

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.3 - phase: 0
(151 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
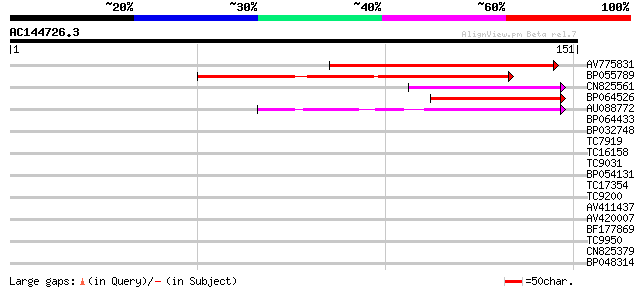
Score E
Sequences producing significant alignments: (bits) Value
AV775831 76 2e-15
BP055789 61 7e-11
CN825561 46 2e-06
BP064526 45 4e-06
AU088772 45 5e-06
BP064433 29 0.31
BP032748 28 0.52
TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus cornicula... 27 1.2
TC16158 27 1.5
TC9031 similar to GB|AAP21369.1|30102902|BT006561 At4g19200 {Ara... 27 1.5
BP054131 27 1.5
TC17354 weakly similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, part... 27 2.0
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 27 2.0
AV411437 27 2.0
AV420007 27 2.0
BF177869 26 2.6
TC9950 similar to UP|AAR08678 (AAR08678) EIN2, partial (11%) 26 2.6
CN825379 26 3.4
BP048314 26 3.4
>AV775831
Length = 475
Score = 76.3 bits (186), Expect = 2e-15
Identities = 36/61 (59%), Positives = 44/61 (72%)
Frame = -3
Query: 86 SSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHK 145
SS SGSSK+ARA++ T L DGC +DLSN R YH RHKVC+LHS+T +VT+ G K
Sbjct: 473 SSFPSGSSKRARAVHYATLTAFCLVDGCSSDLSNCRDYHRRHKVCELHSQTPEVTIAGMK 294
Query: 146 Q 146
Q
Sbjct: 293 Q 291
>BP055789
Length = 455
Score = 61.2 bits (147), Expect = 7e-11
Identities = 36/84 (42%), Positives = 52/84 (61%)
Frame = +3
Query: 51 EFFVDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLA 110
E VDL+LG +S +S KD + + SSS SGSSK++R ++NG+ ++
Sbjct: 216 EATVDLRLGEGSDSVEKSAPDTPKDP---EESKAMSSSPSGSSKRSR-LHNGSLNMACSV 383
Query: 111 DGCQADLSNYRGYHMRHKVCKLHS 134
DGC +DLS+ R YH RH+VC+ HS
Sbjct: 384 DGCTSDLSDCREYHRRHRVCEKHS 455
>CN825561
Length = 646
Score = 46.2 bits (108), Expect = 2e-06
Identities = 20/42 (47%), Positives = 25/42 (58%)
Frame = +2
Query: 107 SSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
S + C ADLS + YH RHKVC+ H+K V + G QRF
Sbjct: 206 SCQVENCDADLSEAKQYHRRHKVCEYHAKAHAVHIAGMHQRF 331
>BP064526
Length = 533
Score = 45.4 bits (106), Expect = 4e-06
Identities = 19/36 (52%), Positives = 25/36 (68%)
Frame = +1
Query: 113 CQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
C+ADLS+ + YH RHKVC+ HSK + +G QRF
Sbjct: 67 CRADLSSAKDYHRRHKVCEGHSKATMALVGNVMQRF 174
>AU088772
Length = 422
Score = 45.1 bits (105), Expect = 5e-06
Identities = 29/82 (35%), Positives = 40/82 (48%)
Frame = +1
Query: 67 ESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMR 126
E G+ K + V T + S S +G S A+ Q+ +GC ADLS + H R
Sbjct: 130 EDGIRKKRVV--VTDLYSKRGSKAGGS----AVPPSCQV-----EGCYADLSGAKAXHRR 276
Query: 127 HKVCKLHSKTSQVTLGGHKQRF 148
HKVC+ H+K L +QRF
Sbjct: 277 HKVCEHHAKAPSXLLSEQQQRF 342
>BP064433
Length = 444
Score = 29.3 bits (64), Expect = 0.31
Identities = 19/66 (28%), Positives = 27/66 (40%)
Frame = +3
Query: 61 VGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNY 120
