

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144609.1 + phase: 0
(269 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
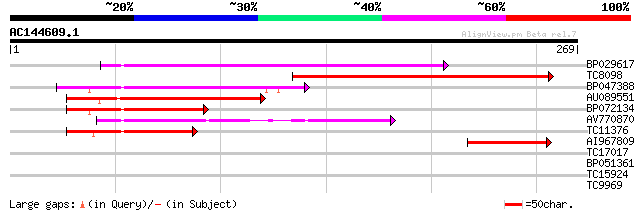
Sequences producing significant alignments: (bits) Value
BP029617 142 5e-35
TC8098 similar to UP|Q94FT5 (Q94FT5) Pectate lyase (Fragment), p... 140 3e-34
BP047388 96 5e-21
AU089551 77 4e-15
BP072134 72 1e-13
AV770870 66 7e-12
TC11376 similar to GB|AAM61400.1|21537059|AY084835 pectate lyase... 64 3e-11
AI967809 57 4e-09
TC17017 similar to UP|PEL8_ARATH (Q9M8Z8) Probable pectate lyase... 37 0.004
BP051361 29 0.74
TC15924 weakly similar to GB|BAC21577.1|24060129|AP005438 G-prot... 27 3.6
TC9969 similar to UP|Q93XJ1 (Q93XJ1) Pectate lyase, partial (22%) 27 3.6
>BP029617
Length = 493
Score = 142 bits (359), Expect = 5e-35
Identities = 69/165 (41%), Positives = 99/165 (59%)
Frame = -1
Query: 44 TLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTIDGRGVDVHIADNACLMIYKATNI 103
TLR+ + + +WI F +M I L L+ +S+ T+DGRG +V I + C+ + +N+
Sbjct: 493 TLRHAVIQDE-PLWIVFAADMTINLRHELIFNSYKTVDGRGANVQITGHGCITLQYISNV 317
Query: 104 IIHGIRVHHCRPQAPGMVMGPDGKIISLGQVDGDAIRLVSASKIWIDHNTLYDCQDGLLD 163
IIH I VHHC+P + + G DGD I + A IWIDH +L C DGL+D
Sbjct: 316 IIHNIHVHHCKPSGNTNIRASPTHVGWRGVSDGDGISIFGARNIWIDHCSLSYCADGLID 137
Query: 164 VTRGSTDITISNNWFREQNKVMLLGHDDGFVRDKNMKVTVVYNYF 208
GST ITISNN F ++VMLLGH+D + D+ M+VT+ +N+F
Sbjct: 136 AIMGSTAITISNNHFAHHDEVMLLGHNDKYTPDRGMQVTIAFNHF 2
>TC8098 similar to UP|Q94FT5 (Q94FT5) Pectate lyase (Fragment), partial
(56%)
Length = 1349
Score = 140 bits (352), Expect = 3e-34
Identities = 67/124 (54%), Positives = 80/124 (64%)
Frame = +3
Query: 135 DGDAIRLVSASKIWIDHNTLYDCQDGLLDVTRGSTDITISNNWFREQNKVMLLGHDDGFV 194
DGDAI + +S IW DHN+L DGL D GST ITISNN N+ +LLGH D +
Sbjct: 9 DGDAISIFGSSHIWDDHNSLPHRADGLADAVMGSTAITISNNHLTHHNEAILLGHSDSYT 188
Query: 195 RDKNMKVTVVYNYFGPNCHQRMPRIRHGYAHVANNMYMGWVQYAIGGSMEPSLKSQSNLF 254
RDK M+VT+ YN+FG QRMPR HGY HV NN Y W YAIGGS P++ SQ N +
Sbjct: 189 RDKQMQVTIAYNHFGEGLIQRMPRCIHGYFHVVNNDYTHWEMYAIGGSGNPTINSQGNRY 368
Query: 255 IAPV 258
AP+
Sbjct: 369 NAPL 380
>BP047388
Length = 590
Score = 96.3 bits (238), Expect = 5e-21
Identities = 51/124 (41%), Positives = 72/124 (57%), Gaps = 4/124 (3%)
Frame = -2
Query: 23 KDLIHYKVTDPSDH-PLNSTPGTLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTID 81
K + Y VTD SDH P+N PGTLRY + + +WI F NM IKL + L+ +S+ TID
Sbjct: 580 KGVQFYIVTDSSDHDPVNPKPGTLRYAVIQNE-PLWIVFPSNMMIKLSQELIFNSYKTID 404
Query: 82 GRGVDVHIADNACLMIYKATNIIIHGIRVHHCRPQAPGM-VMGPDG--KIISLGQVDGDA 138
GRG DVHI C+ + +N+IIH I VHHC P + GP+ ++ + Q+
Sbjct: 403 GRGADVHIVGGGCITLQYISNVIIHNIHVHHCHPSGTIIWACGPEDPMMVVLIFQISAQI 224
Query: 139 IRLV 142
R++
Sbjct: 223 TRMI 212
>AU089551
Length = 599
Score = 76.6 bits (187), Expect = 4e-15
Identities = 45/95 (47%), Positives = 59/95 (61%), Gaps = 1/95 (1%)
Frame = +3
Query: 28 YKVTDPSDHPLNST-PGTLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTIDGRGVD 86
Y VTD SD+ + S PGTLRY + +G +WI F R M I L + LLISS TIDGRG +
Sbjct: 300 YVVTDDSDNDMVSPKPGTLRYAVVQ-KGPLWIIFARXMVITLQQELLISSDKTIDGRGAN 476
Query: 87 VHIADNACLMIYKATNIIIHGIRVHHCRPQAPGMV 121
V I A L + N+I+HGIRV + + + GM+
Sbjct: 477 VQIRGGAGLTMQFVNNVIVHGIRVKNIKARNGGMI 581
>BP072134
Length = 397
Score = 71.6 bits (174), Expect = 1e-13
Identities = 38/68 (55%), Positives = 46/68 (66%), Gaps = 1/68 (1%)
Frame = +2
