

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.12 + phase: 0
(417 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
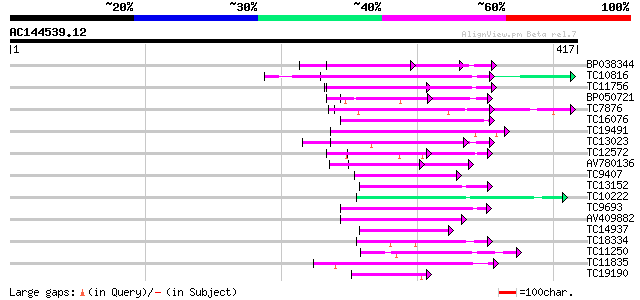
Sequences producing significant alignments: (bits) Value
BP038344 86 1e-17
TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subun... 65 3e-11
TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snR... 64 6e-11
BP050721 62 2e-10
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 61 3e-10
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 53 1e-07
TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 50 9e-07
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 49 1e-06
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 49 2e-06
AV780136 49 2e-06
TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription in... 47 6e-06
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 46 1e-05
TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 45 2e-05
TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit ... 45 2e-05
AV409882 45 3e-05
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 43 8e-05
TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1 (AT3... 42 2e-04
TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle sw... 42 2e-04
TC11835 similar to UP|Q9LG00 (Q9LG00) F20N2.10, partial (35%) 42 2e-04
TC19190 weakly similar to PIR|T46032|T46032 WD-40 repeat regulat... 41 3e-04
>BP038344
Length = 525
Score = 85.5 bits (210), Expect = 1e-17
Identities = 48/145 (33%), Positives = 76/145 (52%)
Frame = +2
Query: 214 IRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETA 273
+RAA A + + +++K + H AV C F G + + S D+ + IWS T
Sbjct: 17 VRAAAMANSSDEPPYKPYRHLKTLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSATL 196
Query: 274 YSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAF 333
L GH I+DLA SS++ + S+S+D +R+W G + LRGHT AV + F
Sbjct: 197 SLLHRLTGHSEGISDLAWSSDSHYICSASDDRTLRIWDATGGDCVKTLRGHTHAVFCVNF 376
Query: 334 SPRPNAVYQLLSSSDDGTCRIWDAR 358
+P+ N ++S S D T R+W+ +
Sbjct: 377 NPQSN---YIVSGSFDETVRVWEVK 442
Score = 53.9 bits (128), Expect = 5e-08
Identities = 26/101 (25%), Positives = 47/101 (45%)
Frame = +2
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+ R+ GH + + Y+ + SDDR ++IW + + RGH + + +
Sbjct: 203 LHRLTGHSEGISDLAWSSDSHYICSASDDRTLRIWDATGGDCVKTLRGHTHAVFCVNFNP 382
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+ + S S D +RVW + G I + HT VT++ F+
Sbjct: 383 QSNYIVSGSFDETVRVWEVKTGRCIHTIIAHTMPVTSVHFN 505
Score = 48.9 bits (115), Expect = 2e-06
Identities = 18/65 (27%), Positives = 37/65 (56%)
Frame = +2
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+K +RGH +AV+C F+ Y+++GS D V++W ++T + + H +T + +
Sbjct: 329 VKTLRGHTHAVFCVNFNPQSNYIVSGSFDETVRVWEVKTGRCIHTIIAHTMPVTSVHFNR 508
Query: 294 NNALV 298
+ L+
Sbjct: 509 DGTLI 523
>TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (20%)
Length = 979
Score = 64.7 bits (156), Expect = 3e-11
Identities = 48/169 (28%), Positives = 74/169 (43%)
Frame = +1
Query: 188 TKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCA 247
TK+N+V GL H + P I A+ ++ + I + H V
Sbjct: 259 TKSNRVKGLSF---------HPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRGV 411
