

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.6 - phase: 0
(178 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
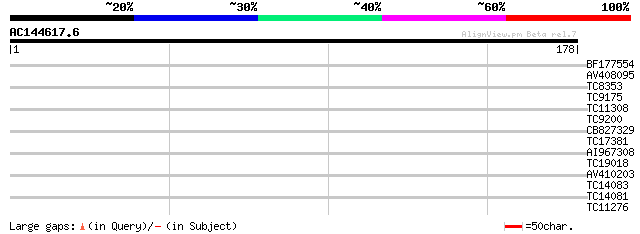
Score E
Sequences producing significant alignments: (bits) Value
BF177554 29 0.41
AV408095 28 1.2
TC8353 homologue to UP|HS7M_PHAVU (Q01899) Heat shock 70 kDa pro... 27 2.0
TC9175 weakly similar to UP|INAD_DROME (Q24008) Inactivation-no-... 27 2.6
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 26 3.4
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 26 4.5
CB827329 25 5.9
TC17381 similar to UP|AAQ93070 (AAQ93070) 3-ketoacyl-CoA thiolas... 25 5.9
AI967308 25 5.9
TC19018 similar to UP|CGS5_YEAST (P30283) S-phase entry cyclin 5... 25 5.9
AV410203 25 5.9
TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor... 25 7.7
TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate c... 25 7.7
TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacety... 25 7.7
>BF177554
Length = 246
Score = 29.3 bits (64), Expect = 0.41
Identities = 15/33 (45%), Positives = 21/33 (63%), Gaps = 1/33 (3%)
Frame = +1
Query: 135 SIEEEAEKEEGTSAKQESDGE-DSSAEKKEDAE 166
S EEE E+EEG ++E DG D A+ +E+ E
Sbjct: 61 SEEEEEEEEEGEEEEEEDDGPGDRDAQPEEEEE 159
>AV408095
Length = 318
Score = 27.7 bits (60), Expect = 1.2
Identities = 16/35 (45%), Positives = 21/35 (59%)
Frame = -1
Query: 26 LNFIDGIWSASTGERLIIFTTNYAEKLDHALICRG 60
LNF G S STG +I+ T ++A +L AL C G
Sbjct: 195 LNF--GGSSGSTGISIILITLSFAPELKAALCCEG 97
>TC8353 homologue to UP|HS7M_PHAVU (Q01899) Heat shock 70 kDa protein,
mitochondrial precursor, partial (74%)
Length = 1752
Score = 26.9 bits (58), Expect = 2.0
Identities = 17/59 (28%), Positives = 29/59 (48%)
Frame = +1
Query: 33 WSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKMLATKYLSLESHFL 91
W+ S +R + F L +L+C+G + +L +L C K+ T LSL +H +
Sbjct: 334 WTNSREQRALTF-------LRISLLCKGFVKLLRKLR*NCLQHLKLRLTCLLSLPTHLV 489
>TC9175 weakly similar to UP|INAD_DROME (Q24008)
Inactivation-no-after-potential D protein, partial (5%)
Length = 692
Score = 26.6 bits (57), Expect = 2.6
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 138 EEAEKEEGTSAKQESDGEDSSAEKKEDAE 166
EEA+KEEG K ++ E E KE+ +
Sbjct: 161 EEAKKEEGGEEKNDNKEEKGGEEGKEEGK 247
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 26.2 bits (56), Expect = 3.4
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -2
Query: 137 EEEAEKEEGTSAKQESDGED 156
EEE E EEG ++E D ED
Sbjct: 299 EEEVEAEEGEEDEEEEDDED 240
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 25.8 bits (55), Expect = 4.5
Identities = 12/31 (38%), Positives = 21/31 (67%), Gaps = 3/31 (9%)
Frame = +1
Query: 137 EEEAEKEEGTSAKQESDGE---DSSAEKKED 164
EEE E+EEG ++E + E ++ AE+++D
Sbjct: 244 EEEEEEEEGEEEEEEEEEEEEVEAEAEEEDD 336
>CB827329
Length = 423
Score = 25.4 bits (54), Expect = 5.9
Identities = 14/40 (35%), Positives = 22/40 (55%)
Frame = -2
Query: 126 LLRLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDA 165
+L L+ RS EEE E E+ S + DG+ + K+ +A
Sbjct: 251 VL*LLLLSRSKEEEQETEDSLSLRG*EDGDSGNEFKQREA 132