+GNS L+K+ V KIT S SS ++ T V++ NY
Sbjct: 219 IGNSQRTLALSKAM*VH*VNKITKQLSDRVASSYFGYSLGGFTVKVTTQGQSTNWSNQNY 398
Query: 121 RGYHMR 126
GYH +
Sbjct: 399 EGYHFQ 416
>BP032748
Length = 453
Score = 28.5 bits (62), Expect = 0.52
Identities = 14/34 (41%), Positives = 20/34 (58%), Gaps = 3/34 (8%)
Frame = +1
Query: 17 IVLFLNNDEVFWR---PWHDSIWNILYGFVCFSM 47
I L L ND FW+ W+ W+++ GF+ FSM
Sbjct: 283 ISLILTNDGFFWQFGEHWYS--WSMMMGFLTFSM 378
>TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus corniculatus var.
japonicus;} , complete
Length = 1219
Score = 27.3 bits (59), Expect = 1.2
Identities = 21/63 (33%), Positives = 30/63 (47%)
Frame = +3
Query: 59 GHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLS 118
GH +SS++S K K+ +SSS S+ SK +R I L G QAD+
Sbjct: 171 GHDSSSSSDSDNEKEKEK------SSSSPSSDFKSKVSRLFGREKPIHHVLGGGKQADVF 332
Query: 119 NYR 121
+R
Sbjct: 333 LWR 341
>TC16158
Length = 708
Score = 26.9 bits (58), Expect = 1.5
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 2/46 (4%)
Frame = +1
Query: 81 KITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSN--YRGYH 124
K+T + + SS+ A A+ NG + + L ++ LSN RG+H
Sbjct: 28 KVTVVAFEANHSSELAMALTNGLRSAAKLIASSESSLSNSVSRGFH 165
>TC9031 similar to GB|AAP21369.1|30102902|BT006561 At4g19200 {Arabidopsis
thaliana;}, partial (26%)
Length = 670
Score = 26.9 bits (58), Expect = 1.5
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 1/43 (2%)
Frame = +3
Query: 58 LGHVGNSS-AESGLTKSKDVGGFTKITSSSSSTSGSSKKARAI 99
L H+GNSS A S + S +G + +S+S SGS + I
Sbjct: 207 LCHMGNSSMASSSMESSASMGNLSMASSASMDYSGSGSDSMYI 335
>BP054131
Length = 500
Score = 26.9 bits (58), Expect = 1.5
Identities = 20/51 (39%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Frame = +3
Query: 44 CFSMCANEFFVDLKL-GHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSS 93
C S C + + LK H+ +SS S + S V + SSSSSTS SS
Sbjct: 111 CISQCQMKHCLYLKQ*NHISSSS--SSIASSSSVYTKSSSISSSSSTSSSS 257
>TC17354 weakly similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, partial (24%)
Length = 638
Score = 26.6 bits (57), Expect = 2.0
Identities = 18/55 (32%), Positives = 30/55 (53%)
Frame = +3
Query: 62 GNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQAD 116
G ++AE+ L + G + TS+SSS+SGS +G++ SS +D +D
Sbjct: 156 GTNAAEA-LRAQVNAEGRDEQTSTSSSSSGSGSSESGSGSGSRSSSSSSDSEGSD 317
Score = 25.8 bits (55), Expect = 3.4
Identities = 15/37 (40%), Positives = 20/37 (53%)
Frame = +3
Query: 83 TSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSN 119
TSSSSS SGSS+ + + SS ++G D N
Sbjct: 222 TSSSSSGSGSSESGSGSGSRSSSSSSDSEGSDEDTVN 332
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 26.6 bits (57), Expect = 2.0
Identities = 15/55 (27%), Positives = 27/55 (48%)
Frame = -3
Query: 83 TSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTS 137
TSSSSS+S SS + ++ + SS + + S+Y + C L+ ++
