

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
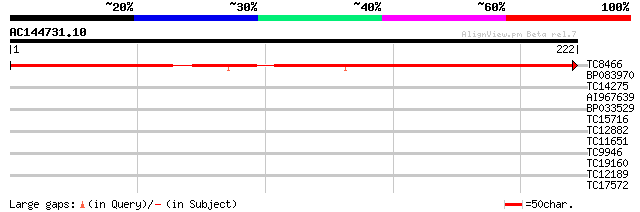
Score E
Sequences producing significant alignments: (bits) Value
TC8466 241 9e-65
BP083970 28 0.96
TC14275 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (... 28 1.3
AI967639 27 2.8
BP033529 27 3.7
TC15716 similar to UP|O78327 (O78327) Transketolase 1 , partial ... 27 3.7
TC12882 similar to UP|PNK1_ARATH (Q8L5Y9) Probable pantothenate ... 26 4.8
TC11651 26 4.8
TC9946 similar to UP|Q9LLM3 (Q9LLM3) MTD1, partial (42%) 26 6.2
TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%) 26 6.2
TC12189 similar to UP|Q9FKV6 (Q9FKV6) Eukaryotic translation ini... 26 6.2
TC17572 similar to UP|O80910 (O80910) At2g38410 protein, partial... 25 8.2
>TC8466
Length = 1265
Score = 241 bits (614), Expect = 9e-65
Identities = 136/236 (57%), Positives = 157/236 (65%), Gaps = 14/236 (5%)
Frame = +3
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
M FLECAPLDRL+DFL +LNLGERTIKGC+EAYSCKH+G DKKLSISL NEILDYLGKSS
Sbjct: 177 MMFLECAPLDRLNDFLCHLNLGERTIKGCLEAYSCKHAGTDKKLSISLDNEILDYLGKSS 356
Query: 61 DNDSPSPVDSLSSPVDSLSARTSP--------AGRHWFT*FLPSITCIQIMISAQLKLTS 112
D+DS SSP D L RTS A H + + S + A T
Sbjct: 357 DSDS-------SSPHDPLITRTSRKTLVYLVLALYHMYPDYDFSA------VKAHQFFTE 497
Query: 113 FSQRNHGTVLSRFLKPTC------LKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADP 166
S + V ++ ++ SLLD L KA+DEVV L DCEIYGY+PD EADP
Sbjct: 498 ESWDSFKQVFDSYMFEASKEWDETVEGLSLLDTLLKAIDEVVKLADCEIYGYVPDYEADP 677
Query: 167 LPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
L + GA+WSFNFLFYNRKLKR+VTF SCFSNLI DGF DE ++YDE+IFADMD
Sbjct: 678 LLDSGAMWSFNFLFYNRKLKRVVTFHFSCFSNLITDGFLADEFHNDYDEQIFADMD 845
>BP083970
Length = 375
Score = 28.5 bits (62), Expect = 0.96
Identities = 23/65 (35%), Positives = 36/65 (55%), Gaps = 2/65 (3%)
Frame = -3
Query: 135 SLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVT--FR 192
S+L+ + + EV L IY +SEAD L +RGAI S + +++N + + + FR
Sbjct: 286 SMLNPIDHWLVEVNCLKVSHIYREA-NSEADKLAKRGAIHSKDLVYFNESIAWMGS*WFR 110
Query: 193 LSCFS 197
L FS
Sbjct: 109 LFVFS 95
>TC14275 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (25%)
Length = 562
Score = 28.1 bits (61), Expect = 1.3
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +1
Query: 37 HSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPAG 86
H +KLSI+++ SS + SPSP S + P DS + AG
Sbjct: 214 HCLGGQKLSINVTGGSSTATPPSSASPSPSPSSSTTPPQDSAAGSLGAAG 363
>AI967639
Length = 335
Score = 26.9 bits (58), Expect = 2.8
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = -1
Query: 44 LSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPA 85
LS+SLS+ +L SS + S S SSP S S+ +SP+
Sbjct: 221 LSLSLSSLMLKSSSSSSSSSSSSSSRLYSSP*SSSSSSSSPS 96
>BP033529
Length = 299
Score = 26.6 bits (57), Expect = 3.7
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Frame = -1
Query: 128 PTC-LKPQSLLDALFKAVDEVVSLDDCEIYGYL 159
P C L+PQ + F +DE+ D C+++GYL
