

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.5 - phase: 0 /pseudo
(443 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
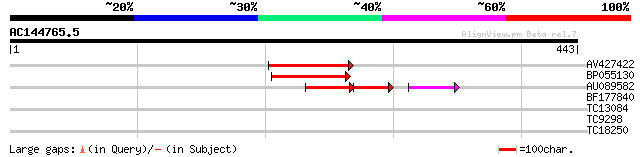
Score E
Sequences producing significant alignments: (bits) Value
AV427422 82 2e-16
BP055130 72 2e-13
AU089582 54 8e-13
BF177840 32 0.21
TC13084 32 0.27
TC9298 UP|CAE02644 (CAE02644) Ornithine decarboxylase , complete 29 1.4
TC18250 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partia... 27 6.8
>AV427422
Length = 417
Score = 81.6 bits (200), Expect = 2e-16
Identities = 34/66 (51%), Positives = 51/66 (76%)
Frame = +1
Query: 203 SRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTS 262
S+G++++ VVVDRL+K +HF+P+ Y +A+I+++ +V+LHGVP SIVSDRDP F S
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 263 RFWRSL 268
FW+ L
Sbjct: 199 NFWKEL 216
>BP055130
Length = 567
Score = 72.0 bits (175), Expect = 2e-13
Identities = 34/62 (54%), Positives = 43/62 (68%)
Frame = +2
Query: 205 GHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRF 264
G I VVVDRL+K HF P +Y +Q+AE ++ IVKLHG+P +IVSDRD FTS F
Sbjct: 179 GFTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAF 358
Query: 265 WR 266
W+
Sbjct: 359 WK 364
>AU089582
Length = 383
Score = 54.3 bits (129), Expect(3) = 8e-13
Identities = 24/38 (63%), Positives = 29/38 (76%)
Frame = +2
Query: 232 AQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
+Q A+IY+ IV LHGVP SI+SDR +FTS FWRS Q
Sbjct: 14 SQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQ 127
Score = 28.1 bits (61), Expect(3) = 8e-13
Identities = 16/32 (50%), Positives = 20/32 (62%)
Frame = +3
Query: 269 QMRWVRN*N*VQLIIRRLMVSRRGQSNL*RIC 300
++ W +* *V L I +LMVS RG *RIC
Sbjct: 126 KLLWELD*K*VPLFILKLMVSPRGLFRS*RIC 221
Score = 26.9 bits (58), Expect(3) = 8e-13
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = +1
Query: 312 GIVIYH**NSHTTTVTIRVLVWHHSKLYMVGSVRLLYVGS 351
GI I * NSH WHH K MVG L VGS
Sbjct: 256 GISICL*WNSHIIIAIGLAFRWHHLKPCMVGDAGHL*VGS 375
>BF177840
Length = 410
Score = 32.0 bits (71), Expect = 0.21
Identities = 13/19 (68%), Positives = 16/19 (83%)
Frame = +2
Query: 250 SSIVSDRDPRFTSRFWRSL 268
+SIVSDRD +F S FWR+L
Sbjct: 11 TSIVSDRDTKFISHFWRTL 67
>TC13084
Length = 508
Score = 31.6 bits (70), Expect = 0.27
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +1
Query: 85 SNQAEESDFKVDEQGVLRFRGRICI 109
SN E+SD+ +E+ RFRGR CI
Sbjct: 157 SNMTEDSDYDEEEEDYPRFRGRYCI 231
>TC9298 UP|CAE02644 (CAE02644) Ornithine decarboxylase , complete
Length = 1522
Score = 29.3 bits (64), Expect = 1.4
Identities = 15/60 (25%), Positives = 24/60 (40%)
Frame = -2
Query: 146 WWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRG 205
WWS +A F + T + P + LD+P+W S ++ S P + G
Sbjct: 930 WWSPKWARIAAFTFVAATSSSGPQVNPPPMSRIDILDIPKWLAISKTLLAASIAPERASG 751
>TC18250 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (3%)
Length = 757
Score = 26.9 bits (58), Expect = 6.8
Identities = 19/51 (37%), Positives = 24/51 (46%), Gaps = 6/51 (11%)
Frame = -2
Query: 203 SRGHDSIWVVVDRLTKSAHFIPINIS------YPVAQLAEIYIQNIVKLHG 247
+R D IW V+ S HFIP+NIS YP A I N + + G
Sbjct: 189 ARNMDVIWTVI-----SFHFIPLNISQGHNETYPAQNSA---INNSITISG 61
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.148 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,323,177
Number of Sequences: 28460
Number of extensions: 94038
Number of successful extensions: 754
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 745
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 754
length of query: 443
length of database: 4,897,600
effective HSP length: 93
effective length of query: 350
effective length of database: 2,250,820
effective search space: 787787000
effective search space used: 787787000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144765.5


