

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144759.9 - phase: 0
(749 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
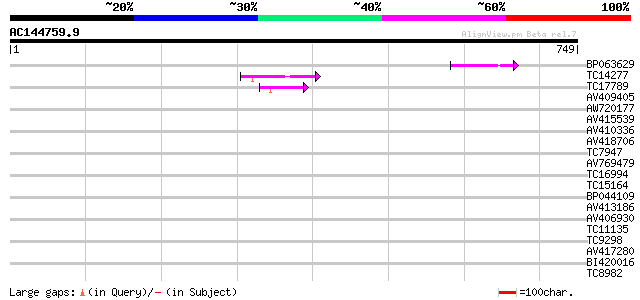
Score E
Sequences producing significant alignments: (bits) Value
BP063629 50 1e-06
TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_1... 47 1e-05
TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 45 4e-05
AV409405 40 0.001
AW720177 40 0.001
AV415539 40 0.002
AV410336 39 0.002
AV418706 39 0.003
TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, parti... 39 0.004
AV769479 39 0.004
TC16994 similar to UP|O22805 (O22805) At2g33550 protein, partial... 38 0.005
TC15164 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, p... 38 0.007
BP044109 38 0.007
AV413186 37 0.009
AV406930 37 0.009
TC11135 homologue to UP|SCRK_SOLTU (P37829) Fructokinase , part... 37 0.009
TC9298 UP|CAE02644 (CAE02644) Ornithine decarboxylase , complete 37 0.012
AV417280 37 0.015
BI420016 36 0.020
TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, parti... 36 0.020
>BP063629
Length = 525
Score = 50.4 bits (119), Expect = 1e-06
Identities = 29/90 (32%), Positives = 51/90 (56%)
Frame = +1
Query: 583 VFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQN 642
+F+ + + + + I + ++ F A+ F+++ GV D++T+ VLKA G + +
Sbjct: 259 IFDHIQQPSLFNYNVMIKAFAKKGSFRRAISLFQQLREDGVWPDNYTYPYVLKAIGCLGD 438
Query: 643 RGSCGEQVHADAIKLGLDSDSYVQCSLIAM 672
G G +VHA IK GL+ D+YV SL+ M
Sbjct: 439 VGQ-GRKVHAFVIKSGLEFDAYVCNSLMDM 525
Score = 38.1 bits (87), Expect = 0.005
Identities = 33/145 (22%), Positives = 64/145 (43%), Gaps = 2/145 (1%)
Frame = +1
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFV--SCGLLENARRVFDVMS 483
SL+K C + + ++ I G++ LN+++ + S G A R+FD +
Sbjct: 106 SLLKSCKSMCELK---QIQALIFCSGLQQDRDTLNKLMAISTDSSIGDFHYALRIFDHIQ 276
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVP 543
++ + ++ + G + AI +F QL G + + +LKA C +V
Sbjct: 277 QPSLFNYNVMIKAFAKKGSFRRAISLFQ----QLREDGVWPDNYTYPYVLKAIGCLGDVG 444
Query: 544 LGMQVHGCLLKLGACDHVLISSSLI 568
G +VH ++K G + +SL+
Sbjct: 445 QGRKVHAFVIKSGLEFDAYVCNSLM 519
>TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_13), partial
(33%)
Length = 941
Score = 47.0 bits (110), Expect = 1e-05
Identities = 35/116 (30%), Positives = 49/116 (42%), Gaps = 10/116 (8%)
Frame = +3
Query: 305 WYDMQFDIKYAYFNF-------LQSIDYRSP-TSPMMPIVAPPPPAPPTRRRNADTTSTT 356
W D F+ K+ Y + L S RSP +S P PPP AP + T S T
Sbjct: 9 WADAWFNFKFLYISLSDSDPWLLSSAASRSPPSSSSSPSPPPPPHAPAAASSSPPTPSAT 188
Query: 357 SPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPPKKSKRRRKCDTT--SHILP 410
