

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
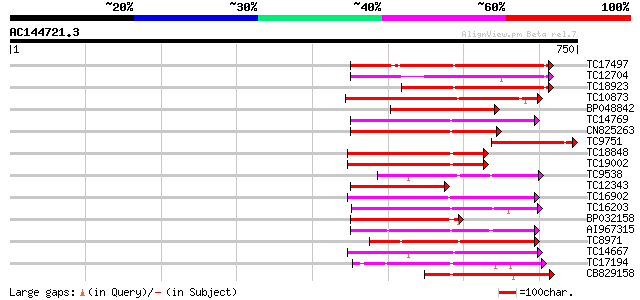
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 285 1e-77
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 243 1e-64
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 225 2e-59
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 198 3e-51
BP048842 189 1e-48
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 184 6e-47
CN825263 174 5e-44
TC9751 weakly similar to UP|O23743 (O23743) SFR1 protein precurs... 172 2e-43
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 166 1e-41
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 165 3e-41
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 160 5e-40
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 159 2e-39
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 156 1e-38
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 153 8e-38
BP032158 150 9e-37
AI967315 148 4e-36
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 147 5e-36
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 144 4e-35
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 144 7e-35
CB829158 136 1e-32
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 285 bits (730), Expect = 1e-77
Identities = 149/269 (55%), Positives = 191/269 (70%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ +GQEIAVKRLS SGQG+EEF NE+ +I++LQHRNLV+L GC V + E N
Sbjct: 1729 LANGQEIAVKRLSNTSGQGMEEFKNEIKLIARLQHRNLVKLFGCSVHQDENSHA-----N 1893
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
K + L D + K +DW KR I++GIARG++YLH+DSRL+IIHRDLK SNILLDDEM
Sbjct: 1894 KKMK-ILLDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDDEMN 2070
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLARI G + EA TKRV+GTYGYMPPEYA+ G FS KSDV+SFGV++LEI+S
Sbjct: 2071 PKISDFGLARIFIGDQV-EARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIIS 2247
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++ FY L+L+ AW+LW+EE + L+D + D + +LR +H+ LLCVQ
Sbjct: 2248 GKKIGRFYDPHHHLNLLSHAWRLWIEERPLELVDELLDDPVIPTEILRYIHVALLCVQRR 2427
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAF 719
P+ RP + ++VLML E LP P AF
Sbjct: 2428 PENRPDMLSIVLMLNGE-KELPKPRLPAF 2511
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 243 bits (619), Expect = 1e-64
Identities = 140/293 (47%), Positives = 177/293 (59%), Gaps = 24/293 (8%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ +GQEIAVKRLS SGQG+EEF NEV +I++LQHRNLV+LLGC + E +L+YEFM N
Sbjct: 1510 LANGQEIAVKRLSNTSGQGMEEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLLIYEFMHN 1689
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
+SLD F+F DSRL+IIHRDLK SNILLD EM
Sbjct: 1690 RSLDYFIF-----------------------------DSRLRIIHRDLKTSNILLDSEMN 1782
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLARI G + EA TKRV+GTYGYM PEYA+ G FS KSDV+SFGV++LEI+S
Sbjct: 1783 PKISDFGLARIFTGDQ-VEAKTKRVMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLEIIS 1959
Query: 631 GRRNNSFYQNEDSLSLVGF------------------------AWKLWLEENTISLIDRE 666
G++ F +L+ AW+LW+EE + L+D
Sbjct: 1960 GKKIGRFCDPHHHRNLLSHSSNFAVFLIKALRICMFENVKNRKAWRLWIEERPLELVDEL 2139
Query: 667 VWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAF 719
+ + + +LR +HI LLCVQ+ P+ RP + +VVLML E LP P AF
Sbjct: 2140 LDGLAIPTEILRYIHIALLCVQQRPEYRPDMLSVVLMLNGE-KELPKPSLPAF 2295
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 225 bits (573), Expect = 2e-59