Query: 248 IFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYII 307
F S ++G DD +K+W+ + L + GH+ I + + + S+S+D I
Sbjct: 412 HFHHSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 591
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
R+W ISVL GH V F PR + V +S+S D T R+WD
Sbjct: 592 RIWNWQSRTCISVLTGHNHYVMCALFHPREDLV---VSASLDQTVRVWD 729
Score = 42.0 bits (97), Expect = 2e-04
Identities = 36/188 (19%), Positives = 67/188 (35%)
Frame = +1
Query: 229 QKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITD 288
Q + + + N V F +++ ++++W + H G +
Sbjct: 229 QSKKMLTKFETKSNRVKGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRG 408
Query: 289 LAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSD 348
+ + L S +DY I+VW + L GH + + F + ++S+SD
Sbjct: 409 VHFHHSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFH---HESPWIVSASD 579
Query: 349 DGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGS 408
D T RIW+ +S S +H + C F+ + V+ S
Sbjct: 580 DQTIRIWN-------------------WQSRTCISVLTGHNHYVMCALFHPREDLVVSAS 702
Query: 409 SDNLARVY 416
D RV+
Sbjct: 703 LDQTVRVW 726
Score = 35.0 bits (79), Expect = 0.023
Identities = 34/148 (22%), Positives = 59/148 (38%)
Frame = +1
Query: 208 HHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKI 267
HH +P I +A Q I + GH + V CA+F V++ S D+ V++
Sbjct: 544 HHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHPREDLVVSASLDQTVRV 723
Query: 268 WSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGA 327
W + + R + D+ S D +++ VL GH
Sbjct: 724 WDIGSLK-----RKNASPADDILRLSQMNTDLFGGVDAVVKY----------VLEGHDRG 858
Query: 328 VTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
V +F P A+ ++S++DD ++W
Sbjct: 859 VNWASFHP---ALPLIVSAADDRQVKLW 933
Score = 28.9 bits (63), Expect = 1.6
Identities = 15/47 (31%), Positives = 25/47 (52%), Gaps = 2/47 (4%)
Frame = +1
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSME--TAYSLASCRG 281
+ GH V A F + +++ +DDR VK+W M A+ + + RG
Sbjct: 838 LEGHDRGVNWASFHPALPLIVSAADDRQVKLWRMNDTKAWEVDTLRG 978
>TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snRNP-specific
40 kDa protein (hPrp8-binding) {Homo sapiens;} , partial
(43%)
Length = 719
Score = 63.5 bits (153), Expect = 6e-11
Identities = 35/126 (27%), Positives = 64/126 (50%), Gaps = 1/126 (0%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSME-TAYSLASCRGHVGDITDLAVS 292
I + GH + +Y F+ +G + +GS DR + +W++ + +GH + DL +
Sbjct: 283 IMLLTGHQSVIYTMKFNPAGTVIASGSHDREIFLWNVHGECKNFMVLKGHKNAVLDLHWT 462
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
++ + S+S D +RVW + G + + H V + S R + ++S SDDGT
Sbjct: 463 TDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLSYVNSCCPSRRGPPL--VVSGSDDGTA 636
Query: 353 RIWDAR 358
++WD R
Sbjct: 637 KLWDMR 654
Score = 41.2 bits (95), Expect = 3e-04
Identities = 21/80 (26%), Positives = 38/80 (47%), Gaps = 1/80 (1%)
Frame = +1
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+N ++GH NAV + G +++ S D+ V++W +ET + H+ +
Sbjct: 406 KNFMVLKGHKNAVLDLHWTTDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLSYVNSCCP 585
Query: 292 SSNN-ALVASSSNDYIIRVW 310
S LV S S+D ++W
Sbjct: 586 SRRGPPLVVSGSDDGTAKLW 645