PP+ P R +P P + S +P++ T PK + + +T SH P
Sbjct: 189 LPPTPSPP-----SPRSDPSPLSTSTTNPTTSSSTTPKTTTNKSNPNTARLSHAAP 341
Score = 28.1 bits (61), Expect = 5.4
Identities = 21/65 (32%), Positives = 25/65 (38%)
Frame = +1
Query: 335 IVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPPK 394
++ PPP PPT R S T P LLRL L P HP + P +
Sbjct: 268 LLRPPPKPPPTNRTPTRRVSPTRPLRVLHRSLLRLHLPP*PHQG-----HPQRRACPPLR 432
Query: 395 KSKRR 399
RR
Sbjct: 433 PWMRR 447
>TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (14%)
Length = 940
Score = 45.1 bits (105), Expect = 4e-05
Identities = 26/71 (36%), Positives = 34/71 (47%), Gaps = 7/71 (9%)
Frame = -3
Query: 331 PMMPIVAPPPPAP-------PTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLI 383
P I +PPPP+P P R + A + + SPPS+ PH L P R P+
Sbjct: 584 PSSGIGSPPPPSPHNSASSPPQRTQAATASPSPSPPSSATPHSLYKPSSRAPRLSRFL*T 405
Query: 384 HPSSQPITPPK 394
SS P TPP+
Sbjct: 404 SSSSNPSTPPE 372
>AV409405
Length = 424
Score = 40.4 bits (93), Expect = 0.001
Identities = 23/75 (30%), Positives = 40/75 (52%)
Frame = +1
Query: 649 QVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQN 708
Q+H+ +KLGL DS++ L +YG+ + A +F+ T +++V NA+L Y
Sbjct: 199 QLHSLCLKLGLTHDSFIVTKLNVLYGKHASIHHAHKLFDET-PDKSVYLWNALLRRYRAE 375
Query: 709 GLYIEAVKFVYQMKA 723
G + E + +M A
Sbjct: 376 GEWAETLSLFRRMHA 420
Score = 37.4 bits (85), Expect = 0.009
Identities = 23/87 (26%), Positives = 38/87 (43%)
Frame = +1
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHN 591
LL+ C C + Q+H LKLG I + L YG+ + A+ +F+ +
Sbjct: 163 LLETCHCEKSTS---QLHSLCLKLGLTHDSFIVTKLNVLYGKHASIHHAHKLFDETPDKS 333
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKM 618
W A + E ++E L F++M
Sbjct: 334 VYLWNALLRRYRAEGEWAETLSLFRRM 414
Score = 31.6 bits (70), Expect = 0.48
Identities = 16/92 (17%), Positives = 42/92 (45%)
Frame = +1
Query: 422 DIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDV 481
D+ +L++ C ++ +LH+ + G+ ++ ++ +++ + +A ++FD
Sbjct: 148 DLIATLLETCHCE---KSTSQLHSLCLKLGLTHDSFIVTKLNVLYGKHASIHHAHKLFDE 318
Query: 482 MSVRDFHSWATLFVSYYENGEYENAIDVFVSM 513
+ + W L Y GE+ + +F M
Sbjct: 319 TPDKSVYLWNALLRRYRAEGEWAETLSLFRRM 414
>AW720177
Length = 505
Score = 40.0 bits (92), Expect = 0.001
Identities = 29/71 (40%), Positives = 35/71 (48%), Gaps = 5/71 (7%)
Frame = +1
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS- 386
PTSP+ P PPPP PPT + T+ST SPP + P P R + P SL S
Sbjct: 214 PTSPISPPSFPPPP-PPTH--SLSTSSTPSPPLSTPPS----PSRNSATPPPTSLAXSSK 372
Query: 387 ----SQPITPP 393
S P +PP
Sbjct: 373 PKATSTPSSPP 405
Score = 28.5 bits (62), Expect = 4.1
Identities = 18/67 (26%), Positives = 27/67 (39%)
Frame = +1