Identities = 113/201 (56%), Positives = 147/201 (72%)
Frame = +1
Query: 519 DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGL 578
D + K LDW KR I++GIARG++YLH+DSRL+IIHRDLK SNILLD+EM PKISDFGL
Sbjct: 1378 DSTRSKLLDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFGL 1557
Query: 579 ARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFY 638
ARI G + EA TKRV+GTYGYMPPEYA+ G FS KSDV+SFGV++LEI+SG++ FY
Sbjct: 1558 ARIFIGDQV-EARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFY 1734
Query: 639 QNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSIS 698
L+L+ AW+LW+E + + L+D+ D+ + +LR +H+ LLCVQ P+ RP +
Sbjct: 1735 DPHHHLNLLSHAWRLWIEGSPLELVDKLFEDSIIPTEILRYIHVALLCVQRRPETRPDML 1914
Query: 699 TVVLMLISEITHLPPPGKVAF 719
++VLML E LP P AF
Sbjct: 1915 SIVLMLNGE-KELPKPSLPAF 1974
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 198 bits (503), Expect = 3e-51
Identities = 112/266 (42%), Positives = 166/266 (62%), Gaps = 6/266 (2%)
Frame = +1
Query: 445 GRFWPG-IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKML 503
G+ + G + G+ +AVK+LS QG +EF+ EV+++S L H NLVRL+G C + +++L
Sbjct: 487 GKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLMLSLLHHTNLVRLIGYCTDGDQRLL 666
Query: 504 VYEFMPNKSLDAFLFD-PIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASN 562
VYE+MP SL+ LF+ K+ L+W R + G ARG+ YLH + +I+RDLK++N
Sbjct: 667 VYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGAARGLEYLHCTADPPVIYRDLKSAN 846
Query: 563 ILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFG 622
ILLD+E PK+SDFGLA++ G+ +T RV+GTYGY PEYAM G + KSD+YSFG
Sbjct: 847 ILLDNEFNPKLSDFGLAKLGPVGDNTHVST-RVMGTYGYCAPEYAMSGKLTLKSDIYSFG 1023
Query: 623 VLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMH- 681
V+LLE+++GRR + +LV +A + + + + F S RC+H
Sbjct: 1024VVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRRRFGHMVDPLLQGRFPS---RCLHQ 1194
Query: 682 ---IGLLCVQELPKERPSISTVVLML 704
I +C+QE PK RP I+ +V+ L
Sbjct: 1195AIAITAMCLQEQPKFRPLITDIVVAL 1272
>BP048842
Length = 524
Score = 189 bits (481), Expect = 1e-48
Identities = 93/144 (64%), Positives = 116/144 (79%)
Frame = -3
Query: 504 VYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNI 563
+YEFM N+SL+ F+FD + K +DW KR I++GIARG++YLH+DSRL+IIHRDLK SNI
Sbjct: 522 IYEFMHNRSLNYFIFDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNI 343
Query: 564 LLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGV 623
LLDDEM PKISDFGLARI G + EA TKRV+GTYGYMPPEYA+ G FS KSDV+SFGV
Sbjct: 342 LLDDEMNPKISDFGLARIFIGDQ-VEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGV 166
Query: 624 LLLEIVSGRRNNSFYQNEDSLSLV 647
++LEI+SG++ FY L+L+
Sbjct: 165 IVLEIISGKKIGRFYDPHHHLNLL 94
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 184 bits (466), Expect = 6e-47
Identities = 107/252 (42%), Positives = 146/252 (57%), Gaps = 1/252 (0%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DGQE+AVK S S QG EF NE+ ++S +QH NLV LLG C E +++LVY FM N
Sbjct: 2220 LNDGQEVAVKVRSSTSTQGTREFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSN 2399
Query: 511 KSLDAFLF-DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SL L+ +P ++K LDW R +I G ARG+ YLH +IHRD+K+SNILLD M
Sbjct: 2400 GSLQDRLYGEPAKRKILDWPTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSM 2579
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
K++DFG ++ EGD + V GT GY+ PEY SEKSDV+SFGV+LLEIV
Sbjct: 2580 CAKVADFGFSKYAP-QEGDSYVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIV 2756
