

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.10 + phase: 0
(337 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
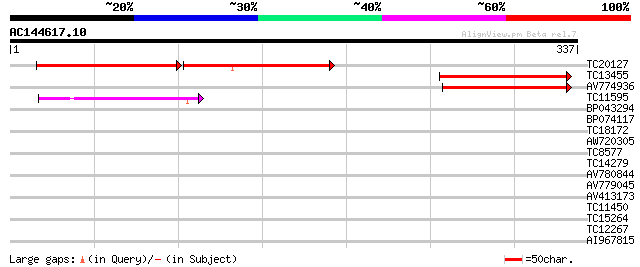
Score E
Sequences producing significant alignments: (bits) Value
TC20127 107 6e-45
TC13455 weakly similar to UP|Q9LG41 (Q9LG41) Dimethylaniline mon... 70 5e-13
AV774936 64 3e-11
TC11595 weakly similar to UP|Q9FKE7 (Q9FKE7) Dimethylaniline mon... 51 2e-07
BP043294 37 0.006
BP074117 35 0.023
TC18172 similar to UP|O81815 (O81815) Monooxygenase, partial (15%) 33 0.052
AW720305 31 0.26
TC8577 similar to UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase,... 30 0.57
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 29 0.98
AV780844 28 2.2
AV779045 28 2.2
AV413173 28 2.2
TC11450 similar to UP|Q9T0I6 (Q9T0I6) Splicing factor-like prote... 28 2.2
TC15264 homologue to UP|DHSA_ARATH (O82663) Succinate dehydrogen... 27 4.9
TC12267 27 6.4
AI967815 26 8.3
>TC20127
Length = 582
Score = 107 bits (267), Expect(2) = 6e-45
Identities = 48/86 (55%), Positives = 61/86 (70%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
IIVG G SGIA A+CL+++ + ++LER DC ASLWQ TYDRL LHL K CELP F
Sbjct: 39 IIVGGGTSGIATASCLTKKSISYIMLEREDCFASLWQKYTYDRLHLHLRKQSCELPHFPF 218
Query: 77 PQTFPKYPTKHQFISYMESYADHFHI 102
P ++P Y K QFI Y+++Y HF+I
Sbjct: 219 PPSYPHYVPKKQFIEYLDNYVKHFNI 296
Score = 90.1 bits (222), Expect(2) = 6e-45
Identities = 43/94 (45%), Positives = 62/94 (65%), Gaps = 4/94 (4%)
Frame = +1
Query: 104 PRFNQTVLSAEFDSTSQIWMVRTKEGDF----QYFSPWLIVATGENAEPVFPTIHGMEHF 159
P +++ V AE D++ Q W V+ K +Y +L+VATGE AEP P + G+E F
Sbjct: 301 PLYHRAVELAEHDNSHQNWRVKAKNRTSGHVEEYAGKFLVVATGETAEPRIPEVEGLEGF 480
Query: 160 HGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVS 193
G V+H++ YK+G E+KN+ VLV+G GNSGME+S
Sbjct: 481 KGKVIHSTGYKNGKEFKNQNVLVVGSGNSGMEIS 582
>TC13455 weakly similar to UP|Q9LG41 (Q9LG41) Dimethylaniline
monooxygenase-like protein, partial (9%)
Length = 535
Score = 70.1 bits (170), Expect = 5e-13
Identities = 36/80 (45%), Positives = 49/80 (61%), Gaps = 1/80 (1%)
Frame = +1
Query: 256 HYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFL-DGQEK 314
HYGI RP GP +K+ GK PV+D G I +IKSG++KV+ E R+ L +G+
Sbjct: 1 HYGIARPNEGPFCMKVKYGKYPVIDTGTIHKIKSGDLKVLPSEIEYVRDKNVLLKNGELH 180
Query: 315 EFDAIILATGYKSNVPSWLK 334
FD+II TG+K + WLK
Sbjct: 181 PFDSIIFCTGFKRSTHKWLK 240
>AV774936
Length = 477
Score = 64.3 bits (155), Expect = 3e-11
Identities = 32/78 (41%), Positives = 47/78 (60%), Gaps = 1/78 (1%)
Frame = -3
Query: 258 GIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEG-VKEITRNGAKFLDGQEKEF 316
GI P GP L++ G+ PV+DVG + QI+ G I+V+ ++ I N F DGQ + F
Sbjct: 475 GIPVPPEGPFPLQMKYGQFPVIDVGTVNQIQFGEIQVLPAEIESIRGNPMLFRDGQSQPF 296
Query: 317 DAIILATGYKSNVPSWLK 334
D+II TG++ + WLK
Sbjct: 295 DSIIFCTGFQRSTKKWLK 242
>TC11595 weakly similar to UP|Q9FKE7 (Q9FKE7) Dimethylaniline monooxygenase
(N-oxide-forming)-like protein, partial (13%)
Length = 435
Score = 51.2 bits (121), Expect = 2e-07
Identities = 32/101 (31%), Positives = 52/101 (50%), Gaps = 3/101 (2%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSFP 77
I+GAG SGIA A LS ++ E +D I +W++ Y+ KL E +P
Sbjct: 110 IIGAGVSGIAAAKQLSHHN--PIVFEATDSIGGIWRHCVYNCTKLQSQTWNYEFSDFPWP 283
Query: 78 QT-FPKYPTKHQFISYMESYADHFHIHP--RFNQTVLSAEF 115
+ YP+ + + Y+E+YA+ F ++ +FN VL +F
Sbjct: 284 KRESTDYPSYLEILEYLENYAERFDLNKLVKFNTKVLEVKF 406
>BP043294
Length = 527
Score = 36.6 bits (83), Expect = 0.006
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = -3
Query: 298 VKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
++ I N F DG+ + FD+II TG+K + WLK
Sbjct: 423 IESIRGNQVLFRDGKSQPFDSIIFCTGFKRSTKKWLK 313
>BP074117
Length = 395
Score = 34.7 bits (78), Expect = 0.023
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = -3
Query: 298 VKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
++ + N F DG+ FD+II TG+K + WLK
Sbjct: 387 IESVRGNQVLFGDGKSHTFDSIIFCTGFKRSTQKWLK 277