Sbjct: 308 TSSSSSSSSSSSSPSSSSSSSSSSSSTSSSTSSSTSSYS*TSASNLFCSLNGSSA 144
Score = 24.3 bits (51), Expect = 9.8
Identities = 16/49 (32%), Positives = 24/49 (48%)
Frame = -3
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSS 93
F M + F++ G +SSA + + S + SSSSS+S SS
Sbjct: 371 FPMGSRSFWI----GSSSSSSASASTSSSSSSSSSSSSPSSSSSSSSSS 237
>AV411437
Length = 417
Score = 26.6 bits (57), Expect = 2.0
Identities = 17/52 (32%), Positives = 27/52 (51%)
Frame = -1
Query: 61 VGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADG 112
V NSS+ ++ S + SSSSS+S SS + AI + ++ + A G
Sbjct: 162 VSNSSSSESVSSSSNCSNSPSTPSSSSSSSSSSSSS-AIVSCAELPETAASG 10
>AV420007
Length = 280
Score = 26.6 bits (57), Expect = 2.0
Identities = 12/33 (36%), Positives = 21/33 (63%)
Frame = -3
Query: 31 WHDSIWNILYGFVCFSMCANEFFVDLKLGHVGN 63
WH S+W+ Y V F C+ E ++L+ G++G+
Sbjct: 236 WHASVWHCSY--VAFVHCSYESIINLR-GNLGS 147
>BF177869
Length = 509
Score = 26.2 bits (56), Expect = 2.6
Identities = 15/42 (35%), Positives = 22/42 (51%)
Frame = +2
Query: 55 DLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKA 96
D+ GH +S+ SG + S G + S S S+SGS +A
Sbjct: 218 DVAAGH---ASSSSGSSSSSSSGSSSSSDSDSGSSSGSDSEA 334
>TC9950 similar to UP|AAR08678 (AAR08678) EIN2, partial (11%)
Length = 853
Score = 26.2 bits (56), Expect = 2.6
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 5/64 (7%)
Frame = +3
Query: 85 SSSSTSGSSKKARAINNGTQIVSSL---ADGC--QADLSNYRGYHMRHKVCKLHSKTSQV 139
SS SGSS K +N + VSS+ +GC +ADL G H+V L S+
Sbjct: 648 SSDGKSGSSMKNNELNCSSFSVSSIPHCGEGCVWRADLIISFGVWCIHRVLDLSLMESRP 827
Query: 140 TLGG 143
L G
Sbjct: 828 ELWG 839
>CN825379
Length = 646
Score = 25.8 bits (55), Expect = 3.4
Identities = 20/72 (27%), Positives = 32/72 (43%)
Frame = -1
Query: 51 EFFVDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLA 110
E +D G + +SS S + + DVGGF SSS+S ++ A + + + L
Sbjct: 382 ETLIDSPTG-LASSSGASCSSSASDVGGFLWFFFFSSSSSSTTTSASSSSYSCSFILFLF 206
Query: 111 DGCQADLSNYRG 122
C A + G
Sbjct: 205 RCCFASRDSCLG 170
>BP048314
Length = 394
Score = 25.8 bits (55), Expect = 3.4
Identities = 16/34 (47%), Positives = 19/34 (55%)
Frame = -3
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKA 96
N + SGLTKS I SSSSS+S SS +
Sbjct: 302 NLNQTSGLTKSSFRKQSLHIASSSSSSSSSSSSS 201
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,408,374
Number of Sequences: 28460
Number of extensions: 53184
Number of successful extensions: 542
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 530
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 540
length of query: 151
length of database: 4,897,600
effective HSP length: 82
effective length of query: 69
effective length of database: 2,563,880
effective search space: 176907720
effective search space used: 176907720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144726.3