Query: 630 SGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQE 689
SGR + + SLV +A ++D + +M R + + L C++
Sbjct: 2757 SGREPLNIKRPRTEWSLVEWATPYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEP 2936
Query: 690 LPKERPSISTVV 701
RPS+ +V
Sbjct: 2937 FSTYRPSMVAIV 2972
>CN825263
Length = 663
Score = 174 bits (441), Expect = 5e-44
Identities = 91/201 (45%), Positives = 133/201 (65%), Gaps = 1/201 (0%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+++AVK L + +G EF+ EV ++S+L HRNLV+L+G C+E+ + L+YE +PN
Sbjct: 28 LNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTRCLIYELVPN 207
Query: 511 KSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
S+++ L ++ LDW R I G ARG+ YLH DS +IHRD K+SNILL+ +
Sbjct: 208 GSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKSSNILLECDF 387
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
PK+SDFGLAR EG++ + V+GT+GY+ PEYAM G KSDVYS+GV+LLE++
Sbjct: 388 TPKVSDFGLARTAL-DEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSYGVVLLELL 564
Query: 630 SGRRNNSFYQNEDSLSLVGFA 650
+G + Q +LV +A
Sbjct: 565 TGTKPVDLSQPPGQENLVTWA 627
>TC9751 weakly similar to UP|O23743 (O23743) SFR1 protein precursor ,
partial (5%)
Length = 550
Score = 172 bits (435), Expect = 2e-43
Identities = 83/113 (73%), Positives = 101/113 (88%)
Frame = +1
Query: 638 YQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSI 697
++N+DSLSL+GFAWKLW EEN ISLID E+WD S ESSMLRC+HIGLLCVQE+P+ERP+I
Sbjct: 1 FRNDDSLSLLGFAWKLWTEENIISLIDPEIWDPSVESSMLRCIHIGLLCVQEIPRERPAI 180
Query: 698 STVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
STVVLMLISEIT LPPPG+VAFV N++S++SSQ+S + +SNNNVTLS+V G
Sbjct: 181 STVVLMLISEITQLPPPGQVAFVQKHNAKSSDSSQKS-QCSSNNNVTLSEVHG 336
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 166 bits (421), Expect = 1e-41
Identities = 86/188 (45%), Positives = 124/188 (65%), Gaps = 1/188 (0%)
Frame = +2
Query: 447 FWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYE 506
+W DG +IAVK+L + + EF EV V+ +++H+NL+ L G CV ++++VY+
Sbjct: 347 YWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEVLGRVRHKNLLGLRGYCVGDDQRLIVYD 526
Query: 507 FMPNKSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILL 565
+MPN SL + L + +L+W+KR I G A GI+YLH + IIHRD+KASN+LL
Sbjct: 527 YMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGSAEGILYLHHEVTPHIIHRDIKASNVLL 706
Query: 566 DDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLL 625
+ + P ++DFG A+++ EG T RV GT GY+ PEYAM G SE DVYSFG+LL
Sbjct: 707 NSDFEPLVADFGFAKLIP--EGVSHMTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILL 880
Query: 626 LEIVSGRR 633
LE+V+GR+
Sbjct: 881 LELVTGRK 904
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 165 bits (417), Expect = 3e-41
Identities = 85/188 (45%), Positives = 124/188 (65%), Gaps = 1/188 (0%)
Frame = +1
Query: 447 FWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYE 506
+W + DG +IAVKRL S + EF EV ++++++H+NL+ L G C E E+++VY+
Sbjct: 109 YWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEILARVRHKNLLSLRGYCAEGQERLIVYD 288
Query: 507 FMPNKSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILL 565
+MPN SL + L + LDW +R NI G A GI+YLH + IIHRD+KASN+LL
Sbjct: 289 YMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGSAEGIVYLHHQATPHIIHRDIKASNVLL 468
Query: 566 DDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLL 625
D + +++DFG A+++ +G T RV GT GY+ PEYAM G +E DV+SFG+LL
