

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
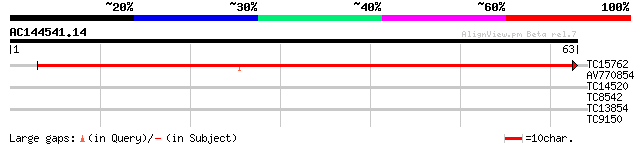
TC15762 similar to GB|BAA25822.1|3088342|AB007158 ribosomal prot... 103 9e-24
AV770854 27 1.0
TC14520 similar to UP|Q8W194 (Q8W194) Zinc finger protein LSD2, ... 25 2.3
TC8542 similar to UP|RK1_PEA (P49208) 50S ribosomal protein L1, ... 25 3.9
TC13854 24 6.6
TC9150 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (Fr... 23 8.6
>TC15762 similar to GB|BAA25822.1|3088342|AB007158 ribosomal protein S23
{Homo sapiens;} , partial (24%)
Length = 572
Score = 103 bits (256), Expect = 9e-24
Identities = 56/65 (86%), Positives = 57/65 (87%), Gaps = 5/65 (7%)
Frame = +1
Query: 4 ERSSSPVQLSKFLGRNKDDGV-----RRSAVSGKKILLKLDKTKEDKEAESKRNELLNFL 58
ERSSSPVQLSKFL R KDDGV RRSAVSGKKILLKL+KTKEDK AESKRNELLNFL
Sbjct: 25 ERSSSPVQLSKFLARGKDDGVKDDGVRRSAVSGKKILLKLEKTKEDKAAESKRNELLNFL 204
Query: 59 NASFD 63
NASFD
Sbjct: 205NASFD 219
>AV770854
Length = 477
Score = 26.6 bits (57), Expect = 1.0
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Frame = +1
Query: 13 SKFLGRN-KDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNF 57
S +LG+ KD+ R SGK + K ++ K S R+EL NF
Sbjct: 241 SSYLGKAIKDENSMRGCRSGKVPIFKTTMRQKLKTHSSSRSELDNF 378
>TC14520 similar to UP|Q8W194 (Q8W194) Zinc finger protein LSD2, partial
(84%)
Length = 1112
Score = 25.4 bits (54), Expect = 2.3
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = +2
Query: 16 LGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRN 52
L +DDG RRS+ KK LK + +ES N
Sbjct: 2 LDEEEDDGSRRSSKEEKKGSLKFSSSSSSFFSESNSN 112
>TC8542 similar to UP|RK1_PEA (P49208) 50S ribosomal protein L1,
chloroplast precursor (Fragment), partial (53%)
Length = 672
Score = 24.6 bits (52), Expect = 3.9
Identities = 11/45 (24%), Positives = 25/45 (55%)
Frame = +3
Query: 7 SSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKR 51
++ V++++ L +DDG + +LLK D+T+ + E ++
Sbjct: 240 AAEVEVAETLDDKEDDGPSKPKTGKAALLLKRDRTRSKRFLEIQK 374
>TC13854
Length = 538
Score = 23.9 bits (50), Expect = 6.6
Identities = 8/22 (36%), Positives = 17/22 (76%)
Frame = +1
Query: 15 FLGRNKDDGVRRSAVSGKKILL 36
F+G+++D+ + S +SGKK+ +
Sbjct: 409 FVGKSRDNLISASVLSGKKLCM 474
>TC9150 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (Fragment),
partial (65%)
Length = 1191
Score = 23.5 bits (49), Expect = 8.6
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +3
Query: 7 SSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEA 47
S+P + S+ LG + +SGK + + L + KED+ A
Sbjct: 99 STPEEASRALGE-----MNGKMISGKPLYVALAQRKEDRRA 206
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 721,095
Number of Sequences: 28460
Number of extensions: 5449
Number of successful extensions: 22
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 21
length of query: 63
length of database: 4,897,600
effective HSP length: 39
effective length of query: 24
effective length of database: 3,787,660
effective search space: 90903840
effective search space used: 90903840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC144541.14