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
+P SP PPP + + T++ +SPPS +P S++ S
Sbjct: 307 TPPSPSRNSATPPPTSLAXSSKPKATSTPSSPPST----------ATSPTATYYSVVDSS 456
Query: 387 SQPITPP 393
S P PP
Sbjct: 457 STPPLPP 477
>AV415539
Length = 419
Score = 39.7 bits (91), Expect = 0.002
Identities = 31/99 (31%), Positives = 41/99 (41%), Gaps = 15/99 (15%)
Frame = +3
Query: 328 PTSPMMPIVAPPPPAP-PTRRRNADTTSTTSPPSN----HQPHLLR-----LPLRRNPKP 377
P P +P+ PPPP P PTR PP N H PHL R LRR P
Sbjct: 75 PPPPPLPLAPPPPPPPHPTR----------PPPGNPQPLHVPHLRRYFHRNTRLRRRRDP 224
Query: 378 KNLSLIHPSSQ-----PITPPKKSKRRRKCDTTSHILPL 411
++ ++ P P+ PP+ R + +LPL
Sbjct: 225 RHALVLAPLQNPF*MAPLFPPRNPTLRARVLLVLPLLPL 341
>AV410336
Length = 358
Score = 39.3 bits (90), Expect = 0.002
Identities = 29/87 (33%), Positives = 39/87 (44%), Gaps = 3/87 (3%)
Frame = +1
Query: 327 SPTSPMMPI--VAPPPPAPPTRRRNADTTSTTSP-PSNHQPHLLRLPLRRNPKPKNLSLI 383
SP +P APPP APP N + S++SP P++ PH RN P +
Sbjct: 22 SPPTPTTSTSSTAPPPHAPPPPTPNPKSPSSSSPTPTSPSPH-----SGRNSSPATTTFT 186
Query: 384 HPSSQPITPPKKSKRRRKCDTTSHILP 410
+S P TPP S T + +LP
Sbjct: 187 TSTSTP-TPPPPSSLPAASSTAASLLP 264
>AV418706
Length = 374
Score = 38.9 bits (89), Expect = 0.003
Identities = 16/35 (45%), Positives = 23/35 (65%)
Frame = +1
Query: 326 RSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPS 360
RS P P +APPPP RR+N ++T+ T+PP+
Sbjct: 10 RSTLQPSPPAMAPPPPTSHYRRQNPNSTAPTAPPT 114
>TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, partial (81%)
Length = 1294
Score = 38.5 bits (88), Expect = 0.004
Identities = 29/89 (32%), Positives = 36/89 (39%), Gaps = 11/89 (12%)
Frame = +2
Query: 327 SPTSPMMPIV----------APPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPK 376
SP+SP P +PPPP PP+ T+ T P S Q P
Sbjct: 167 SPSSPQPPNSPSSLSSPDPSSPPPPPPPSPPNANPPTTLTPPSSTPQTQKKTSPSTHQKS 346
Query: 377 PKNLSLIHPS-SQPITPPKKSKRRRKCDT 404
PK SL+ PS + P P KS R + T
Sbjct: 347 PKASSLLLPSTTAPSKPRTKSPPRMRKST 433
Score = 31.2 bits (69), Expect = 0.63
Identities = 31/94 (32%), Positives = 43/94 (44%), Gaps = 3/94 (3%)
Frame = +2
Query: 330 SPMMPIVAPPPPAPPTRRRNADTTSTTSP-PSNHQPHLLRLPLRRNPKPKNLSLIHPSSQ 388
SP+ P APP P+P + + +S +SP PS+ P P NP +L PSS
Sbjct: 134 SPLSPS-APPHPSPSSPQPPNSPSSLSSPDPSSPPPPPPPSPPNANPP---TTLTPPSST 301
Query: 389 PITPPK--KSKRRRKCDTTSHILPLMDALHFPIT 420
P T K S ++ +S +LP A P T
Sbjct: 302 PQTQKKTSPSTHQKSPKASSLLLPSTTAPSKPRT 403
>AV769479
Length = 287
Score = 38.5 bits (88), Expect = 0.004
Identities = 17/76 (22%), Positives = 42/76 (54%)
Frame = +1
Query: 531 CLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRH 590
C + C+ +++ LG Q+H +LK +++ ++ + FY + A F+R+++
