

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
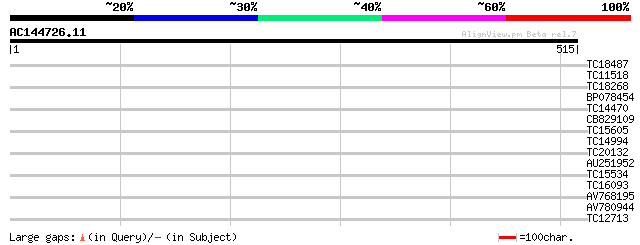
Query= AC144726.11 + phase: 0
(515 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18487 similar to UP|O80953 (O80953) At2g39280 protein, partial... 30 0.73
TC11518 homologue to UP|FTSH_MEDSA (Q9BAE0) Cell division protei... 30 0.95
TC18268 weakly similar to UP|Q9C9S6 (Q9C9S6) Kinesin-related pro... 30 0.95
BP078454 30 1.2
TC14470 similar to UP|AAQ56728 (AAQ56728) Polygalacturonase inhi... 30 1.2
CB829109 29 1.6
TC15605 29 1.6
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 29 2.1
TC20132 similar to UP|Q7X9L1 (Q7X9L1) Holocarboxylase synthetase... 28 2.8
AU251952 28 2.8
TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28... 28 3.6
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 28 3.6
AV768195 28 4.7
AV780944 27 8.0
TC12713 27 8.0
>TC18487 similar to UP|O80953 (O80953) At2g39280 protein, partial (13%)
Length = 592
Score = 30.4 bits (67), Expect = 0.73
Identities = 30/119 (25%), Positives = 50/119 (41%), Gaps = 8/119 (6%)
Frame = +1
Query: 61 FSFSEHPFSFSSI----SLNHFRTYGSVAAEAIESTDTEDDCSG----SEEVVQKLLDQM 112
F F P + S SL+ + + A E + S E D E+VV ++
Sbjct: 142 FGFKHDPKNEQSADMLGSLSRSESGSTNADEILISLTGEGDIDSVPDLHEQVVGLKVELC 321
Query: 113 VIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLA 171
+ EEK++ L ++ K RRQ+ + E EE + ++ L D +EQ+ A
Sbjct: 322 RLLEEKRSAILRAEELETALMEMVKQDNRRQLSAKVEQLEEEIAQLRQALSDKQEQETA 498
>TC11518 homologue to UP|FTSH_MEDSA (Q9BAE0) Cell division protein ftsH
homolog, chloroplast precursor , partial (16%)
Length = 719
Score = 30.0 bits (66), Expect = 0.95
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +3
Query: 18 SQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTS 52
+Q+ TS QH E +H F R +Q+GIFTS
Sbjct: 228 NQTITSSTLMSQHMVSETQEKHQFLRTTQIGIFTS 332
>TC18268 weakly similar to UP|Q9C9S6 (Q9C9S6) Kinesin-related protein,
partial (5%)
Length = 617
Score = 30.0 bits (66), Expect = 0.95
Identities = 29/121 (23%), Positives = 53/121 (42%), Gaps = 4/121 (3%)
Frame = +2
Query: 85 AAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSY----KYKMLR 140
A EA+ + TE++ EVV + + Q+ + + + EEKKK + + K K+
Sbjct: 41 ALEALAAGATEEN-----EVVTRWVQQLKFTMQLEQTKFEEKKKLEEQDFSRLKKDKVRN 205
Query: 141 RRQIKIETEAWEEAAREYQELLEDMREQKLAPNLPYMKSLFLGWFEPLKNAIAADQEICK 200
+I + + A R Y+E + + Q Y K + LK+ +A ++ K
Sbjct: 206 EIEISALKQDLDMAKRTYEEHVSQLELQAAESKAEYEKRIV-----ELKHQLADARKQVK 370
Query: 201 D 201
D
Sbjct: 371 D 373
>BP078454
Length = 385
Score = 29.6 bits (65), Expect = 1.2
Identities = 20/61 (32%), Positives = 30/61 (48%), Gaps = 10/61 (16%)
Frame = -3
Query: 273 ATSHKSDSDSVPAPVKGENL--------TAEENEKLAKEEKR--LRNKVASLIKKQKKQQ 322
AT + D DS G + +EN+K KEEK+ +NKV +KK+K++
Sbjct: 317 ATDPEEDGDSSSGSETGSPVDKKAARKEARKENKKKVKEEKKEARKNKVPKAVKKRKQKM 138
Query: 323 A 323
A
Sbjct: 137 A 135
>TC14470 similar to UP|AAQ56728 (AAQ56728) Polygalacturonase inhibiting
protein, partial (93%)
Length = 1195
Score = 29.6 bits (65), Expect = 1.2
Identities = 23/78 (29%), Positives = 34/78 (43%), Gaps = 4/78 (5%)
Frame = +3
Query: 17 PSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSNFSFS----EHPFSFSS 72
P +S L ++ +F +L S V T L R+ +F E P S S
Sbjct: 618 PIPTSLGLIDPNRIDFSRNMLEGDASMLFGVNKTTEMIDLSRNLLTFDMSKVELPTSLIS 797
Query: 73 ISLNHFRTYGSVAAEAIE 90
+ LNH + YGS+ E I+
Sbjct: 798 LDLNHNQIYGSIPQELIK 851
>CB829109
Length = 514
Score = 29.3 bits (64), Expect = 1.6
Identities = 15/48 (31%), Positives = 27/48 (56%)
Frame = +2
Query: 85 AAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNN 132
AA + ST ++ S S ++Q ++ KK+P+++E K+N NN
Sbjct: 350 AARSTGSTGANEESSRSHAILQLVV--------KKHPEVKESKRNNNN 469
>TC15605
Length = 576
Score = 29.3 bits (64), Expect = 1.6
