

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144592.1 - phase: 0 /pseudo
(486 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
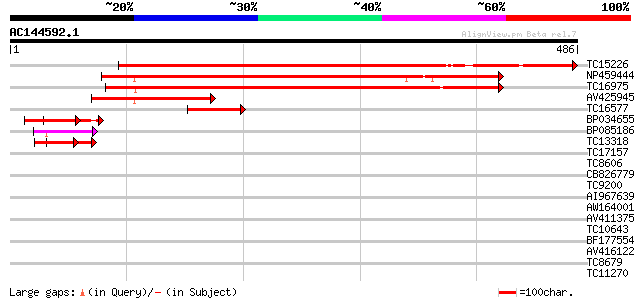
Score E
Sequences producing significant alignments: (bits) Value
TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Frag... 564 e-161
NP459444 rac GTPase activating protein 3 [Lotus japonicus] 377 e-105
TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, part... 376 e-105
AV425945 142 9e-35
TC16577 similar to UP|O82458 (O82458) Rac GTPase activating prot... 77 8e-15
BP034655 45 3e-05
BP085186 40 9e-04
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 40 9e-04
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 38 0.003
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 36 0.012
CB826779 36 0.016
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 35 0.021
AI967639 35 0.021
AW164001 35 0.021
AV411375 35 0.027
TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear phospho... 35 0.036
BF177554 34 0.047
AV416122 34 0.061
TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partia... 34 0.061
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 34 0.061
>TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Fragment),
partial (92%)
Length = 1436
Score = 564 bits (1453), Expect = e-161
Identities = 294/394 (74%), Positives = 324/394 (81%), Gaps = 1/394 (0%)
Frame = +3
Query: 94 KSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKV 153
KS+VTCSV+REDVSSLDISWPTEVRHVSHVTFDRFNGFLGLP+E QPEVP +VPSAS KV
Sbjct: 3 KSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKV 182
Query: 154 FGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNK 213
FGVSAKSMQCSYD+RGNSVPTILL MQ +LYSEGGLKAEGIFRI A+NSQE FVR QLN+
Sbjct: 183 FGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNR 362
Query: 214 GVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALL 273
G+VP G++VHCLSGLIKAWFRELPTGVLDSLTPEQVM CN+E+DCTNLVKLLPSTEAALL
Sbjct: 363 GLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALL 542
Query: 274 DWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKML 333
DWAINLMADVVE+EQFNKMNARN+AMVFAPNMTQM DPLTALIHAVQVMNFLKTLILK L
Sbjct: 543 DWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTL 722
Query: 334 REREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDACDKNSSTTKRKFSRTS 393
RER+ES+ AR LS ++ SC + PF N+EESS + + D C K +FS
Sbjct: 723 RERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPV-DTC-ATMPPDKSEFS--- 887
Query: 394 TLGRIEWSV-EKLRSSEEKRNREEVFRSFSSDSVTPRYESGPLENSYSYRRRYDSEHWSR 452
R+EW V EK+ SSEEK S S S RYESGPLE+ YR YDSEHW R
Sbjct: 888 ---RMEWCVDEKVWSSEEKGTGGGALESVSGGSSPSRYESGPLES--RYRGIYDSEHWLR 1052
Query: 453 LRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG 486
LR GVR+LC+HPVFQLSK +KK A LGIVNTREG
Sbjct: 1053LRKGVRRLCQHPVFQLSKSTKKRADLGIVNTREG 1154
>NP459444 rac GTPase activating protein 3 [Lotus japonicus]
Length = 1300
Score = 377 bits (967), Expect = e-105
Identities = 204/356 (57%), Positives = 254/356 (71%), Gaps = 11/356 (3%)
Frame = +2
Query: 79 QNQFAILDIVMAALKKSIVTCSVERED-----VSSLDISWPTEVRHVSHVTFDRFNGFLG 133
QNQ + ++AALKKS+V CSV+ D V ++I WPT V+HV+HVTFDRFNGFLG
Sbjct: 14 QNQGSPAAFLLAALKKSMVACSVDSPDDVISAVHPMEIGWPTNVKHVTHVTFDRFNGFLG 193
Query: 134 LPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEG 193
LP E + VP VPSASV VFGVSA+SM CSYD +GNSVPTILL MQ++LYS+GGL AEG
Sbjct: 194 LPLELEVHVPAPVPSASVSVFGVSAESMHCSYDSKGNSVPTILLLMQERLYSQGGLMAEG 373
Query: 194 IFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCN 253
