

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.14 - phase: 0
(234 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
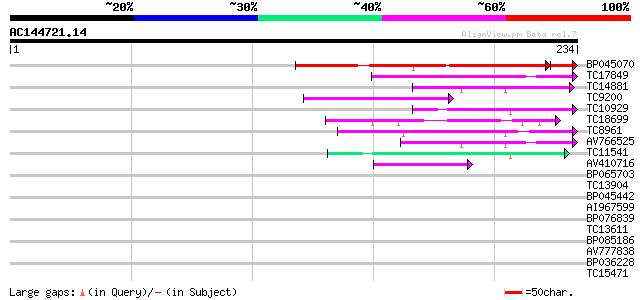
Sequences producing significant alignments: (bits) Value
BP045070 123 3e-30
TC17849 49 1e-06
TC14881 46 6e-06
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 43 4e-05
TC10929 similar to UP|Q94AX6 (Q94AX6) AT3g23280/K14B15_17, parti... 43 5e-05
TC18699 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromo... 41 2e-04
TC8961 similar to GB|BAB71852.1|16902294|AB062739 kinesin-relate... 41 2e-04
AV766525 40 3e-04
TC11541 weakly similar to UP|O64756 (O64756) At2g34920 protein, ... 40 5e-04
AV410716 40 5e-04
BP065703 38 0.002
TC13904 weakly similar to UP|Q9ZU51 (Q9ZU51) RING-H2 finger prot... 37 0.002
BP045442 36 0.005
AI967599 36 0.007
BP076839 36 0.007
TC13611 weakly similar to UP|AAQ82840 (AAQ82840) At1g30860, part... 35 0.009
BP085186 35 0.011
AV777838 35 0.015
BP036228 35 0.015
TC15471 35 0.015
>BP045070
Length = 386
Score = 123 bits (308), Expect(2) = 3e-30
Identities = 62/108 (57%), Positives = 79/108 (72%), Gaps = 3/108 (2%)
Frame = -3
Query: 119 GEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREE---EEEREEEGEG 175
GE + V+MSLM++LEE+E++ E V GDD + ++E +++ EE EE+G
Sbjct: 378 GEATTTVGLVNMSLMEMLEETEVDFE----VRVNGDDEDNDDECQKQGGGEEGEEEQGGV 211
Query: 176 EGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLC 223
E + V V +NCCVCMVR KGAAFIPCGHTFCRMCSRELWV+RGNCPLC
Sbjct: 210 EETGV-VAYNCCVCMVRHKGAAFIPCGHTFCRMCSRELWVNRGNCPLC 70
Score = 24.3 bits (51), Expect(2) = 3e-30
Identities = 9/11 (81%), Positives = 11/11 (99%)
Frame = -2
Query: 224 NHFILEVLDIF 234
+HFILE+LDIF
Sbjct: 70 HHFILEILDIF 38
>TC17849
Length = 589
Score = 48.5 bits (114), Expect = 1e-06
Identities = 23/85 (27%), Positives = 38/85 (44%)
Frame = +2
Query: 150 VDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMC 209
+D ++++E+ E+E+ E E K C VC+ + +PCGH CR C
Sbjct: 59 IDLQENLKESQPAFLYEQEKVERATKEADTAKAAWVCRVCLSAEVDITIVPCGHVLCRKC 238
Query: 210 SRELWVSRGNCPLCNHFILEVLDIF 234
S + CP C + + + IF
Sbjct: 239 SSAV----SKCPFCRLQVTKAIRIF 301
>TC14881
Length = 609
Score = 45.8 bits (107), Expect = 6e-06
Identities = 24/73 (32%), Positives = 35/73 (47%), Gaps = 6/73 (8%)
Frame = +3
Query: 167 EEREEEGEGEGSNVKVEHN-----CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNC 220
E RE G G + + N C +CM K A +PC H C C++EL + C
Sbjct: 27 ELRELYGIGSSTAADFDDNDPQKECVICMTEPKDTAVLPCRHMCMCSDCAKELRLQSNKC 206
Query: 221 PLCNHFILEVLDI 233
P+C I E+++I
Sbjct: 207 PICRQPIEELIEI 245
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 43.1 bits (100), Expect = 4e-05
Identities = 23/62 (37%), Positives = 34/62 (54%)
Frame = +1
Query: 122 SDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVK 181
S ++A++ L + LE E E + + V+E + EE E+ EEE E EEE E E V+
Sbjct: 133 SSQSAELPFKLQNKLEAEVQEYEEVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVE 312
Query: 182 VE 183
E
Sbjct: 313 AE 318
>TC10929 similar to UP|Q94AX6 (Q94AX6) AT3g23280/K14B15_17, partial (16%)
