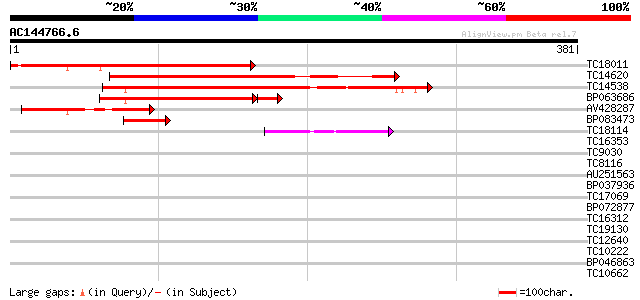
Query= AC144766.6 - phase: 0
(381 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18011 247 2e-66
TC14620 207 2e-54
TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization ... 199 6e-52
BP063686 164 3e-44
AV428287 103 5e-23
BP083473 59 2e-09
TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D... 50 5e-07
TC16353 weakly similar to UP|HQGT_ARATH (Q9M156) Probable hydroq... 37 0.007
TC9030 similar to GB|AAM63479.1|21554372|AY086477 phospholipase-... 36 0.009
TC8116 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), partia... 34 0.046
AU251563 33 0.079
BP037936 33 0.10
TC17069 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), parti... 32 0.13
BP072877 31 0.30
TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal pro... 31 0.39
TC19130 similar to UP|Q7YSZ8 (Q7YSZ8) GE-rich protein precursor,... 31 0.39
TC12640 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase prec... 30 0.51
TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 30 0.67
BP046863 30 0.87
TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CW... 30 0.87
>TC18011
Length = 542
Score = 247 bits (631), Expect = 2e-66
Identities = 114/175 (65%), Positives = 138/175 (78%), Gaps = 10/175 (5%)
Frame = +3
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSND-------DYSQFYMQEALSAFQH 53
+M+GE R NPTVHVPPW ++PTAEI+SP + D ++S YM EAL+A Q
Sbjct: 21 LMIGEH-RRANPTVHVPPWTIHDDPTAEIYSPYFAADGIAVDGGEFSPQYMHEALAALQR 197
Query: 54 YVNENN---DSDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGE 110
Y+ N +SDSD+ + VD+YS DHFRM+EFK+RRCARGRSHDWTECP++HPGE
Sbjct: 198 YLPSNEVDAESDSDALGRDSDAPVDAYSCDHFRMYEFKVRRCARGRSHDWTECPYAHPGE 377
Query: 111 KARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDG 165
KARRRDPRK++YSGT+CPDFRKGSCKK D+CE+AHGVFECWLHP+RYRTQPCKDG
Sbjct: 378 KARRRDPRKFHYSGTACPDFRKGSCKKADACEYAHGVFECWLHPARYRTQPCKDG 542
>TC14620
Length = 1260
Score = 207 bits (528), Expect = 2e-54
Identities = 103/195 (52%), Positives = 120/195 (60%)
Frame = +3
Query: 68 FPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSC 127
F T + D S DHFRMF+FK+ C RGR HDWT+CP++HPGEKARRRDPRK++YS T C
Sbjct: 132 FATTPTEDFSSTDHFRMFDFKVNLCQRGRPHDWTDCPYAHPGEKARRRDPRKHHYSATPC 311
Query: 128 PDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQ 187
PD RKG C +GDSC FAHGVFECWLHPSRYRTQPCKDGT CRR VCFFAHT QLR
Sbjct: 312 PDLRKGYCNRGDSCHFAHGVFECWLHPSRYRTQPCKDGTFCRRRVCFFAHTANQLRVLNH 491
Query: 188 QSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGS 247
+ S SP + SS + T F V +
Sbjct: 492 EHVLS---------SPTSVLNSSSTLTTKSFRYG----------------------VVSA 578
Query: 248 SSVNEMVASLRNLQL 262
SV+E+V S+RN+QL
Sbjct: 579 FSVDEIVRSMRNVQL 623
>TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization 1 homolog a
{Mus musculus;}, partial (3%)
Length = 2631
Score = 199 bits (506), Expect = 6e-52
Identities = 108/233 (46%), Positives = 145/233 (61%), Gaps = 11/233 (4%)
Frame = +2
Query: 63 SDSEIFPTHESVDS-----YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDP 117
SD + +P S+ Y +D FRM+ FK++ C+R SHDWTECPF HPGE ARRRDP
Sbjct: 698 SDKKEYPVDVSLPDINNGLYGSDEFRMYSFKVKPCSRAYSHDWTECPFVHPGENARRRDP 877
Query: 118 RKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAH 177