IFRI EN QE +RDQLN+GVVP IDVHCL+GLIKAWFRELP+GVLD L+PEQV++CN
Sbjct: 374 IFRINPENGQEEHLRDQLNRGVVPDNIDVHCLAGLIKAWFRELPSGVLDGLSPEQVLECN 553
Query: 254 TEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLT 313
TE++ LVK L TE ALL+WA++LMADVVE E+ NKM+ARN+AMVFAPNMTQM+DPLT
Sbjct: 554 TEEEFVQLVKQLKPTELALLNWALDLMADVVEEEEHNKMDARNIAMVFAPNMTQMSDPLT 733
Query: 314 ALIHAVQVMNFLKTLILKMLREREE--SIDNARLLSHSMDFASCNDEFPP----FSFNKE 367
AL+HAVQVMN LKTLILK L EREE + + + SHS D S DE+ ++ +
Sbjct: 734 ALMHAVQVMNLLKTLILKTLSEREEATTAGYSSMSSHSSDRQS-EDEYDSQLEMYTSAEL 910
Query: 368 ESSCEQIEDACDKNSSTTKRKFSRTSTLGRIEWSVEKLRSSEEKRNREEVFRSFSS 423
S +D + NS ++ + S E +++L +++ EE RS S
Sbjct: 911 RGSQSDCDDHVNNNSLNSEEEVDSASVSEIEECFLKQLNENKQGGFAEEPARSTCS 1078
>TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, partial (88%)
Length = 1686
Score = 376 bits (965), Expect = e-105
Identities = 200/347 (57%), Positives = 248/347 (70%), Gaps = 6/347 (1%)
Frame = +1
Query: 83 AILDIVMAALKKSIV-TCSVEREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLGLPS 136
++L +++A +KS++ +CS D SS++I WP+ VRHV+HVTFDRF+GFLGLP
Sbjct: 4 SLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPV 183
Query: 137 EFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFR 196
EF+PEVP R PSAS VFGVS +SMQ S+D RGNSVPTILL MQ+ LY+ GGL+AEGIFR
Sbjct: 184 EFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFR 363
Query: 197 ITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTED 256
I AENSQE VR+QLN+GVVP+G+DVHCL+GLIKAWFRELPTG+LD L+PE+VMQ +E+
Sbjct: 364 INAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEE 543
Query: 257 DCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALI 316
+C LV+LLP TEAALLDWAINLMADV + E FNKMNARN+AMVFAPNMT MADPLTAL+
Sbjct: 544 ECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALM 723
Query: 317 HAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIED 376
+AVQVMNFLKTL++K LR REESI + + + F + K+ S E D
Sbjct: 724 YAVQVMNFLKTLVVKTLRVREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGS--ENGND 897
Query: 377 ACDKNSSTTKRKFSRTSTLGRIEWSVEKLRSSEEKRNREEVFRSFSS 423
D+++ + S+ S E E SE E F S S
Sbjct: 898 CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGS 1038
>AV425945
Length = 415
Score = 142 bits (359), Expect = 9e-35
Identities = 67/111 (60%), Positives = 87/111 (78%), Gaps = 5/111 (4%)
Frame = +1
Query: 71 TKGSNNNHQNQFAILDIVMAALKKSIVTCSVERED-----VSSLDISWPTEVRHVSHVTF 125
T + QNQ +++ +++AAL+KS+V C V+R D V ++I WPT+V+H++HVTF
Sbjct: 82 TAEEEQDRQNQLSLVALLLAALRKSMVACRVDRPDEAISTVHQMEIGWPTDVQHITHVTF 261
Query: 126 DRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTIL 176
DRFNGFLGLP EF+ E+P RVPSASV VFGVSA+SMQCSYD +GNSVPTIL
Sbjct: 262 DRFNGFLGLPVEFEVEIPGRVPSASVSVFGVSAESMQCSYDSKGNSVPTIL 414
>TC16577 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (16%)
Length = 726
Score = 76.6 bits (187), Expect = 8e-15
Identities = 35/50 (70%), Positives = 44/50 (88%)
Frame = +1
Query: 153 VFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENS 202
+FGVS +SMQ SYD RGNSVPTILL+MQ+ LY++GGL+AEGIFRI +N+
Sbjct: 172 MFGVSTESMQLSYDTRGNSVPTILLQMQRHLYAQGGLQAEGIFRINTDNT 321
>BP034655
Length = 517
Score = 45.1 bits (105), Expect = 3e-05
Identities = 19/48 (39%), Positives = 34/48 (70%)
Frame = +1
Query: 13 AGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
A F +S+ + + +++EE++E+EEE+EEEE E DD +++D+D I
Sbjct: 118 ADFGLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYI 261
Score = 44.7 bits (104), Expect = 3e-05
Identities = 22/51 (43%), Positives = 35/51 (68%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQN 80