Sbjct: 194 PDCFLRPQL*FNMNFVELDELREKDVCKLWGYL 96
>TC15716 similar to UP|O78327 (O78327) Transketolase 1 , partial (8%)
Length = 567
Score = 26.6 bits (57), Expect = 3.7
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = -1
Query: 175 SFNFLFYNRKLKRIVTFRLSCFSNL 199
SFNF+ + L + T R++C SNL
Sbjct: 393 SFNFISSSHPLHDVATLRITCKSNL 319
>TC12882 similar to UP|PNK1_ARATH (Q8L5Y9) Probable pantothenate kinase 1
(Pantothenic acid kinase 1) , partial (8%)
Length = 647
Score = 26.2 bits (56), Expect = 4.8
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = +2
Query: 16 LSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEIL 53
+ N L E+ +KG I CK+ A+ KL + ++I+
Sbjct: 149 VKNQRLAEKLVKGNIYDCVCKYEPAN*KLCVQTQSQIM 262
>TC11651
Length = 480
Score = 26.2 bits (56), Expect = 4.8
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +3
Query: 174 WSFNFLFYNRKLKRIVTFRLSCFS 197
W F+ L ++ LK IV F LSC +
Sbjct: 303 WLFSILCFSLNLKLIVNFFLSCLA 374
>TC9946 similar to UP|Q9LLM3 (Q9LLM3) MTD1, partial (42%)
Length = 676
Score = 25.8 bits (55), Expect = 6.2
Identities = 17/48 (35%), Positives = 22/48 (45%)
Frame = +3
Query: 37 HSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSP 84
H G + SISLS L ++ +DS S + S S S S SP
Sbjct: 156 HGGGISQRSISLSRSSLALAVAATHSDSSSSITSEDSASSSNSLSPSP 299
>TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%)
Length = 557
Score = 25.8 bits (55), Expect = 6.2
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 3/55 (5%)
Frame = -1
Query: 38 SGADKKLS---ISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPAGRHW 89
SG+ + L+ + LS + SS+ PSP +L SP + + GR W
Sbjct: 386 SGSARSLAFVEVKLSKSSSSFRRDSSEESPPSPSSALWSPAPQSAP*RTLVGRTW 222
>TC12189 similar to UP|Q9FKV6 (Q9FKV6) Eukaryotic translation initiation
factor 3 subunit 7, partial (30%)
Length = 658
Score = 25.8 bits (55), Expect = 6.2
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = +1
Query: 56 LGKSSDNDSPSPVDSLSSPVDSLSARTSPAGR 87
L S SP+P S SP+ ++ T PA R
Sbjct: 115 LSTSPSPPSPAPTSSAGSPIGPATSTTRPAPR 210
>TC17572 similar to UP|O80910 (O80910) At2g38410 protein, partial (28%)
Length = 772
Score = 25.4 bits (54), Expect = 8.2
Identities = 14/36 (38%), Positives = 22/36 (60%)
Frame = +2
Query: 45 SISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSA 80
S +LS+E+ DY + S SPV+S+S+P S+
Sbjct: 608 SSTLSSEV-DYSANQTQVKSSSPVESVSTPNSKASS 712
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,008,572
Number of Sequences: 28460
Number of extensions: 56507
Number of successful extensions: 291
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 288
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 291
length of query: 222
length of database: 4,897,600
effective HSP length: 87
effective length of query: 135
effective length of database: 2,421,580
effective search space: 326913300
effective search space used: 326913300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144731.10