>TC17381 similar to UP|AAQ93070 (AAQ93070) 3-ketoacyl-CoA thiolase ,
partial (39%)
Length = 547
Score = 25.4 bits (54), Expect = 5.9
Identities = 9/17 (52%), Positives = 14/17 (81%)
Frame = -2
Query: 54 HALICRGRMDMLIELPY 70
HAL+CR + DML++ P+
Sbjct: 228 HALLCRWQYDMLLQ*PH 178
>AI967308
Length = 443
Score = 25.4 bits (54), Expect = 5.9
Identities = 13/39 (33%), Positives = 21/39 (53%)
Frame = +1
Query: 128 RLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDAE 166
+L +A+ + E KEEG + + GE+ A K +AE
Sbjct: 304 QLNEAVSKVSEAETKEEGMKLQLKELGEELEAASKLNAE 420
>TC19018 similar to UP|CGS5_YEAST (P30283) S-phase entry cyclin 5, partial
(4%)
Length = 828
Score = 25.4 bits (54), Expect = 5.9
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = -1
Query: 89 HFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPL 126
HFL DK+ + T++ P + NL + N V TPL
Sbjct: 711 HFLADKL**ACI*TSLHPNRLKGNLYKALSNHGV*TPL 598
>AV410203
Length = 339
Score = 25.4 bits (54), Expect = 5.9
Identities = 10/32 (31%), Positives = 19/32 (59%)
Frame = +1
Query: 55 ALICRGRMDMLIELPYCCFDGFKMLATKYLSL 86
A+ C + +L+E P+CC D + L+ L++
Sbjct: 130 AM*CIHVLSLLMEFPHCCLDEYLKLSLISLAI 225
>TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor, partial
(92%)
Length = 1693
Score = 25.0 bits (53), Expect = 7.7
Identities = 25/96 (26%), Positives = 42/96 (43%), Gaps = 6/96 (6%)
Frame = +1
Query: 82 KYLSLE-----SHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALRSI 136
KY+ +E S LFD + L+ + VAE K + + A +A +
Sbjct: 1054 KYVGIELWQVKSGTLFDNV--LITDDPEYAKQVAEETWGKQKDAEKAA----FEEAEKKR 1215
Query: 137 EEEAEKEEGTSAKQESDGE-DSSAEKKEDAEMVTSS 171
EEE KE+ + E + + D ++ +DAE T +
Sbjct: 1216 EEEESKEDPADSDAEDEEDTDEASHDSDDAESKTEA 1323
>TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate
carboxylase/oxygenase activase, chloroplast precursor
(RuBisCO activase) (RA), partial (98%)
Length = 1625
Score = 25.0 bits (53), Expect = 7.7
Identities = 10/39 (25%), Positives = 19/39 (48%)
Frame = -2
Query: 70 YCCFDGFKMLATKYLSLESHFLFDKIACLLVETNMTPAD 108
+ C F++ +T + ++SH L DK+ + P D
Sbjct: 664 FSCISTFQLSSTHHDGVDSHLLEDKLTLERFSLSFAPPD 548
>TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacetylase 1
(HD1), partial (5%)
Length = 636
Score = 25.0 bits (53), Expect = 7.7
Identities = 11/42 (26%), Positives = 23/42 (54%)
Frame = +3
Query: 137 EEEAEKEEGTSAKQESDGEDSSAEKKEDAEMVTSSMLRLESY 178
+EEA+KEE +++ D + +KKE + + +++Y
Sbjct: 111 KEEAKKEEPKKEEEKKDEKKEEEKKKEQPAQIDPVLEWVKAY 236
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.130 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,440,453
Number of Sequences: 28460
Number of extensions: 30271
Number of successful extensions: 135
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 132
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 135
length of query: 178
length of database: 4,897,600
effective HSP length: 84
effective length of query: 94
effective length of database: 2,506,960
effective search space: 235654240
effective search space used: 235654240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144617.6