Score = 26.9 bits (58), Expect = 6.3
Identities = 15/50 (30%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +1
Query: 228 VQKMQNIKRIRGHCNAVY-CAIFDRSGRYVITGSDDRLVKIWSMETAYSL 276
V+ + +K++ H + V C R V++GSDD K+W M S+
Sbjct: 520 VETGKQVKKMVEHLSYVNSCCPSRRGPPLVVSGSDDGTAKLWDMRQRGSI 669
>BP050721
Length = 562
Score = 62.0 bits (149), Expect = 2e-10
Identities = 35/114 (30%), Positives = 54/114 (46%), Gaps = 2/114 (1%)
Frame = -1
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDI--TDLAVSSNNALVASS 301
V C G ++ G D V++ + L +GH D +++ +A VA
Sbjct: 502 VNCISMSNDGNCILAGCLDSTVRLLDRTSGELLQEYKGHTNKSYKLDCCLTNTDAHVAGG 323
Query: 302 SNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
S D I W L D +S R HT VT++++ P+ N +LS+S DGT R+W
Sbjct: 322 SEDGFIYFWDLVDAYVVSSFRAHTSVVTSVSYHPKENC---MLSASVDGTIRVW 170
Score = 43.5 bits (101), Expect = 6e-05
Identities = 23/81 (28%), Positives = 40/81 (48%), Gaps = 3/81 (3%)
Frame = -1
Query: 234 IKRIRGHCNAVY---CAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
++ +GH N Y C + + +V GS+D + W + AY ++S R H +T ++
Sbjct: 406 LQEYKGHTNKSYKLDCCLTNTDA-HVAGGSEDGFIYFWDLVDAYVVSSFRAHTSVVTSVS 230
Query: 291 VSSNNALVASSSNDYIIRVWR 311
+ S+S D IRVW+
Sbjct: 229 YHPKENCMLSASVDGTIRVWK 167
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 61.2 bits (147), Expect = 3e-10
Identities = 46/181 (25%), Positives = 85/181 (46%), Gaps = 4/181 (2%)
Frame = +3
Query: 240 HCNAVYCAIFDRSGRY--VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNAL 297
H + V C F S +++ S DR VK+W++ + GH G + +AVS + +L
Sbjct: 513 HSDWVSCVRFSPSTLQPTIVSASWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAVSPDGSL 692
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDA 357
AS D +I +W L +G + L + + A+ FSP L ++ + + +IWD
Sbjct: 693 CASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPN----RYWLCAATETSIKIWDL 857
Query: 358 RYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFN--ANGTVFVTGSSDNLARV 415
L V +++ + G ++ + I+C + N A+G+ +G +D + RV
Sbjct: 858 ESKSIVEDLKVDLKTEADATTGGNTN----KKKVIYCTSLNWSADGSTLFSGYTDGVIRV 1025
Query: 416 Y 416
+
Sbjct: 1026W 1028
Score = 56.6 bits (135), Expect = 7e-09
Identities = 31/124 (25%), Positives = 61/124 (49%), Gaps = 2/124 (1%)
Frame = +3
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
+R+ GH + V + G++ ++GS D +++W + S GH D+ +A S +
Sbjct: 240 RRLTGHSHFVQDVVLSSDGQFALSGSWDGELRLWDLAAGTSARRFVGHTKDVLSVAFSID 419
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVL--RGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N + S+S D I++W ++ H+ V+ + FSP ++S+S D T
Sbjct: 420 NRQIVSASRDRTIKLWNTLGECKYTIQDNDAHSDWVSCVRFSP-STLQPTIVSASWDRTV 596
Query: 353 RIWD 356
++W+
Sbjct: 597 KVWN 608
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 52.8 bits (125), Expect = 1e-07
Identities = 27/114 (23%), Positives = 59/114 (51%), Gaps = 1/114 (0%)
Frame = +3
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
V C F G+ + + DD+ V +W+M+T + ++ H I+D+ N++ +A++S
Sbjct: 3 VTCCHFSSDGKMLASAGDDKKVVLWNMDTLQTESTPEQHKSVISDVRFRPNSSQLATASI 182
Query: 304 DYIIRVWRLPD-GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
D +R+W + + GH+ A+ ++ F P+ ++ S+++ R W+