RKY YS CP+FRKG+C+KGDSCE+AHGVFE WLHP++YRT+ CKD T C R VCFFAH
Sbjct: 878 RKYPYSCVPCPEFRKGACQKGDSCEYAHGVFESWLHPAQYRTRLCKDETGCGRKVCFFAH 1057
Query: 178 TTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPM 237
E+LR + ++PS SY + A + S + L SS +S V +PPMSP+
Sbjct: 1058RPEELRPVYASTGSAMPSPKSYSSN----AVDMSVMSPLALSSSSLPMS-TVSTPPMSPL 1222
Query: 238 TRSLGRSVGSSSVNEMVASLR--NLQL--GTMKSLPSS--WNVQMGSPRFGSP 284
SL G+ N++ +L +LQL +K+ S+ ++++M GSP
Sbjct: 1223AGSLSPKSGNLWQNKINLNLTPPSLQLPGSRLKTALSARDFDLEMEMLGLGSP 1381
>BP063686
Length = 514
Score = 164 bits (415), Expect(2) = 3e-44
Identities = 70/111 (63%), Positives = 85/111 (76%), Gaps = 5/111 (4%)
Frame = -2
Query: 61 SDSDSEIFPTHESVDS-----YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRR 115
S S+ + +P S+ YS D FRM+ FK+R C+R SHDWTECPF HPGE ARRR
Sbjct: 420 SVSEKKEYPVDPSLPDIKNSIYSTDEFRMYSFKVRPCSRAYSHDWTECPFVHPGENARRR 241
Query: 116 DPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGT 166
DPRKY+YS CPDFRKG+C++GD CE+AHGVFECWLHP++YRT+ CKDGT
Sbjct: 240 DPRKYHYSCVPCPDFRKGACRRGDMCEYAHGVFECWLHPAQYRTRLCKDGT 88
Score = 31.2 bits (69), Expect(2) = 3e-44
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = -1
Query: 167 SCRRPVCFFAHTTEQLR 183
+C R +CFFAHT E+LR
Sbjct: 88 NCSRRICFFAHTAEELR 38
>AV428287
Length = 286
Score = 103 bits (257), Expect = 5e-23
Identities = 53/92 (57%), Positives = 64/92 (68%), Gaps = 3/92 (3%)
Frame = +1
Query: 9 RTNPTVHVPPWPTLNNPTAEIFSPLTSND---DYSQFYMQEALSAFQHYVNENNDSDSDS 65
R NPTVHVPPWP +N+PT EIFSPLT+ + D+S FY+QEAL + +D+DSD
Sbjct: 37 RANPTVHVPPWPMVNDPTTEIFSPLTATNADYDHSPFYLQEALEQY-----APSDADSD- 198
Query: 66 EIFPTHESVDSYSNDHFRMFEFKIRRCARGRS 97
P +S D YS D FRMFEFK+R CARGRS
Sbjct: 199 ---PGRDSADVYSFDQFRMFEFKVRNCARGRS 285
>BP083473
Length = 214
Score = 58.5 bits (140), Expect = 2e-09
Identities = 22/32 (68%), Positives = 25/32 (77%)
Frame = -3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHP 108
Y+ D FRMF FK+R C+R SHDWTECPF HP
Sbjct: 98 YATDEFRMFSFKVRPCSRAYSHDWTECPFVHP 3
>TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;}, partial (4%)
Length = 576
Score = 50.4 bits (119), Expect = 5e-07
Identities = 32/87 (36%), Positives = 45/87 (50%)
Frame = +2
Query: 172 VCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVES 231
VCFFAH E+LR + ++PS SY + A E V + F SP S+ PPV +
Sbjct: 5 VCFFAHKLEELRPLYVSTGSAIPSPRSYSAN--NSALEMGSVSPIAF-GSPSSMMPPVST 175
Query: 232 PPMSPMTRSLGRSVGSSSVNEMVASLR 258
PP+SP S + G+ N V +L+
Sbjct: 176 PPLSPSGASSPVAGGAMWQNVSVPALQ 256
>TC16353 weakly similar to UP|HQGT_ARATH (Q9M156) Probable hydroquinone
glucosyltransferase (Arbutin synthase) , partial (52%)
Length = 885
Score = 36.6 bits (83), Expect = 0.007
Identities = 38/139 (27%), Positives = 52/139 (37%), Gaps = 11/139 (7%)
Frame = +2
Query: 154 PSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQ----------SPRSVPSVDSYDGS- 202
P + Q K +S R + F + +PT + SP S P S S
Sbjct: 176 PPKLHRQKLKSPSSNRFRIPFHTFSFLPSTSPTSRKTP*LKYESPSPYSAPFRPSATPSA 355
Query: 203 PLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQL 262
P SS L +SP + +PP+ S + P RSL S + +AS N
Sbjct: 356 PSPPPHSSSTSSALTPSTSPENSAPPLTSSSLPPPPRSLSSSTSLAWTGRFIASSENSPN 535
Query: 263 GTMKSLPSSWNVQMGSPRF 281
+ PS VQ S RF
Sbjct: 536 RSKSPAPSRSTVQTFSTRF 592