Sbjct: 10 CHMNLCSKRVDLALGKQIHAHILK-SKWRNLIADNAFVNFYAECGKISSAFRTFDRMAKR 186
Query: 591 NTLTWTAKIVSSCRER 606
+ + WT I+++C ++
Sbjct: 187 DVVCWTT-IITACSQQ 231
Score = 34.3 bits (77), Expect = 0.075
Identities = 21/84 (25%), Positives = 36/84 (42%)
Frame = +1
Query: 431 CTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSW 490
C+ D ++H I+ + L N + + CG + +A R FD M+ RD W
Sbjct: 25 CSKRVDLALGKQIHAHIL-KSKWRNLIADNAFVNFYAECGKISSAFRTFDRMAKRDVVCW 201
Query: 491 ATLFVSYYENGEYENAIDVFVSML 514
T+ + + G A+ + ML
Sbjct: 202 TTIITACSQQGLGHEALLILSQML 273
>TC16994 similar to UP|O22805 (O22805) At2g33550 protein, partial (36%)
Length = 680
Score = 38.1 bits (87), Expect = 0.005
Identities = 29/90 (32%), Positives = 35/90 (38%), Gaps = 9/90 (10%)
Frame = -1
Query: 331 PMMPIVAPPPP----APPTRRRN-----ADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLS 381
P P +P PP A P RR + +T +T+SP SNH P
Sbjct: 644 PRAPSQSPEPP*QQPATPARREHHTPPGQNTPATSSPSSNHSSSAKTTPF---------- 495
Query: 382 LIHPSSQPITPPKKSKRRRKCDTTSHILPL 411
HPS TP RRR C T S P+
Sbjct: 494 --HPSPTTPTP*SSYSRRRGCSTASGTEPV 411
>TC15164 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(57%)
Length = 1185
Score = 37.7 bits (86), Expect = 0.007
Identities = 47/180 (26%), Positives = 71/180 (39%), Gaps = 19/180 (10%)
Frame = +2
Query: 330 SPMMPIVAPPPPAP------PTRRRNADTTSTTSPPSNHQPHLLRL-PLRRNPKPKNLSL 382
SP+ PP P+P P+RR TT+TT+PP++ P LL P++ P P
Sbjct: 293 SPVTTTSTPPTPSPAATNASPSRR---STTTTTTPPTSPTPMLLTSPPMKLKPLPGYFLP 463
Query: 383 IHPSSQPITPPKKSKRRRKCDTTSHILPLMDALHF---PITIDIYTSLVKECTLSTDPET 439
+ PI+ P + K +S I P + P + + + +L + T
Sbjct: 464 LPTPKAPISTPLTTP-TPKSTPSSSISPKLTRQSITAPPPPTESFRCNHRARSLQSSTTT 640
Query: 440 AIELHTQIITRGIELPL--TLLNRILIM-------FVSCGLLENARRVFDVMSVRDFHSW 490
L T+ I+ + L L T LNR + V C + RR V R W
Sbjct: 641 TTTLTTKSISLPLSLSLTTTTLNRTTTLCRRRRWRLVLCRMGTRCRRFRTVAMERRRRRW 820
Score = 32.3 bits (72), Expect = 0.28
Identities = 21/80 (26%), Positives = 28/80 (34%)
Frame = +2
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
S SP P PP +PP + S + PP L LP + P + P+
Sbjct: 155 SSASPATPPSTPPTSSPPATLASLSARSASKPPPPSPARLTPLP---SASPVTTTSTPPT 325
Query: 387 SQPITPPKKSKRRRKCDTTS 406
P RR TT+
Sbjct: 326 PSPAATNASPSRRSTTTTTT 385
Score = 32.0 bits (71), Expect = 0.37
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Frame = -3
Query: 317 FNFLQSIDYRSPTSPMMP--IVAPPPPAPPTRRRNADTTSTTSPPSNHQP 364
F+ S+ P P P I PPPPAP + + ++ S P+ HQP
Sbjct: 892 FDSSSSLCTSKP*LPYPPPRIAPPPPPAPAFHSNSTKSPTSRSHPAQHQP 743
Score = 27.7 bits (60), Expect = 7.0
Identities = 25/82 (30%), Positives = 33/82 (39%), Gaps = 4/82 (4%)
Frame = +2
