

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144645.10 - phase: 0 /pseudo
(70 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
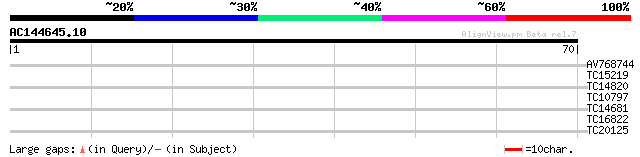
Score E
Sequences producing significant alignments: (bits) Value
AV768744 29 0.19
TC15219 similar to PIR|T00986|T00986 yeast pheromone receptor-li... 25 2.8
TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate p... 24 4.8
TC10797 weakly similar to PIR|C86287|C86287 F9L1.24 protein - Ar... 24 4.8
TC14681 similar to UP|Q9SLZ4 (Q9SLZ4) Retinoblastoma-related pro... 24 6.2
TC16822 23 8.2
TC20125 23 8.2
>AV768744
Length = 526
Score = 28.9 bits (63), Expect = 0.19
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 9/60 (15%)
Frame = +2
Query: 9 GASVHVEASKYVDSSVL-----PLQAHEVDTARF----FQTTLSRKIEKNCLNGHVVKQI 59
G+++HV A++ V+SSVL P VD R F + +++ +N H + QI
Sbjct: 317 GSTIHVVANQLVESSVLVF*MEPAARSTVDAPRVARIGFVAVIVQRVSAEVVNAHALLQI 496
>TC15219 similar to PIR|T00986|T00986 yeast pheromone receptor-like protein
AR781 [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (11%)
Length = 571
Score = 25.0 bits (53), Expect = 2.8
Identities = 17/41 (41%), Positives = 21/41 (50%), Gaps = 6/41 (14%)
Frame = +3
Query: 1 VEAKPNTDGASVHVEA-----SKYVD-SSVLPLQAHEVDTA 35
VEA P T S H+ S V S+ +PLQAH+ TA
Sbjct: 186 VEAPPGTISISCHISVPLLHPSVLVKISTSVPLQAHQASTA 308
>TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate protein 80,
partial (53%)
Length = 1180
Score = 24.3 bits (51), Expect = 4.8
Identities = 13/34 (38%), Positives = 16/34 (46%)
Frame = +2
Query: 17 SKYVDSSVLPLQAHEVDTARFFQTTLSRKIEKNC 50
S +V+SSV PL +R QT S NC
Sbjct: 167 SSFVNSSVTPLMLWTRSVSRVSQTRASSMDNPNC 268
>TC10797 weakly similar to PIR|C86287|C86287 F9L1.24 protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (10%)
Length = 657
Score = 24.3 bits (51), Expect = 4.8
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = -1
Query: 19 YVDSSVLPLQAHEVDTARFFQTTLSRKIEK 48
Y+ PL A ++DT F + RKI+K
Sbjct: 291 YIQGPPPPLVARKIDTDPFLGFSYQRKIQK 202
>TC14681 similar to UP|Q9SLZ4 (Q9SLZ4) Retinoblastoma-related protein,
partial (22%)
Length = 911
Score = 23.9 bits (50), Expect = 6.2
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = -3
Query: 35 ARFFQTTLSRKIEKNCLN 52
+R TTL + I K+CLN
Sbjct: 735 SRIIYTTLQKNIHKHCLN 682
>TC16822
Length = 802
Score = 23.5 bits (49), Expect = 8.2
Identities = 7/24 (29%), Positives = 16/24 (66%)
Frame = -2
Query: 45 KIEKNCLNGHVVKQIRWDLQLLHK 68
K+E+ C++ H + + ++ +LHK
Sbjct: 681 KVEQGCISDHPGQVLTFNFDILHK 610
>TC20125
Length = 401
Score = 23.5 bits (49), Expect = 8.2
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = -1
Query: 42 LSRKIEKNCLNGHVVKQIRWDLQLLH 67
L + +EKN HV+ + W LQ+LH
Sbjct: 380 LRKCLEKN----HVILMVVWFLQILH 315
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,168,968
Number of Sequences: 28460
Number of extensions: 10989
Number of successful extensions: 38
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 38
length of query: 70
length of database: 4,897,600
effective HSP length: 46
effective length of query: 24
effective length of database: 3,588,440
effective search space: 86122560
effective search space used: 86122560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 48 (23.1 bits)
Medicago: description of AC144645.10