Sbjct: 469 DSDFQARVADFGFAKLIP--DGATHVTTRVKGTLGYLAPEYAMLGKANECCDVFSFGILL 642
Query: 626 LEIVSGRR 633
LE+ SG++
Sbjct: 643 LELASGKK 666
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 160 bits (406), Expect = 5e-40
Identities = 94/232 (40%), Positives = 137/232 (58%), Gaps = 12/232 (5%)
Frame = +2
Query: 487 NLVRLLGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKK----------LDWRKRSNIVE 536
N+VRLL C +LVYE++ N SLD +L + LDW KR I
Sbjct: 5 NIVRLLCCISNEASMLLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAI 184
Query: 537 GIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLAR-IVKGGEGDEANTKRV 595
G A+G+ Y+H D I+HRD+K SNILLD + K++DFGLAR ++K GE + +T V
Sbjct: 185 GAAQGLSYMHHDCSPPIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMST--V 358
Query: 596 VGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWK-LW 654
+GT+GY+ PEY SEK DVYSFGV+LLE+ +G+ N Y ++ S SL +AW+ +
Sbjct: 359 IGTFGYIAPEYVQTTRISEKVDVYSFGVVLLELTTGKEAN--YGDQHS-SLAEWAWRHIL 529
Query: 655 LEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLIS 706
+ N L+ ++V +AS+ M +G++C LP RPS+ V+ +L+S
Sbjct: 530 IGSNVXDLLXKDVMEASYIDEMCSVFKLGVMCTATLPATRPSMKEVLPILLS 685
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 159 bits (402), Expect = 2e-39
Identities = 72/131 (54%), Positives = 104/131 (78%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+EIAVK+LS+ S QG +F+NE +++++QHRN+V L G C EK+LVYE++P
Sbjct: 300 LNDGREIAVKKLSRRSNQGRTQFINEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPR 479
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
+SLD LF +K++LDW++R +I+ G+ARG++YLH DS IIHRD+KA+NILLD++ +
Sbjct: 480 ESLDKLLFRSQKKEQLDWKRRFDIISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWV 659
Query: 571 PKISDFGLARI 581
PKI+DFGLARI
Sbjct: 660 PKIADFGLARI 692
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 156 bits (395), Expect = 1e-38
Identities = 92/256 (35%), Positives = 146/256 (56%), Gaps = 1/256 (0%)
Frame = +3
Query: 447 FWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYE 506
++ + G ++A+KR + S QG+ EF E+ ++SKL+HR+LV L+G C E E +LVY+
Sbjct: 42 YYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEENTEMILVYD 221
Query: 507 FMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLD 566
M +L L+ QK L W++R I G ARG+ YLH ++ IIHRD+K +NILLD
Sbjct: 222 HMAYGTLREHLYKT-QKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRDVKTTNILLD 398
Query: 567 DEMIPKISDFGLARIVKGGEGDEANTKRVV-GTYGYMPPEYAMGGLFSEKSDVYSFGVLL 625
++ + K+SDFGL++ G D + VV G++GY+ PEY ++KSDVYSFGV+L
Sbjct: 399 EKWVAKVSDFGLSK--TGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDVYSFGVVL 572
Query: 626 LEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLL 685
EI+ R + ++ +SL +A + + ++D + + +
Sbjct: 573 FEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFKKFAETAMK 752
Query: 686 CVQELPKERPSISTVV 701
CV + ERPS+ V+
Sbjct: 753 CVSDQGIERPSMGDVL 800
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 153 bits (387), Expect = 8e-38
Identities = 93/260 (35%), Positives = 149/260 (56%), Gaps = 8/260 (3%)
Frame = +2
Query: 453 DGQEIAVKRL-SKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNK 511
+G ++A+KRL + SG+ F E+ + K++HRN++RLLG + +L+YE+MPN
Sbjct: 2255 NGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGYVSNKDTNLLLYEYMPNG 2434
Query: 512 SLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIP 571
SL +L + L W R I ARG+ Y+H D IIHRD+K++NILLD +
Sbjct: 2435 SLGEWLHGA-KGGHLRWEMRYKIAVEAARGLCYMHHDCSPLIIHRDVKSNNILLDADFEA 2611