>TC9030 similar to GB|AAM63479.1|21554372|AY086477 phospholipase-like
protein {Arabidopsis thaliana;}, partial (53%)
Length = 596
Score = 36.2 bits (82), Expect = 0.009
Identities = 42/140 (30%), Positives = 52/140 (37%), Gaps = 9/140 (6%)
Frame = +1
Query: 151 WLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRL---- 206
W SR RT TS R TT+ + AP+ SP S S G P L
Sbjct: 55 WCTQSRRRTSTTPSATSLRTSSTL---TTQ*VTAPSS-SPIPAASSFSPSGGPHSLRPNQ 222
Query: 207 -----AFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQ 261
+ +S SSP SP +SPP TR GSS + M S +
Sbjct: 223 SEPSPSCTASPASQAGSSSSPPYTSPKPDSPPAPSTTRDTASPTGSSRTSPM--STPSSM 396
Query: 262 LGTMKSLPSSWNVQMGSPRF 281
+ S+PS V PRF
Sbjct: 397 TASRSSIPS---VPASIPRF 447
>TC8116 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), partial (31%)
Length = 595
Score = 33.9 bits (76), Expect = 0.046
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +2
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFE 149
N+ C +F KG+C G+ C FAHG E
Sbjct: 278 NFKTKLCENFAKGTCTFGERCHFAHGPAE 364
>AU251563
Length = 350
Score = 33.1 bits (74), Expect = 0.079
Identities = 23/66 (34%), Positives = 31/66 (46%)
Frame = +2
Query: 15 HVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSEIFPTHESV 74
HVPP+P + LTSNDD E++SA ++DSD DSE +
Sbjct: 119 HVPPFPPSSFSADTRDGALTSNDDL------ESISAR----GADSDSDEDSEDAVVDDEE 268
Query: 75 DSYSND 80
D Y N+
Sbjct: 269 DDYDNE 286
>BP037936
Length = 518
Score = 32.7 bits (73), Expect = 0.10
Identities = 17/63 (26%), Positives = 28/63 (43%)
Frame = +3
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTE 180
+Y T C + + C KGD+C F H + + R+ + CR C + HT E
Sbjct: 267 SYRQTVCRHWLRSLCMKGDACGFLHQYDKSRMPVCRF----FRLYGECREQDCVYKHTNE 434
Query: 181 QLR 183
++
Sbjct: 435 DIK 443
>TC17069 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), partial (33%)
Length = 577
Score = 32.3 bits (72), Expect = 0.13
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +2
Query: 127 CPDFRKGSCKKGDSCEFAHGVFE 149
C +F +GSC G+ C FAHG E
Sbjct: 281 CENFAQGSCTFGERCHFAHGTDE 349
>BP072877
Length = 433
Score = 31.2 bits (69), Expect = 0.30
Identities = 20/81 (24%), Positives = 40/81 (48%), Gaps = 2/81 (2%)
Frame = -1
Query: 167 SCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFES--SCVKTLQFMSSPGS 224
SC +C + T + T+ S R V S +S G+ + + F+ S +++ +PG+
Sbjct: 334 SCSSELCISSRTKRNWKKRTKSSKR-VSSSNSSSGASVPILFQKQRSSELSVKLKIAPGT 158
Query: 225 VSPPVESPPMSPMTRSLGRSV 245
++P + PP + L +S+
Sbjct: 157 LNPEILPPPQIRASGGLLKSI 95
>TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal protein S1
{Arabidopsis thaliana;}, partial (5%)
Length = 691
Score = 30.8 bits (68), Expect = 0.39
Identities = 23/97 (23%), Positives = 42/97 (42%), Gaps = 8/97 (8%)
Frame = +3
Query: 170 RPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSP-- 227
+P C F+ + + + P P+ SPL + + V + SSP S+ P
Sbjct: 252 KPPCTFSSSAYSSQLWLSEQPSPSPATSRRISSPLPASVSPTPVASKNAGSSPASLEPTR 431
Query: 228 -PVES-----PPMSPMTRSLGRSVGSSSVNEMVASLR 258
P+E+ SP+ + L R GS ++ + + +R
Sbjct: 432 SPIETHREPFGASSPIVKGLNRIAGSKTLPILPSPVR 542
>TC19130 similar to UP|Q7YSZ8 (Q7YSZ8) GE-rich protein precursor, partial
(20%)
Length = 603
Score = 30.8 bits (68), Expect = 0.39
Identities = 10/30 (33%), Positives = 18/30 (59%)
Frame = +2