+++EE++E+EEE+EEEE E +++D +D+DE Y S + S N HQN
Sbjct: 163 EEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRV---SPNVHQN 306
Score = 31.2 bits (69), Expect = 0.40
Identities = 15/56 (26%), Positives = 30/56 (52%)
Frame = +1
Query: 4 LFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
LF + + + F +E A + + E+EEE+EEEE E ++ + ++++E
Sbjct: 46 LFATPMASPSEFHRNEIAAAHLTAADFGLSDDDEEEEEEEEEEEEEEEEEEEEEEE 213
>BP085186
Length = 388
Score = 40.0 bits (92), Expect = 9e-04
Identities = 20/64 (31%), Positives = 35/64 (54%), Gaps = 9/64 (14%)
Frame = -1
Query: 21 NPAAQVTNNK---------KKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFT 71
NP A++ N K + E++E+EEE+EEEE E ++ + + ++E N + Q
Sbjct: 325 NPYARIVNGKLISPDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQEG 146
Query: 72 KGSN 75
+G N
Sbjct: 145 EGEN 134
Score = 34.3 bits (77), Expect = 0.047
Identities = 13/49 (26%), Positives = 30/49 (60%)
Frame = -1
Query: 12 IAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+ G +S N + +++EE++E+EEE+EEEE + + ++++ +
Sbjct: 307 VNGKLISPDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENV 161
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 40.0 bits (92), Expect = 9e-04
Identities = 17/43 (39%), Positives = 29/43 (66%)
Frame = +3
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGS 74
+EE+ + EEED+E+E + DD+D+DDD E ++ + +GS
Sbjct: 252 EEEEDDDEEEDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEGS 380
Score = 40.0 bits (92), Expect = 9e-04
Identities = 15/38 (39%), Positives = 27/38 (70%)
Frame = +3
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
P V + ++EE ++EE+DE+E+ + DD+D+DD +E
Sbjct: 225 PPKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEE 338
Score = 37.0 bits (84), Expect = 0.007
Identities = 12/38 (31%), Positives = 29/38 (75%)
Frame = +3
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
P T+N+++E+ E+E++++E++ + DD+D+D ++E
Sbjct: 228 PKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEEE 341
Score = 26.9 bits (58), Expect = 7.5
Identities = 12/48 (25%), Positives = 26/48 (54%), Gaps = 13/48 (27%)
Frame = +3
Query: 25 QVTNNKKKEEQQEQEEEDE-------------EEELESDDNDNDDDDE 59
Q N +++++ +E+++EDE EEE + DD++ ++ E
Sbjct: 240 QTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEGSE 383
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 38.1 bits (87), Expect = 0.003
Identities = 23/69 (33%), Positives = 36/69 (51%), Gaps = 5/69 (7%)
Frame = +2
Query: 16 SVSEFNPAAQVTNNKKKEEQQEQEEEDEEE-----ELESDDNDNDDDDEIYSSNNASRQF 70
+VS F A+ ++ + EE E +EDE+E E E DD+D+D+DDE + S+
Sbjct: 722 AVSWFTGEAEQSDFEDIEEDDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSG 901
Query: 71 TKGSNNNHQ 79
+K Q
Sbjct: 902 SKARPGKDQ 928
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 36.2 bits (82), Expect = 0.012
Identities = 14/27 (51%), Positives = 21/27 (76%)
Frame = -2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
EE E EE+++ E+ E DD+D++DDDE
Sbjct: 443 EENGEDEEDEDGEDQEDDDDDDEDDDE 363
Score = 35.4 bits (80), Expect = 0.021
Identities = 13/50 (26%), Positives = 29/50 (58%)
Frame = -2
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQ 81
+E +++E+ED E++ + DD+D DDD+E + + + +N + +
Sbjct: 443 EENGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEEDNEDEEE 294
Score = 31.6 bits (70), Expect = 0.30
Identities = 10/32 (31%), Positives = 25/32 (77%)
Frame = -2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++ +E+ + E++EEE +E +DN+++++DE
Sbjct: 383 DDEDDDEEDDGGEDEEEEGVEEEDNEDEEEDE 288