Query: 572 KISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSG 631
++DFGLA+ + G + + G+YGY+ PEYA EKSDVYSFGV+LLE++ G
Sbjct: 2612 HVADFGLAKFLY-DPGASQSMSSIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIG 2788
Query: 632 RRNNSFYQNEDSLSLVGFAWKLWLEEN-------TISLIDREVWDASFESSMLRCMHIGL 684
R+ + D + +VG+ K E + ++++D + +S++ +I +
Sbjct: 2789 RKPVGEF--GDGVDIVGWVNKTMSELSQPSDTALVLAVVDPRLSGYPL-TSVIHMFNIAM 2959
Query: 685 LCVQELPKERPSISTVVLML 704
+CV+E+ RP++ VV ML
Sbjct: 2960 MCVKEMGPARPTMREVVHML 3019
>BP032158
Length = 555
Score = 150 bits (378), Expect = 9e-37
Identities = 80/154 (51%), Positives = 106/154 (67%), Gaps = 4/154 (2%)
Frame = +2
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ +G IAVK+LS S QG EF+NE+ +IS LQH NLV+L GCC+E + +LVYE+M N
Sbjct: 107 LSEGDVIAVKQLSSKSKQGNREFINEIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMEN 286
Query: 511 KSLDAFLF-DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SL LF + QK L+WR R I GIA+G+ YLH +SRLKI+HRD+KA+N+LLD ++
Sbjct: 287 NSLARALFGNEEQKLNLNWRTRMKICVGIAKGLAYLHEESRLKIVHRDIKATNVLLDKDL 466
Query: 570 IPKISDFGLARIVKGGEGDEANT---KRVVGTYG 600
KISDFGLA++ +E NT R+ GT G
Sbjct: 467 KAKISDFGLAKL-----DEEENTHISTRIAGTIG 553
>AI967315
Length = 1308
Score = 148 bits (373), Expect = 4e-36
Identities = 91/253 (35%), Positives = 145/253 (56%), Gaps = 2/253 (0%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKA--SGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFM 508
++ G EIAVKRL++ + +EF+ E+ I + H N++ LLGCC++ G LV+E
Sbjct: 205 LESGDEIAVKRLTRTCRDERKEKEFLTEIGTIGHVCHSNVMPLLGCCIDNG-LYLVFELS 381
Query: 509 PNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDE 568
S+ + + D + LDW+ R IV G ARG+ YLH+ + +IIHRD+KASNILL ++
Sbjct: 382 TVGSVASLIHDE-KMAPLDWKTRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNILLTED 558
Query: 569 MIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEI 628
P+ISDFGLA+ + + + + GT+G++ PEY M G+ EK+DV++FGV LLE+
Sbjct: 559 FEPQISDFGLAKWLP-SQWTHHSIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEV 735
Query: 629 VSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQ 688
+SGR+ + SL +A + + L+D + + R LC++
Sbjct: 736 ISGRKP----VDGSHQSLHTWAKPILSKWEIEKLVDPRLEGCYDVTQFNRVAFAASLCIR 903
Query: 689 ELPKERPSISTVV 701
RP++S V+
Sbjct: 904 ASSTWRPTMSEVL 942
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 147 bits (372), Expect = 5e-36
Identities = 84/226 (37%), Positives = 138/226 (60%), Gaps = 1/226 (0%)
Frame = +3
Query: 476 EVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIV 535
EV +++K+ HRNLV+LLG + E++L+ E++PN +L L D ++ K LD+ +R I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL-DGLRGKILDFNQRLEIA 182
Query: 536 EGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVK-GGEGDEANTKR 594
+A G+ YLH + +IIHRD+K+SNILL + M K++DFG AR+ G+ +TK
Sbjct: 183 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK- 359
Query: 595 VVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLW 654
V GT GY+ PEY + KSDVYSFG+LLLEI++GRR + D + +A++ +
Sbjct: 360 VKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY 539
Query: 655 LEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTV 700
E + + L+D + +A +++ + + C + +RP++ +V
Sbjct: 540 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 677
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 144 bits (364), Expect = 4e-35
Identities = 93/265 (35%), Positives = 145/265 (54%), Gaps = 7/265 (2%)