Length = 783
Score = 42.7 bits (99), Expect = 5e-05
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 1/69 (1%)
Frame = +2
Query: 167 EEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNH 225
E+ EEG+ G ++ C +C+ A IPCGH C C E+ + CP+C
Sbjct: 221 EKLPEEGQSAG---EIGSTCVICLDAPAEGACIPCGHVAGCMSCLNEVKTKKWGCPVCRA 391
Query: 226 FILEVLDIF 234
I +V+ ++
Sbjct: 392 KIDQVIKLY 418
>TC18699 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromosome 5, TAC
clone:K17O22 (AT5g45100/K17O22_9), partial (28%)
Length = 537
Score = 40.8 bits (94), Expect = 2e-04
Identities = 33/107 (30%), Positives = 48/107 (44%), Gaps = 10/107 (9%)
Frame = +3
Query: 131 SLMDLLEESEIELERISD-----VVDEGDDVEEN---NEKREEEEEREEEGEGEGSNVKV 182
+++ L + E L +SD +E DD E + N REEEEE E V
Sbjct: 126 AVISLRSDLEQVLAHVSDSHRGAAAEEEDDAESSCGSNHHREEEEEENE---------AV 278
Query: 183 EHNCCVCMVRDKGAAFIPCGHTFCRMCS-RELWVSR-GNCPLCNHFI 227
+ C C ++ G +PC H +C ++WV NCPLC+ I
Sbjct: 279 GNLCRECGAQESGVLLLPCRH----LCL*YDVWVQPIRNCPLCHSAI 407
>TC8961 similar to GB|BAB71852.1|16902294|AB062739 kinesin-related protein
{Arabidopsis thaliana;} , partial (8%)
Length = 735
Score = 40.8 bits (94), Expect = 2e-04
Identities = 28/105 (26%), Positives = 46/105 (43%), Gaps = 6/105 (5%)
Frame = +1
Query: 136 LEESEIELERISDVVDEGDDVEENNE-----KREEEEEREEEGEGEGSNVKVEHNCCVCM 190
+E I E+I D + G+++ + K +E +E+E + G+ H C VC
Sbjct: 88 IENDTIPKEQILDGSEPGNELPKEEPLVVRLKARMQEMKEKELKHLGNGDANSHVCKVCF 267
Query: 191 VRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
A +PC H C+ CS ++ CP+C I + L F
Sbjct: 268 ESSTAAILLPCRHFCLCKSCS----LACSECPICRTSIADRLFAF 390
>AV766525
Length = 564
Score = 40.0 bits (92), Expect = 3e-04
Identities = 24/78 (30%), Positives = 37/78 (46%), Gaps = 5/78 (6%)
Frame = -1
Query: 162 KREEEEEREEEGEGEGSNVKVEHN----CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVS 216
KR + E+ +G + K E C +C+ ++ + F+PCGH C CS L
Sbjct: 498 KRLGHDNEGEKADGLVDSAKRERLIPDLCVICLEKEYNSVFVPCGHMCCCTGCSSHL--- 328
Query: 217 RGNCPLCNHFILEVLDIF 234
+CPLC I +V+ F
Sbjct: 327 -TSCPLCRRQIEKVVKTF 277
>TC11541 weakly similar to UP|O64756 (O64756) At2g34920 protein, partial
(6%)
Length = 553
Score = 39.7 bits (91), Expect = 5e-04
Identities = 27/101 (26%), Positives = 41/101 (39%), Gaps = 1/101 (0%)
Frame = +1
Query: 132 LMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMV 191
+++ + ++EL+R V + N E E +G + CCVC
Sbjct: 22 MLEXCMDMQLELQRS---VRQEVSAALNRSAGENGSVAETSDDGSKWGHVKKGTCCVCCD 192
Query: 192 RDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVL 231
+ CGH C C+ EL G CPLC I+EV+
Sbjct: 193 SHIDSLLYRCGHMCTCSKCANELIRGGGKCPLCRAPIVEVV 315
>AV410716
Length = 424
Score = 39.7 bits (91), Expect = 5e-04
Identities = 18/41 (43%), Positives = 24/41 (57%)
Frame = +2
Query: 151 DEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMV 191
+E ++VEE E+ EEEEE E+ G + V EH C C V
Sbjct: 155 EEEEEVEEEEEEEEEEEEAEKNGG*SRNRVSTEHGCVGCEV 277
>BP065703
Length = 480
Score = 37.7 bits (86), Expect = 0.002
