

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144656.5 + phase: 0 /pseudo
(2301 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
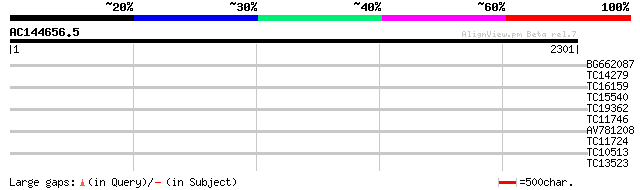
Score E
Sequences producing significant alignments: (bits) Value
BG662087 38 0.016
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 35 0.17
TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, parti... 34 0.23
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 31 2.5
TC19362 30 3.3
TC11746 similar to UP|Q40983 (Q40983) Metalloendopeptidase , par... 30 4.2
AV781208 29 7.2
TC11724 weakly similar to UP|Q9M1Q2 (Q9M1Q2) Serine/threonine pr... 29 7.2
TC10513 similar to GB|AAN15531.1|23198008|BT000212 expressed pro... 29 9.5
TC13523 homologue to UP|R1AD_ARATH (Q9FKC0) 60S ribosomal protei... 29 9.5
>BG662087
Length = 373
Score = 38.1 bits (87), Expect = 0.016
Identities = 34/113 (30%), Positives = 47/113 (41%)
Frame = +3
Query: 1318 WQSQNVC*LP*P*QS*SKR*FPITSY*CTG*QHCKIQSVLPHGWFLRLQSDQDGARRQRE 1377
W+ +V L Q KR I Y TG + P G LRL +QD R +
Sbjct: 15 WEMAHVGGLHGSEQGMPKRLLSIAKYRQTGGWSVRQ*VAKPDGCLLRLSPNQDAPIR*GQ 194
Query: 1378 DVFHHTLGHFLLQSNAVRFDQCRSYLSKGYDYYLSRHDTQRD*GLCG*HDRQV 1430
+ H G LL N+VR ++CR +S L ++ L HDR+V
Sbjct: 195 NGLHDRQGKLLLPDNSVRVEECRGDISTADG*GLX*PXGEKHGSLPRQHDRKV 353
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 34.7 bits (78), Expect = 0.17
Identities = 32/102 (31%), Positives = 47/102 (45%), Gaps = 2/102 (1%)
Frame = +2
Query: 38 HLLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLL 97
H LH + HLPLL + H +LH++P +HL ++ +PLL +
Sbjct: 71 HHLHTTTNHLHHHLPLLPPHTTTNLPHHHHILHHLPTTTNHLHHHPRH-LPLLT-----I 232
Query: 98 LNMQLMFLRL--LRQFLKLL*HTQHLRFMLSRKITNPFSIQR 137
N+ + L L L K+L H HL F++ TNP QR
Sbjct: 233 TNLPHLHLHLHRLHTTTKVLLH--HLPFLILPTTTNPLHHQR 352
>TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, partial (19%)
Length = 605
Score = 34.3 bits (77), Expect = 0.23
Identities = 33/84 (39%), Positives = 39/84 (46%), Gaps = 2/84 (2%)
Frame = +3
Query: 40 LHLLSELRSKHLPLLSLNGQYVPTLR--HTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLL 97
LH L LR +HL LL Y+P R H L H HLLLL +V LL++ L
Sbjct: 123 LHRLC-LRPRHLWLLPRGNLYLPRQRFLHLLRHR-----RHLLLLRWRAVLLLLRPPSTL 284
Query: 98 LNMQLMFLRLLRQFLKLL*HTQHL 121
L + LM L LL H HL
Sbjct: 285 LRLPLMLLLLLP-------HLSHL 335
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 30.8 bits (68), Expect = 2.5
Identities = 29/67 (43%), Positives = 37/67 (54%), Gaps = 3/67 (4%)
Frame = -2
Query: 54 LSLNGQYVP--TLRHTLLHNVPRLGSHLLLLAKYSVPL-LVKLRCLLLNMQLMFLRLLRQ 110
LSL P TL L H+ P +HLLLL + + L L++LR LLL +QL L LL+
Sbjct: 313 LSLRNHL*PPRTLNICLTHSCP---NHLLLLLRKLLML*LLQLRKLLLRLQLRELLLLKL 143
Query: 111 FLKLL*H 117
LL H
Sbjct: 142 RELLLLH 122