Query: 329 TSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQ 388
T P A PP PPT A S ++ ++ P P R P P + S + +S
Sbjct: 146 TQPSSASPATPPSTPPTSSPPATLASLSARSASKPPP--PSPARLTPLP-SASPVTTTST 316
Query: 389 PITP----PKKSKRRRKCDTTS 406
P TP S RR TT+
Sbjct: 317 PPTPSPAATNASPSRRSTTTTT 382
>BP044109
Length = 497
Score = 37.7 bits (86), Expect = 0.007
Identities = 29/85 (34%), Positives = 39/85 (45%), Gaps = 18/85 (21%)
Frame = +3
Query: 328 PTSPMMPIVA-PPPPAPP--------------TRRRNADTTSTTSPPSNHQPHLLRLPLR 372
PT P++ + PPP +PP TR R T T PP+ H+ RL LR
Sbjct: 39 PTPPLLHFLTLPPPHSPPLTSSLLRHHHHHHRTRTRQPHHTLPTLPPNQHRAPKPRLHLR 218
Query: 373 RNPKPKNLSLIH---PSSQPITPPK 394
R P + SL+H P + P+ P K
Sbjct: 219 R-LLPHHPSLLHQPSPQNPPLLPRK 290
Score = 29.3 bits (64), Expect = 2.4
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +2
Query: 337 APPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKP 377
AP P+PP+ ++ TST SPPS L P R P P
Sbjct: 317 APTLPSPPS---SSTPTSTPSPPSPPNGSTLPSPPRATPTP 430
Score = 28.5 bits (62), Expect = 4.1
Identities = 22/79 (27%), Positives = 33/79 (40%), Gaps = 4/79 (5%)
Frame = +2
Query: 322 SIDYRSPTSPMMPIVAP-PPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNL 380
S+ + S TS P PPP PP +N T S S ++ + P P P +
Sbjct: 56 SLSHSSSTSLSSSHFFPSPPPPPPPPNKNKTTPSHASNATSKSTPRTQTPTTPPPSPTSS 235
Query: 381 SLIHPS---SQPITPPKKS 396
PS ++P T P ++
Sbjct: 236 PKPPPSTFATKPSTSPPET 292
>AV413186
Length = 398
Score = 37.4 bits (85), Expect = 0.009
Identities = 22/72 (30%), Positives = 34/72 (46%)
Frame = +1
Query: 672 MYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEP 731
MY + G L A VF + N+++V N M+ +G EA+ + M +GV+P+
Sbjct: 7 MYSKCGSLETAWRVFNLVGNKQDVVLWNTMISALAHHGFGTEAMVMLNDMIRSGVEPNRA 186
Query: 732 LLEKLRIACGSS 743
L AC S
Sbjct: 187 TFVVLLNACSLS 222
>AV406930
Length = 433
Score = 37.4 bits (85), Expect = 0.009
Identities = 32/102 (31%), Positives = 42/102 (40%), Gaps = 24/102 (23%)
Frame = +1
Query: 335 IVAPPPPAPPTRRRNADTTSTTSPPSN--HQP--HLLRLPLRRNPKPKNLSLI------- 383
+ PPPP PP + S ++ P H P H LPL PKP ++ L
Sbjct: 82 LAPPPPPQPPNPPSISQNLSISTHPITPFHTPLSHSFNLPL---PKPASIPLSFSALPSP 252
Query: 384 --HPSSQPI---TPPKKSKRRRKCD--------TTSHILPLM 412
PSS P PP S+RR C ++ ILPL+
Sbjct: 253 SPSPSSSPTLIPPPPSSSRRRGSCSPMNSPLFVSSKRILPLL 378
>TC11135 homologue to UP|SCRK_SOLTU (P37829) Fructokinase , partial (33%)
Length = 402
Score = 37.4 bits (85), Expect = 0.009
Identities = 24/65 (36%), Positives = 31/65 (46%)
Frame = +2
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSS 387
P P P PPPAP + +A ST+SPPS P RLP P + + P+S
Sbjct: 44 PRKPWPPTTLSPPPAPASSSASARC*STSSPPSPASPS-PRLPASSKPP----AALPPTS 208