Identities = 26/87 (29%), Positives = 34/87 (38%), Gaps = 18/87 (20%)
Frame = -3
Query: 155 DVEENNEKREEEEERE--------EEGEGEGSNVKVEHN---------CCVCMVRDKGAA 197
D E ++ K+ +EE RE GE + K+E C VC R K
Sbjct: 463 DRERSSRKKLQEELRELNNXIAELSSETGEAAIQKLEEEIRACKNMIKCTVCSDRPKEVV 284
Query: 198 FIPCGHTFCRMC-SRELWVSRGNCPLC 223
+ C H FC C R L + CP C
Sbjct: 283 IVKCYHLFCNPCIQRNLELRHRKCPAC 203
>TC13904 weakly similar to UP|Q9ZU51 (Q9ZU51) RING-H2 finger protein RHA2b,
partial (32%)
Length = 535
Score = 37.4 bits (85), Expect = 0.002
Identities = 24/62 (38%), Positives = 30/62 (47%), Gaps = 5/62 (8%)
Frame = +1
Query: 172 EGEGEGSNVKVEHNCCVCMVRDKGAAFI---PCGHTFCRMCSRELWVS--RGNCPLCNHF 226
+G GE HNC VC K + PC HTF R C E W+ + NCPLC +
Sbjct: 1 DGAGE------RHNCVVCQRTFKDGDQVRRLPCRHTFHRRCF-EGWLHHYKFNCPLCRYS 159
Query: 227 IL 228
+L
Sbjct: 160 LL 165
>BP045442
Length = 488
Score = 36.2 bits (82), Expect = 0.005
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 5/46 (10%)
Frame = -1
Query: 183 EHNCCVCMVRDKGAAFI---PCGHTFCRMCSRELWVSRGN--CPLC 223
EH C VC+ + + + I PCGH F + C E W+ N CPLC
Sbjct: 446 EHECSVCLTKFEPESEINCLPCGHLFHKAC-LEKWLDYWNITCPLC 312
>AI967599
Length = 412
Score = 35.8 bits (81), Expect = 0.007
Identities = 22/79 (27%), Positives = 34/79 (42%), Gaps = 2/79 (2%)
Frame = +1
Query: 158 ENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-TFCRMCSRELW-V 215
E + +E + E+ E E S+ C +CM + F+PC H C CS E
Sbjct: 1 EFGTRLQELDNLEDMSEKEVSS---NRECIICMKDEVSVVFLPCAHQVMCASCSDEYGRK 171
Query: 216 SRGNCPLCNHFILEVLDIF 234
+ CP C I + + +F
Sbjct: 172 GKAACPCCRVQIQQRIRVF 228
>BP076839
Length = 392
Score = 35.8 bits (81), Expect = 0.007
Identities = 17/47 (36%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Frame = -3
Query: 186 CCVCM-VRDKGAAF--IPCGHTFCRMCSRELWVSRGNCPLCNHFILE 229
CC+C+ D G +PCGH F C + CPLC + IL+
Sbjct: 360 CCICLSAYDDGVELRQLPCGHHFHCACVDKWLCMNATCPLCKYNILK 220
>TC13611 weakly similar to UP|AAQ82840 (AAQ82840) At1g30860, partial (7%)
Length = 519
Score = 35.4 bits (80), Expect = 0.009
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Frame = +3
Query: 186 CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVL 231
CCVC + CGH C C+ EL + CP+C ++EV+
Sbjct: 147 CCVCCESHIDSLLYRCGHLCTCSKCANELLPNGRKCPMCPAPVVEVI 287
>BP085186
Length = 388
Score = 35.0 bits (79), Expect = 0.011
Identities = 18/46 (39%), Positives = 25/46 (54%)
Frame = -1
Query: 137 EESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKV 182
EE E E E + +E ++ E EK EEE +EGEGE ++V
Sbjct: 256 EEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQEGEGENP*LQV 119
Score = 35.0 bits (79), Expect = 0.011
Identities = 17/32 (53%), Positives = 20/32 (62%)
Frame = -1
Query: 152 EGDDVEENNEKREEEEEREEEGEGEGSNVKVE 183
EG++ EE E+ EEEEE EEE E E K E
Sbjct: 268 EGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNE 173
Score = 30.8 bits (68), Expect = 0.21
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = -1
Query: 152 EGDDVEENNEKREEEEEREEEGEGE 176
+G+ EE E+ EEEEE EEE E E
Sbjct: 274 DGEGNEEEEEEEEEEEEEEEEEEEE 200