Sbjct: 183 DKSVRLWDAANPTYCVQEYNGHSSAIMSLDFHPKKTDIFCFCDSANE--IRYWN 338
Score = 31.2 bits (69), Expect = 0.33
Identities = 19/78 (24%), Positives = 38/78 (48%)
Frame = +3
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
I+ + N Y +F S ++ +++W+M S+ + H I+ LA S
Sbjct: 597 IQELSSSGNQFYSCVFHPSYSTLLVIGGFSSLELWNMADNKSM-TISAHESVISALAQSP 773
Query: 294 NNALVASSSNDYIIRVWR 311
+VAS+S+D +++W+
Sbjct: 774 VTGMVASASHDNSVKLWK 827
>TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (39%)
Length = 527
Score = 49.7 bits (117), Expect = 9e-07
Identities = 36/138 (26%), Positives = 61/138 (44%), Gaps = 7/138 (5%)
Frame = +1
Query: 237 IRGHCNAVYCAIFDR-SGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNN 295
+ GH V ++ R + T SDD V ++ E + + GH + + VS +
Sbjct: 4 LEGHFMPVRSLVYSPYDPRLLFTASDDGNVHMYDAEGKALVGTMSGHASWVLCVDVSPDG 183
Query: 296 ALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVY---QLLSSSDDGTC 352
+A+ S+D +R+W L + + HT V +AF P +L S SDD +
Sbjct: 184 GAIATGSSDRTVRLWDLAMRASVQTMSNHTDQVWGVAFRPPGGTDVRGGRLASVSDDKSI 363
Query: 353 RIWD---ARYTHSTPRLY 367
++D + + H T LY
Sbjct: 364 SLYDYS*S*FRHCTQCLY 417
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 49.3 bits (116), Expect = 1e-06
Identities = 22/101 (21%), Positives = 44/101 (42%)
Frame = +1
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
++G +V + + R +++W + T + S +GH G + ++ +
Sbjct: 271 LQGDSESVTALALSPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGPVMCMSCHPSGG 450
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
L+A+ D + VW + G +GH G V+ + F P P
Sbjct: 451 LLATGGADRKVLVWDVDGGYCTHFFKGHGGVVSCVMFHPDP 573
Score = 41.2 bits (95), Expect = 3e-04
Identities = 34/149 (22%), Positives = 62/149 (40%), Gaps = 8/149 (5%)
Frame = +1
Query: 216 AACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAI-FDRSGRYVITGSDDRL-------VKI 267
A+C+ +P ++ + R++ + V F G YV++ + +KI
Sbjct: 58 ASCFFFRRPGPLLSESMEALRLKSNYRCVPALQQFYTGGPYVVSSDGSFIACACGESIKI 237
Query: 268 WSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGA 327
A ++ +G +T LA+S ++ L+ SS + IRVW L + +GH G
Sbjct: 238 VDSANASIRSTLQGDSESVTALALSPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGP 417
Query: 328 VTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
V ++ P L + D +WD
Sbjct: 418 VMCMSCHPSGGL---LATGGADRKVLVWD 495
Score = 34.3 bits (77), Expect = 0.039
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDIT 287
++ +GH V C SG + TG DR V +W ++ Y +GH G ++
Sbjct: 388 VRSWKGHEGPVMCMSCHPSGGLLATGGADRKVLVWDVDGGYCTHFFKGHGGVVS 549
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 48.9 bits (115), Expect = 2e-06
Identities = 27/110 (24%), Positives = 52/110 (46%), Gaps = 3/110 (2%)
Frame = +3
Query: 249 FDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGD---ITDLAVSSNNALVASSSNDY 305
F + ++++ G+ D +++W+ T L + GHV I+ ++N V S D+
Sbjct: 192 FSPNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDH 371
Query: 306 IIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
I +W L + L GH+ V +++ P N + + +D T +IW
Sbjct: 372 GIYLWELQTRKIVQKLEGHSDTVVSVSCHPTENMIAS-GALGNDKTVKIW 518
Score = 47.4 bits (111), Expect = 4e-06
Identities = 21/82 (25%), Positives = 41/82 (49%), Gaps = 5/82 (6%)