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +3
Query: 361 PANKFGDGGPDIFPAFKHTLKTISNDSNNGSRRYG 395
PAN F DGG FPA +IS+ NN + +G
Sbjct: 225 PANPFADGGFGAFPA-----NSISHPQNNNNNPFG 314
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 28.9 bits (63), Expect = 2.1
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = +3
Query: 116 EEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETE 149
+EKK +L++KKK K K K ++R++ E E
Sbjct: 156 QEKKKKKLKKKKKKKKKKKKKKKKKQRKMVAEAE 257
>TC20132 similar to UP|Q7X9L1 (Q7X9L1) Holocarboxylase synthetase
(Fragment), partial (11%)
Length = 524
Score = 28.5 bits (62), Expect = 2.8
Identities = 10/30 (33%), Positives = 20/30 (66%)
Frame = -3
Query: 120 NPQLEEKKKNKNNSYKYKMLRRRQIKIETE 149
NP E++ K + YK+K+L++++ K+ E
Sbjct: 519 NPDFEKEGSGKQSRYKFKILKKKEKKLYKE 430
>AU251952
Length = 323
Score = 28.5 bits (62), Expect = 2.8
Identities = 15/44 (34%), Positives = 28/44 (63%)
Frame = -1
Query: 290 ENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQA 333
EN +N K K++K+L+N+ + KQK++ ++ KGRD++
Sbjct: 200 ENKKTTKNHKKKKKKKKLQNR----LTKQKEEGSVLDQKGRDKS 81
>TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28040
{Arabidopsis thaliana;}, partial (8%)
Length = 937
Score = 28.1 bits (61), Expect = 3.6
Identities = 21/90 (23%), Positives = 35/90 (38%), Gaps = 9/90 (10%)
Frame = -3
Query: 275 SHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKV-----ASLIKKQKKQQALGIVKG 329
+HK + PAP K A E+ A + KR + K A+ ++ + G G
Sbjct: 521 THKPKAQPSPAPAKSAAKRAAESNANAGDSKRAKKKAVDSAPAAAAGSDEEMEEDGKKSG 342
Query: 330 RDQAKP----WGQEAQVKVGSRLIQLLIET 355
D K W E ++ + L + +T
Sbjct: 341 NDSKKQFTRLWSDEDEIAILKGLADFISKT 252
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 28.1 bits (61), Expect = 3.6
Identities = 13/22 (59%), Positives = 17/22 (77%)
Frame = +2
Query: 10 SSRRFILPSQSSTSLRFSHQHN 31
+S +FIL S SSTS+ FS+Q N
Sbjct: 95 NSSKFILRSTSSTSINFSNQFN 160
>AV768195
Length = 334
Score = 27.7 bits (60), Expect = 4.7
Identities = 20/65 (30%), Positives = 31/65 (46%), Gaps = 11/65 (16%)
Frame = -1
Query: 26 FSHQHNFPEKILRHPFSRFSQV----GIFTSEYK-----LGRSNFSFS--EHPFSFSSIS 74
F HQH P +L + FS+ V G Y +GR NFS++ H F+F+++
Sbjct: 196 FGHQHGSPS*LLEYVFSQDHYVYYV*GNSFKNYS*DKIMMGRRNFSWTSVHHDFTFTAMF 17
Query: 75 LNHFR 79
F+
Sbjct: 16 KRVFK 2
>AV780944
Length = 542
Score = 26.9 bits (58), Expect = 8.0
Identities = 20/84 (23%), Positives = 33/84 (38%), Gaps = 6/84 (7%)
Frame = -3
Query: 1 MWSKLAKRASSRRFILPSQSSTSLRFSHQHNF---PEKILRHPFSRFSQVGIFTSEYKLG 57
+W L RAS + P SS H H P + ++ P + Y L
Sbjct: 429 LWEDLQVRASLYEQVQPKLSSLHAMNQHPHQVKTAPTQRMKPPHKGKMSIAKLCVRYLLL 250
Query: 58 RSN---FSFSEHPFSFSSISLNHF 78
+++ + + H + S +SL HF
Sbjct: 249 QTHAQQYPYPSHQITTSQLSLLHF 178
>TC12713
Length = 412
Score = 26.9 bits (58), Expect = 8.0
Identities = 30/111 (27%), Positives = 45/111 (40%), Gaps = 2/111 (1%)
Frame = +3
Query: 63 FSEHPFSFSSISLNHFRTY--GSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKN 120
F+EHP +SI LN+ R GS+ C +VV+ L
Sbjct: 27 FTEHPLRKASIFLNNTRKSRKGSLRKNKFMGV-----CGSRPKVVEDL------------ 155
Query: 121 PQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLA 171
+ KKNK+ + ++LRRR + EA +AA + L R +A
Sbjct: 156 ----KLKKNKHRRRRRRILRRRVSSQKIEA--DAAHSHSVLQTSNRASDVA 290
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,244,909
Number of Sequences: 28460
Number of extensions: 106320
Number of successful extensions: 614
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 607
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 614
length of query: 515
length of database: 4,897,600
effective HSP length: 94
effective length of query: 421
effective length of database: 2,222,360
effective search space: 935613560
effective search space used: 935613560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144726.11