Query: 388 QPITP 392
Q +P
Sbjct: 209 QSPSP 223
>TC9298 UP|CAE02644 (CAE02644) Ornithine decarboxylase , complete
Length = 1522
Score = 37.0 bits (84), Expect = 0.012
Identities = 27/74 (36%), Positives = 33/74 (44%), Gaps = 3/74 (4%)
Frame = +3
Query: 322 SIDYRSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLS 381
S+ S +P P AP P P A S +S PS L P R NP P + S
Sbjct: 327 SLSTPSSATPTSPSSAPSPH*APASTAPAKLRSKSSCPSASHRTALSTPTRANPSPTS-S 503
Query: 382 LIHPS---SQPITP 392
+ PS SQP+TP
Sbjct: 504 MPQPSASTSQPMTP 545
>AV417280
Length = 427
Score = 36.6 bits (83), Expect = 0.015
Identities = 23/56 (41%), Positives = 31/56 (55%)
Frame = +1
Query: 49 SNSISHWPSLSLWSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASA 104
S S++ PSL LW+ SL+ST+ +PT SA SL P +SP S + SA
Sbjct: 247 SPSLAPTPSLPLWA------TSLTSTSSPSATSPTSSAPSLQPSQNSPHSTTYTSA 396
>BI420016
Length = 559
Score = 36.2 bits (82), Expect = 0.020
Identities = 26/70 (37%), Positives = 33/70 (47%)
Frame = +2
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
+P+ P P + PPP +PP+ NA S +SPPS P NP P PS
Sbjct: 236 TPSPPSSPTLTPPP-SPPSPTPNA-PLSPSSPPSPPTPPSPPPSPAPNPTPSATGSTTPS 409
Query: 387 SQPITPPKKS 396
S P+TP S
Sbjct: 410 S-PLTPTSAS 436
Score = 30.4 bits (67), Expect = 1.1
Identities = 25/80 (31%), Positives = 31/80 (38%)
Frame = +2
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
SP+SP P P PP P T +T+P S PL P + SL PS
Sbjct: 311 SPSSPPSPPTPPSPPPSPAPNPTPSATGSTTPSS---------PL----TPTSASLSSPS 451
Query: 387 SQPITPPKKSKRRRKCDTTS 406
S +P S TT+
Sbjct: 452 SL-FSPASTSPASTTATTTT 508
>TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, partial (10%)
Length = 901
Score = 36.2 bits (82), Expect = 0.020
Identities = 26/68 (38%), Positives = 33/68 (48%), Gaps = 3/68 (4%)
Frame = +3
Query: 341 PAPPTR---RRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPPKKSK 397
P+ PTR R + +TSTTSPP+ + P PLRR P P S P P +S
Sbjct: 279 PSSPTRVATRTLSGSTSTTSPPTLNSPRSRASPLRRGTSP-------PRSPP--PLSRSS 431
Query: 398 RRRKCDTT 405
R +C T
Sbjct: 432 PR*RCRRT 455
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,720,066
Number of Sequences: 28460
Number of extensions: 329414
Number of successful extensions: 5498
Number of sequences better than 10.0: 446
Number of HSP's better than 10.0 without gapping: 3701
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4713
length of query: 749
length of database: 4,897,600
effective HSP length: 97
effective length of query: 652
effective length of database: 2,136,980
effective search space: 1393310960
effective search space used: 1393310960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144759.9