Score = 28.5 bits (62), Expect = 2.6
Identities = 10/32 (31%), Positives = 23/32 (71%)
Frame = -2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++ ++++ E EEE+ EE +++D + D++DE
Sbjct: 374 DDDEEDDGGEDEEEEGVEEEDNEDEEEDEEDE 279
Score = 26.9 bits (58), Expect = 7.5
Identities = 9/32 (28%), Positives = 23/32 (71%)
Frame = -2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ + +++EEE EEE D+ ++++D+E
Sbjct: 371 DDEEDDGGEDEEEEGVEEEDNEDEEEDEEDEE 276
Score = 26.6 bits (57), Expect = 9.8
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = -2
Query: 27 TNNKKKEEQQEQEEEDEEEELESDDNDNDDDD 58
T + + Q E ++ D E E +DD ++DDD
Sbjct: 671 TTKGETKSQSEDKQHDAEGEDGNDDEGDEDDD 576
>CB826779
Length = 469
Score = 35.8 bits (81), Expect = 0.016
Identities = 17/65 (26%), Positives = 39/65 (59%), Gaps = 4/65 (6%)
Frame = +1
Query: 27 TNNKKKEEQQEQEE----EDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQF 82
T++KKK + QE ++ ED +E + D++D++DD++ + + +K S + +++
Sbjct: 151 TDDKKKPQPQEYDDAPVVEDVHDEADEDEDDDEDDEDDEDGAQGTTEGSKQSRSEKKSRK 330
Query: 83 AILDI 87
A+L +
Sbjct: 331 AMLKL 345
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 35.4 bits (80), Expect = 0.021
Identities = 13/43 (30%), Positives = 29/43 (67%)
Frame = +1
Query: 18 SEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+E +V ++E ++E+EEE+EEEE E ++ + ++++E+
Sbjct: 181 AEVQEYEEVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEV 309
Score = 34.3 bits (77), Expect = 0.047
Identities = 12/28 (42%), Positives = 24/28 (84%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDD 57
+++E ++E+EEE+EEEE+E++ + DD+
Sbjct: 256 EEEEGEEEEEEEEEEEEVEAEAEEEDDE 339
Score = 33.9 bits (76), Expect = 0.061
Identities = 14/30 (46%), Positives = 23/30 (76%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE+ E+EEE+EEEE E + ++DDE
Sbjct: 250 EEEEEEGEEEEEEEEEEEEVEAEAEEEDDE 339
Score = 32.0 bits (71), Expect = 0.23
Identities = 13/31 (41%), Positives = 23/31 (73%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+++EE +E+EEE+EEEE + + +DD+ I
Sbjct: 253 EEEEEGEEEEEEEEEEEEVEAEAEEEDDEPI 345
Score = 30.8 bits (68), Expect = 0.52
Identities = 12/41 (29%), Positives = 26/41 (63%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
E + +++EE++ +EEE+EEEE E + + +++D+
Sbjct: 214 EVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDD 336
>AI967639
Length = 335
Score = 35.4 bits (80), Expect = 0.021
Identities = 11/34 (32%), Positives = 26/34 (76%)
Frame = +1
Query: 26 VTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ + +E+ E+E++ +E LE ++ D+DDDD+
Sbjct: 82 ISSSSEGDEEDEEEDQGDEYNLEDEEEDDDDDDD 183
Score = 33.1 bits (74), Expect = 0.10
Identities = 15/45 (33%), Positives = 30/45 (66%), Gaps = 3/45 (6%)
Frame = +1
Query: 27 TNNKKKEEQQEQEEEDE---EEELESDDNDNDDDDEIYSSNNASR 68
++++ EE +E+++ DE E+E E DD+D+DDD I +++ +
Sbjct: 88 SSSEGDEEDEEEDQGDEYNLEDEEEDDDDDDDDDLSISEDSDSDK 222
>AW164001
Length = 447
Score = 35.4 bits (80), Expect = 0.021
Identities = 16/42 (38%), Positives = 27/42 (64%), Gaps = 1/42 (2%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQE-QEEEDEEEELESDDNDNDDDDE 59
EF + V ++ K+ E +EED +E+ + DD+D+DDDD+
Sbjct: 1 EFGTSDVVGDSVNKQNDDEVYKEEDVDEDFDDDDDDDDDDDD 126
Score = 33.1 bits (74), Expect = 0.10
Identities = 12/32 (37%), Positives = 23/32 (71%)
Frame = +1
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
N + +E ++E+ DE+ + + DD+D+DDDD+
Sbjct: 37 NKQNDDEVYKEEDVDEDFDDDDDDDDDDDDDD 132
>AV411375
Length = 418
Score = 35.0 bits (79), Expect = 0.027
Identities = 15/32 (46%), Positives = 22/32 (67%)
Frame = +1