Score = 30.4 bits (67), Expect = 0.28
Identities = 20/47 (42%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Frame = -1
Query: 137 EESEIELERISDVVDEGDDVEENNEKREEEEEREEEG----EGEGSN 179
EE E E E +E ++ EE E EEE+ EEE EGEG N
Sbjct: 259 EEEEEEEEE-----EEEEEEEEEEEAEPEEEKNEEENVNTQEGEGEN 134
Score = 30.0 bits (66), Expect = 0.36
Identities = 15/29 (51%), Positives = 17/29 (57%)
Frame = -1
Query: 155 DVEENNEKREEEEEREEEGEGEGSNVKVE 183
D E N E+ EEEEE EEE E E + E
Sbjct: 274 DGEGNEEEEEEEEEEEEEEEEEEEEAEPE 188
Score = 28.9 bits (63), Expect = 0.80
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = -1
Query: 158 ENNEKREEEEEREEEGEGEGSNVKVEHN 185
E E+ EEEEE EEE E E + + E N
Sbjct: 259 EEEEEEEEEEEEEEEEEEEEAEPEEEKN 176
Score = 28.5 bits (62), Expect = 1.0
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = -1
Query: 148 DVVDEGDDVEENNEKREEEEEREEEGE 174
D D + EE E+ EEEEE EEE E
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEEE 203
Score = 27.3 bits (59), Expect = 2.3
Identities = 13/26 (50%), Positives = 14/26 (53%)
Frame = -1
Query: 151 DEGDDVEENNEKREEEEEREEEGEGE 176
D D E+ EEEEE EEE E E
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEE 206
Score = 26.2 bits (56), Expect = 5.2
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = -1
Query: 154 DDVEENNEKREEEEEREEEGEGEGSNVKVE 183
D+ + + EEEEE EEE E E + E
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEEEEAE 194
>AV777838
Length = 565
Score = 34.7 bits (78), Expect = 0.015
Identities = 24/66 (36%), Positives = 31/66 (46%)
Frame = -3
Query: 117 PGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
P G +E + + L+EE E E + V + E E+ EEEEE EEE E E
Sbjct: 395 PLGSCDEEEEVMV*PAVALMEEEEEEGGAVMVVWPAAVALMEEEEEEEEEEEEEEEEEEE 216
Query: 177 GSNVKV 182
G V V
Sbjct: 215 GGPVMV 198
>BP036228
Length = 626
Score = 34.7 bits (78), Expect = 0.015
Identities = 17/45 (37%), Positives = 19/45 (41%), Gaps = 2/45 (4%)
Frame = +2
Query: 185 NCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRG--NCPLCNHFI 227
NC CM PCGH FC C E W+ +G C C I
Sbjct: 419 NCSFCMQLPDRPVTTPCGHNFCLKCF-EKWIGQGKRTCANCRRDI 550
>TC15471
Length = 642
Score = 34.7 bits (78), Expect = 0.015
Identities = 13/49 (26%), Positives = 23/49 (46%), Gaps = 1/49 (2%)
Frame = +3
Query: 186 CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVLDI 233
C +C+ + +PC H C C++ L CP+C + +L+I
Sbjct: 174 CVICLSEPRDTTVLPCRHMCMCSGCAKVLRFQTNRCPICRQPVERLLEI 320
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,064,424
Number of Sequences: 28460
Number of extensions: 59964
Number of successful extensions: 1093
Number of sequences better than 10.0: 195
Number of HSP's better than 10.0 without gapping: 738
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 909
length of query: 234
length of database: 4,897,600
effective HSP length: 87
effective length of query: 147
effective length of database: 2,421,580
effective search space: 355972260
effective search space used: 355972260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144721.14