Frame = +2
Query: 447 FWPGIQDGQEIAVKRLSKASG-QGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVY 505
++ + DG +AVK+L +S + EF+ +V ++S+L++ N V L G CVE ++L Y
Sbjct: 404 YYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSMVSRLKNDNFVELHGYCVEGNLRVLAY 583
Query: 506 EFMPNKSLDAFLFDP--IQKKK----LDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EF SL L +Q + LDW +R I ARG+ YLH + IIHRD++
Sbjct: 584 EFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAVDAARGLEYLHEKVQPAIIHRDIR 763
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
+SN+L+ ++ KI+DF L+ ++ RV+GT+GY PEYAM G ++KSDVY
Sbjct: 764 SSNVLIFEDYKAKIADFNLSNQAPDMAA-RLHSTRVLGTFGYHAPEYAMTGQLTQKSDVY 940
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
SFGV+LLE+++GR+ SLV +A E+ +D ++ + +
Sbjct: 941 SFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSEDKVKQCVDPKLKGEYPPKGVAKL 1120
Query: 680 MHIGLLCVQELPKERPSISTVVLML 704
+ LCVQ + RP++S VV L
Sbjct: 1121AAVAALCVQYEAEFRPNMSIVVKAL 1195
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 144 bits (362), Expect = 7e-35
Identities = 95/265 (35%), Positives = 149/265 (55%), Gaps = 9/265 (3%)
Frame = +1
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G++ A+K++ Q EF+ E+ V++ + H NLVRL+G CVE G LVYE + N +L
Sbjct: 1150 GKKTAIKKMDV---QASTEFLCELKVLTHVHHLNLVRLIGYCVE-GSLFLVYEHIDNGNL 1317
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
+L K+ L W R I ARG+ Y+H + IHRD+K++NIL+D + K+
Sbjct: 1318 GQYLHGS-GKEPLPWSSRVQIALDAARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKV 1494
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
+DFGL ++++ G+ R+VGT+GYMPPEYA G S K DVY+FGV+L E++S +
Sbjct: 1495 ADFGLTKLIE--VGNSTLQTRLVGTFGYMPPEYAQYGDISPKIDVYAFGVVLFELISA-K 1665
Query: 634 NNSFYQNE---DSLSLVGFAWKLWLEENTI----SLIDREVWDASFESSMLRCMHIGLLC 686
N E +S LV + + + L+D + + S+L+ +G C
Sbjct: 1666 NAVLKTGELVAESKGLVALFEEALNKSDPCDALRKLVDPRLGENYPIDSVLKIAQLGRAC 1845
Query: 687 VQELPKERPSISTVV--LMLISEIT 709
++ P RPS+ ++V LM +S +T
Sbjct: 1846 TRDNPLLRPSMRSLVVALMTLSSLT 1920
>CB829158
Length = 553
Score = 136 bits (343), Expect = 1e-32
Identities = 78/177 (44%), Positives = 110/177 (62%), Gaps = 5/177 (2%)
Frame = +1
Query: 549 SRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAM 608
S+++IIHRD+K SNILLD ++ KI+DFGLAR + E + + GT GYM PEY
Sbjct: 1 SKVRIIHRDIKVSNILLDAKLRAKIADFGLARSFQ--EDKSHISTAIAGTLGYMAPEYLA 174
Query: 609 GGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLID---- 664
G +EK DVYSFGVLLLEIV+GR+NN ++E + SL+ AW+ + L D
Sbjct: 175 HGQLTEKVDVYSFGVLLLEIVTGRQNNRSKESEYTDSLILVAWEHFQTGTAEQLFDPNIE 354
Query: 665 -REVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFV 720
E + + ++ +LR +HIGLLC+QE+P RP++S V+ ML + L P F+
Sbjct: 355 LHEAHNGNVKNEILRVIHIGLLCIQEIPSLRPTMSKVLQMLTKKEELLVAPSNPPFL 525
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,786,472
Number of Sequences: 28460
Number of extensions: 229075
Number of successful extensions: 3275
Number of sequences better than 10.0: 298
Number of HSP's better than 10.0 without gapping: 2395
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 659
Number of HSP's gapped (non-prelim): 2556
length of query: 750
length of database: 4,897,600
effective HSP length: 97
effective length of query: 653
effective length of database: 2,136,980
effective search space: 1395447940
effective search space used: 1395447940
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144721.3