Query: 28 YKVTDPSDH-PLNSTPGTLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTIDGRGVD 86
Y VTD D P+N PGTLRYGA + + +WI FKR+M I L + LL++SF TIDGRG
Sbjct: 197 YVVTDSGDDDPVNPRPGTLRYGAIQDE-PLWIIFKRDMVINLKQELLVNSFKTIDGRGAS 373
Query: 87 VHIADNAC 94
VHIA C
Sbjct: 374 VHIAGGGC 397
>AV770870
Length = 528
Score = 65.9 bits (159), Expect = 7e-12
Identities = 49/142 (34%), Positives = 73/142 (50%)
Frame = +2
Query: 42 PGTLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTIDGRGVDVHIADNACLMIYKAT 101
PG+LR G + + +WI F+ + I L L +SS+ T+DGRG + + L + +
Sbjct: 158 PGSLREGCRRKE-PLWIVFQVSGTIHLQSYLSVSSYKTVDGRGQRIKLTGKG-LRLKECE 331
Query: 102 NIIIHGIRVHHCRPQAPGMVMGPDGKIISLGQVDGDAIRLVSASKIWIDHNTLYDCQDGL 161
+II+ + + R G D VDG I+ ++ IWID +L D DGL
Sbjct: 332 HIIVCNLELEGGR--------GHD--------VDGIQIK-PNSRHIWIDRCSLRDYDDGL 460
Query: 162 LDVTRGSTDITISNNWFREQNK 183
+D+TR STDITIS F +K
Sbjct: 461 IDITRQSTDITISRCHFASHDK 526
>TC11376 similar to GB|AAM61400.1|21537059|AY084835 pectate lyase-like
protein {Arabidopsis thaliana;} , partial (26%)
Length = 636
Score = 63.9 bits (154), Expect = 3e-11
Identities = 34/63 (53%), Positives = 43/63 (67%), Gaps = 1/63 (1%)
Frame = +1
Query: 28 YKVTDPSDHPL-NSTPGTLRYGASKIQGKVWITFKRNMNIKLVRPLLISSFTTIDGRGVD 86
Y VTD SD+ + N PGTLRYG + + +WI F NM IKL + L+ +S+ TIDGRG D
Sbjct: 451 YLVTDSSDNDVVNPKPGTLRYGVIQNE-PLWIVFPSNMMIKLSQELIFNSYKTIDGRGAD 627
Query: 87 VHI 89
VHI
Sbjct: 628 VHI 636
>AI967809
Length = 482
Score = 56.6 bits (135), Expect = 4e-09
Identities = 24/40 (60%), Positives = 28/40 (70%)
Frame = +2
Query: 218 RIRHGYAHVANNMYMGWVQYAIGGSMEPSLKSQSNLFIAP 257
R RHGY HV NN Y W YAIGGS P++ SQ+N F+AP
Sbjct: 65 RCRHGYFHVVNNDYTHWEMYAIGGSANPTINSQANRFLAP 184
>TC17017 similar to UP|PEL8_ARATH (Q9M8Z8) Probable pectate lyase 8
precursor , partial (28%)
Length = 670
Score = 37.0 bits (84), Expect = 0.004
Identities = 23/48 (47%), Positives = 30/48 (61%), Gaps = 2/48 (4%)
Frame = +3
Query: 28 YKVTDP-SDHPLNSTPGTLRYGASKIQGK-VWITFKRNMNIKLVRPLL 73
Y VTD D P+N PGTLR+ IQ + +WI FKR+M I L + L+
Sbjct: 531 YVVTDSRDDDPVNPRPGTLRHAV--IQDRPLWIVFKRDMVITLKQELI 668
>BP051361
Length = 458
Score = 29.3 bits (64), Expect = 0.74
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -2
Query: 146 KIWIDHNTLYDCQDGLLDVT 165
K WI+H T Y+C DG D T
Sbjct: 394 KPWIEHVTTYNCNDGPCDAT 335
>TC15924 weakly similar to GB|BAC21577.1|24060129|AP005438 G-protein beta
family-like protein {Oryza sativa (japonica
cultivar-group);}, partial (11%)
Length = 1262
Score = 26.9 bits (58), Expect = 3.6
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = -1
Query: 25 LIHYKVTDPSDHPLNSTP 42
LI + + DPS++PLNS+P
Sbjct: 164 LISHTLIDPSEYPLNSSP 111
>TC9969 similar to UP|Q93XJ1 (Q93XJ1) Pectate lyase, partial (22%)
Length = 485
Score = 26.9 bits (58), Expect = 3.6
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +1
Query: 239 IGGSMEPSLKSQSNLFIAP 257
IGGS P+L SQ + F+AP
Sbjct: 1 IGGSANPTLHSQGHSFVAP 57
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,976,186
Number of Sequences: 28460
Number of extensions: 66157
Number of successful extensions: 302
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 297
length of query: 269
length of database: 4,897,600
effective HSP length: 89
effective length of query: 180
effective length of database: 2,364,660
effective search space: 425638800
effective search space used: 425638800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144609.1