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDEIYSS 63
+E QQ++EEEDEE E+E ++ DDD + S
Sbjct: 196 EEPQQQEEEEDEEVEVEEEEASYSDDDTSFLS 291
>TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear
phosphoprotein-like protein, partial (5%)
Length = 789
Score = 34.7 bits (78), Expect = 0.036
Identities = 14/40 (35%), Positives = 25/40 (62%)
Frame = +2
Query: 38 QEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNN 77
+E+EDEEEE +S+++ +DD+DE + + N+N
Sbjct: 38 EEDEDEEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDN 157
Score = 28.5 bits (62), Expect = 2.6
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +2
Query: 34 EQQEQEEEDEEEELESDDNDNDDDDE 59
E+ E EEE+E+ E ES D+D D+ E
Sbjct: 38 EEDEDEEEEEDSE-ESSDDDEDESAE 112
>BF177554
Length = 246
Score = 34.3 bits (77), Expect = 0.047
Identities = 15/39 (38%), Positives = 23/39 (58%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDD 57
EF + Q + +EE E EE+EEEE E ++ + +DD
Sbjct: 1 EFGTSEQQDEDSCEEEAGESSEEEEEEEEEGEEEEEEDD 117
Score = 32.3 bits (72), Expect = 0.18
Identities = 17/44 (38%), Positives = 24/44 (53%)
Frame = +1
Query: 14 GFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDD 57
G S + + + + EE++E+EEE EEEE E DD D D
Sbjct: 7 GTSEQQDEDSCEEEAGESSEEEEEEEEEGEEEE-EEDDGPGDRD 135
>AV416122
Length = 414
Score = 33.9 bits (76), Expect = 0.061
Identities = 13/27 (48%), Positives = 20/27 (73%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
EE+++ EE+ EEE+ E DD D+D + E
Sbjct: 303 EEEEDDEEDKEEEDEEEDDEDDDSNGE 383
Score = 33.5 bits (75), Expect = 0.080
Identities = 19/56 (33%), Positives = 30/56 (52%)
Frame = +3
Query: 18 SEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKG 73
S+ AA N + Q E EEE+++EE + ++ D ++DDE SN + KG
Sbjct: 240 SQPKEAANDDNEPDNDVQFEGEEEEDDEE-DKEEEDEEEDDEDDDSNGEAEVDRKG 404
>TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partial (70%)
Length = 972
Score = 33.9 bits (76), Expect = 0.061
Identities = 14/37 (37%), Positives = 26/37 (69%)
Frame = +2
Query: 34 EQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQF 70
E +E++EED EEE+E ++++ D+ D+ +AS +F
Sbjct: 635 EGEEEDEEDVEEEVEEEEDEEDEYDKRVKEVHASGEF 745
Score = 26.9 bits (58), Expect = 7.5
Identities = 11/23 (47%), Positives = 18/23 (77%)
Frame = +2
Query: 31 KKEEQQEQEEEDEEEELESDDND 53
++E++++ EEE EEEE E D+ D
Sbjct: 641 EEEDEEDVEEEVEEEEDEEDEYD 709
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 33.9 bits (76), Expect = 0.061
Identities = 13/28 (46%), Positives = 21/28 (74%)
Frame = +3
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDE 59
+EE+ +EEE+EEE + D+ N+D+DE
Sbjct: 432 EEEENGEEEEEEEEVVVEDEQGNEDEDE 515
Score = 28.9 bits (63), Expect = 2.0
Identities = 12/36 (33%), Positives = 23/36 (63%)
Frame = +3
Query: 24 AQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
AQ + +++ ++E+EEE+ E E + D D+D+E
Sbjct: 417 AQEEDEEEENGEEEEEEEEVVVEDEQGNEDEDEDEE 524
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.129 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,387,067
Number of Sequences: 28460
Number of extensions: 93157
Number of successful extensions: 1537
Number of sequences better than 10.0: 144
Number of HSP's better than 10.0 without gapping: 757
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1116
length of query: 486
length of database: 4,897,600
effective HSP length: 94
effective length of query: 392
effective length of database: 2,222,360
effective search space: 871165120
effective search space used: 871165120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144592.1