>TC19362
Length = 513
Score = 30.4 bits (67), Expect = 3.3
Identities = 16/50 (32%), Positives = 23/50 (46%)
Frame = +2
Query: 36 PKHLLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHLLLLAKY 85
P H +H S +H P+ S Q +L HN L +H LLL+ +
Sbjct: 98 PHHHVHFFSSFNVRHTPITSSFSQAPFSLLQHHTHNPSSLKTHFLLLSPW 247
>TC11746 similar to UP|Q40983 (Q40983) Metalloendopeptidase , partial (3%)
Length = 535
Score = 30.0 bits (66), Expect = 4.2
Identities = 26/71 (36%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = -1
Query: 29 KLRLMPKPKHLLHLLSELRSKHLPLLSLNGQYVPTLRH-TLLHNVPRLGSHLLLLAKYSV 87
+L+L P L EL S L LL++ TLR + L + P + S+LLLL S+
Sbjct: 475 RLQLQPLSPSL-----ELASPSLLLLTITATATVTLRQLSPLDSAPIVPSYLLLLHSPSL 311
Query: 88 PLLVKLRCLLL 98
P + + RCL L
Sbjct: 310 PDVKEDRCLKL 278
>AV781208
Length = 524
Score = 29.3 bits (64), Expect = 7.2
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = +1
Query: 1366 QSDQDGARRQREDVFHHTLGHFLLQSNAVRFDQ 1398
QSD + Q+ D FHH+L F LQ +RF Q
Sbjct: 184 QSDHPALQFQKHDDFHHSLTDF-LQHTVLRFQQ 279
>TC11724 weakly similar to UP|Q9M1Q2 (Q9M1Q2) Serine/threonine protein
kinase-like protein, partial (20%)
Length = 535
Score = 29.3 bits (64), Expect = 7.2
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = -1
Query: 1366 QSDQDGARRQREDVFHHTLGHFLLQSNAVRFDQ 1398
QSD + Q+ D FHH+L F LQ +RF Q
Sbjct: 367 QSDHPALQFQKHDDFHHSLTDF-LQHTVLRFQQ 272
>TC10513 similar to GB|AAN15531.1|23198008|BT000212 expressed protein
{Arabidopsis thaliana;}, partial (40%)
Length = 647
Score = 28.9 bits (63), Expect = 9.5
Identities = 17/33 (51%), Positives = 20/33 (60%)
Frame = -2
Query: 651 VQYLILLRRQFLTSTMTL**KVVVKFQYHLCLL 683
VQ++ILLR Q LTS TL K +H CLL
Sbjct: 226 VQFVILLRSQSLTSRKTL------KSYHHACLL 146
>TC13523 homologue to UP|R1AD_ARATH (Q9FKC0) 60S ribosomal protein L13a-4,
partial (52%)
Length = 351
Score = 28.9 bits (63), Expect = 9.5
Identities = 13/48 (27%), Positives = 26/48 (54%)
Frame = -3
Query: 33 MPKPKHLLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHLL 80
+P+ H+LHLL + ++H LL+ N + + L H+ + H++
Sbjct: 220 LPEKPHVLHLLPDETTRHANLLTPNHHHFLAVEKLLRHDRSKPTQHVV 77
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,660,012
Number of Sequences: 28460
Number of extensions: 663679
Number of successful extensions: 7850
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 5631
Number of HSP's successfully gapped in prelim test: 187
Number of HSP's that attempted gapping in prelim test: 1880
Number of HSP's gapped (non-prelim): 6290
length of query: 2301
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2196
effective length of database: 1,909,300
effective search space: 4192822800
effective search space used: 4192822800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144656.5