Query: 7 PHRTNPTVHVPPWPTLNNPTAEIFSPLTSN 36
PH + T++ PPW NP+ +F P++ +
Sbjct: 248 PHHHHVTMNEPPWSPPQNPSRHLFPPISGD 337
>TC12640 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase precursor ,
partial (16%)
Length = 738
Score = 30.4 bits (67), Expect = 0.51
Identities = 18/58 (31%), Positives = 28/58 (48%)
Frame = +1
Query: 221 SPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGS 278
SP +++ P +P + T S R+ SSS NE ++ + T +PSS Q S
Sbjct: 565 SPATMASP*SAPTRTASTTSTTRAATSSSRNESKSTSQRCSAPTSHQVPSSPRTQTSS 738
Score = 28.5 bits (62), Expect = 1.9
Identities = 24/103 (23%), Positives = 39/103 (37%)
Frame = +3
Query: 177 HTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSP 236
H T+Q + P + ++PS S SC LQF P S P + +SP
Sbjct: 336 H*TQQQQQPKMMASSTIPSSSS------------SCFHQLQF---PSSTRKPTKPFSVSP 470
Query: 237 MTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSP 279
+ S+ S SV + S ++ +P + + P
Sbjct: 471 IRASVSEKPSSISVPSVSVSSPEPTKLPIRKIPGDHGIPLIGP 599
>TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (55%)
Length = 557
Score = 30.0 bits (66), Expect = 0.67
Identities = 34/143 (23%), Positives = 54/143 (36%), Gaps = 10/143 (6%)
Frame = +2
Query: 151 WLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFES 210
W+ +R+RT+ C +H Q + S + + S G P L+ +
Sbjct: 20 WVDSNRWRTRTTTP--------CGQSHGHPQPPIALRSSSPVLLTRPSGSGGPTSLSSNA 175
Query: 211 SCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPS 270
T S P ++P SPP P T S S+ ++ + L L+ G S+P
Sbjct: 176 PTPATAS-ASLPSPLTPSAPSPPPLPSTASFASSMLIPTLPSPPSRLHPLKYGKCVSIPR 352
Query: 271 ----SWNVQ------MGSPRFGS 283
W V+ G+P GS
Sbjct: 353 VPF*QWLVEAVHQ*SFGTPPHGS 421
>BP046863
Length = 580
Score = 29.6 bits (65), Expect = 0.87
Identities = 21/75 (28%), Positives = 34/75 (45%)
Frame = -3
Query: 296 LPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGS 355
LP + SR +N F + E ++E+ GR V + + S GSG+G
Sbjct: 317 LPLNSSNNASRSSINIF-----TASTESLIEQKCGGRRPVVTRQQVVGNGRSGAGSGLGR 153
Query: 356 GSGLGEVVEDPDVGW 370
G+ +V E ++GW
Sbjct: 152 SYGIVKVRELSEIGW 108
>TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CWP1
precursor, partial (6%)
Length = 937
Score = 29.6 bits (65), Expect = 0.87
Identities = 35/130 (26%), Positives = 49/130 (36%), Gaps = 2/130 (1%)
Frame = +2
Query: 158 RTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVP--SVDSYDGSPLRLAFESSCVKT 215
+TQ D S A T + + +SP S P + S+ + + +S
Sbjct: 65 QTQMPSDSESESESTTSIASLTLSSSSSSTRSPTSKP*VAAASFPAASTPSSLKSKTSSF 244
Query: 216 LQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQ 275
SSP + P PP +P S G+SS + AS +L SSW VQ
Sbjct: 245 ASTASSPTTTHLPPPPPPTNPAAPS-----GTSSASSSAASPNPSRLWV-----SSW-VQ 391
Query: 276 MGSPRFGSPR 285
PR PR
Sbjct: 392 RELPRLPPPR 421
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,150,364
Number of Sequences: 28460
Number of extensions: 142337
Number of successful extensions: 1212
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 1114
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1202
length of query: 381
length of database: 4,897,600
effective HSP length: 92
effective length of query: 289
effective length of database: 2,279,280
effective search space: 658711920
effective search space used: 658711920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144766.6