>TC18172 similar to UP|O81815 (O81815) Monooxygenase, partial (15%)
Length = 597
Score = 33.5 bits (75), Expect = 0.052
Identities = 15/30 (50%), Positives = 20/30 (66%)
Frame = +2
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSD 46
+IVG G G+A A L + + SL+LERSD
Sbjct: 95 VIVGGGICGLATALALHRKSIKSLVLERSD 184
>AW720305
Length = 577
Score = 31.2 bits (69), Expect = 0.26
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = +3
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASL 51
+VG+GP+G+A A L++ G + ER+D I L
Sbjct: 312 VVGSGPAGLAAADQLNKMGHTVTVFERADRIGGL 413
>TC8577 similar to UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, partial
(53%)
Length = 790
Score = 30.0 bits (66), Expect = 0.57
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILER 44
+VG GP+G A A LS+ G+ + ++ER
Sbjct: 221 VVGGGPAGGAAAETLSKAGIETFLIER 301
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 29.3 bits (64), Expect = 0.98
Identities = 14/43 (32%), Positives = 19/43 (43%)
Frame = +2
Query: 63 HLPKHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPR 105
HL H LP P T P H + ++ + +H H HPR
Sbjct: 95 HLHHHLPLLP----PHTTTNLPHHHHILHHLPTTTNHLHHHPR 211
>AV780844
Length = 563
Score = 28.1 bits (61), Expect = 2.2
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = -3
Query: 231 YKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPK 263
Y+W ++L F LLV+ +L TNH+ ++ +
Sbjct: 111 YQWSIIQLTISFCLLVNIMYLNVTNHFCMRNER 13
>AV779045
Length = 522
Score = 28.1 bits (61), Expect = 2.2
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = -1
Query: 231 YKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPK 263
Y+W ++L F LLV+ +L TNH+ ++ +
Sbjct: 156 YQWSIIQLTISFCLLVNIMYLNVTNHFCMRNER 58
>AV413173
Length = 404
Score = 28.1 bits (61), Expect = 2.2
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +1
Query: 168 DYKSGSEYKNKKVLVIGCGNSGMEVS 193
D+ G + K+V +IG GN GMEV+
Sbjct: 256 DFPLGCKLSGKRVGIIGLGNIGMEVA 333
>TC11450 similar to UP|Q9T0I6 (Q9T0I6) Splicing factor-like protein, partial
(14%)
Length = 1097
Score = 28.1 bits (61), Expect = 2.2
Identities = 20/70 (28%), Positives = 33/70 (46%)
Frame = +3
Query: 199 HNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLGNTNHYG 258
HN ++ A+N+V + +DM ++YG+ GL +V + L++ LG T
Sbjct: 147 HNIADYVTAKNNVVLSYKDMSHTNSYGLIRGLQ--FASFVVQYYGLVLDLLLLGLTRASE 320
Query: 259 IKRPKTGPIE 268
I P P E
Sbjct: 321 IAGPPQMPNE 350
>TC15264 homologue to UP|DHSA_ARATH (O82663) Succinate dehydrogenase
[ubiquinone] flavoprotein subunit, mitochondrial (FP)
(Flavoprotein subunit of complex II) , partial (77%)
Length = 1643
Score = 26.9 bits (58), Expect = 4.9
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +1
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTY 57
I+VGAG +G+ A LSE G + CI L+ R++
Sbjct: 232 IVVGAGGAGLRAAIGLSEHGF------NTACITKLFPTRSH 336
>TC12267
Length = 364
Score = 26.6 bits (57), Expect = 6.4
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = -1
Query: 95 SYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTK 127
+++ F +P FN + S++ STS I ++R K
Sbjct: 253 TFSSGFSCNPNFNPSFKSSKLSSTSDIELLRVK 155
>AI967815
Length = 343
Score = 26.2 bits (56), Expect = 8.3
Identities = 10/33 (30%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 119 SQIWMVRTKE-GDFQYFSPWLIVATGENAEPVF 150
+++W++R K F SPW ++ T ++ P F
Sbjct: 142 ARVWLIRHKPLACFPLLSPWPVLHTPHHSHPAF 44
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,812,313
Number of Sequences: 28460
Number of extensions: 107561
Number of successful extensions: 537
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 530
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 534
length of query: 337
length of database: 4,897,600
effective HSP length: 91
effective length of query: 246
effective length of database: 2,307,740
effective search space: 567704040
effective search space used: 567704040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144617.10