Frame = +3
Query: 234 IKRIRGHCNAVYC---AIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
+K GH N+ YC +G+YV+ GS+D + +W ++T + GH + ++
Sbjct: 273 LKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDHGIYLWELQTRKIVQKLEGHSDTVVSVS 452
Query: 291 VSSNNALVASSS--NDYIIRVW 310
++AS + ND +++W
Sbjct: 453 CHPTENMIASGALGNDKTVKIW 518
Score = 38.5 bits (88), Expect = 0.002
Identities = 24/124 (19%), Positives = 51/124 (40%), Gaps = 1/124 (0%)
Frame = +3
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGD-ITDLAVS 292
+K + H + V F+R G +++ S D L +IW T + + + ++ + S
Sbjct: 18 LKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPVSFVKFS 197
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N + + D +R+W G + GH + I+ + ++ S+D
Sbjct: 198 PNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDHGI 377
Query: 353 RIWD 356
+W+
Sbjct: 378 YLWE 389
Score = 36.6 bits (83), Expect = 0.008
Identities = 20/47 (42%), Positives = 27/47 (56%)
Frame = +3
Query: 315 GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTH 361
G + VL H+ VTA+ F+ + + +SSS DG CRIWDA H
Sbjct: 9 GKCLKVLPAHSDPVTAVDFNRDGSLI---VSSSYDGLCRIWDASTGH 140
Score = 31.2 bits (69), Expect = 0.33
Identities = 31/131 (23%), Positives = 55/131 (41%), Gaps = 1/131 (0%)
Frame = +3
Query: 282 HVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPI-SVLRGHTGAVTAIAFSPRPNAV 340
H +T + + + +L+ SSS D + R+W G + +++ V+ + FS PNA
Sbjct: 36 HSDPVTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPVSFVKFS--PNAK 209
Query: 341 YQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNAN 400
+ L+ + D+ R+W+ STGR + + + I N
Sbjct: 210 FILVGTLDN-NLRLWNY----------------STGRFLKTYTGHVNSKYCISSTFSTTN 338
Query: 401 GTVFVTGSSDN 411
G V GS D+
Sbjct: 339 GKYVVGGSEDH 371
>AV780136
Length = 558
Score = 48.9 bits (115), Expect = 2e-06
Identities = 23/92 (25%), Positives = 45/92 (48%)
Frame = -1
Query: 250 DRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRV 309
+ G +++G ++++++W + RGH +I L + S S S+D +IR+
Sbjct: 546 NEGGTVLVSGGTEKVIRVWDPRSGSKTLKLRGHADNIRALLLDSTGRYCLSGSSDSMIRL 367
Query: 310 WRLPDGLPISVLRGHTGAVTAIAFSPRPNAVY 341
W + + HT +V A+A +P + VY
Sbjct: 366 WDIGQPRCLHTYAVHTDSVWALASTPTFSHVY 271
Score = 42.4 bits (98), Expect = 1e-04
Identities = 18/70 (25%), Positives = 36/70 (50%)
Frame = -1
Query: 236 RIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNN 295
++RGH + + + D +GRY ++GS D ++++W + L + H + LA +
Sbjct: 462 KLRGHADNIRALLLDSTGRYCLSGSSDSMIRLWDIGQPRCLHTYAVHTDSVWALASTPTF 283
Query: 296 ALVASSSNDY 305
+ V S D+
Sbjct: 282 SHVYSGGRDF 253
Score = 35.8 bits (81), Expect = 0.013
Identities = 20/68 (29%), Positives = 30/68 (43%)
Frame = -1
Query: 289 LAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSD 348
LA++ ++ S + +IRVW G LRGH + A+ + LS S
Sbjct: 555 LAMNEGGTVLVSGGTEKVIRVWDPRSGSKTLKLRGHADNIRALLLD---STGRYCLSGSS 385
Query: 349 DGTCRIWD 356
D R+WD
Sbjct: 384 DSMIRLWD 361
>TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 530
Score = 47.0 bits (110), Expect = 6e-06
Identities = 26/79 (32%), Positives = 40/79 (49%)
Frame = +2
Query: 254 RYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLP 313
RY+ T S D VKIW+++ + GH + D S + A + ++S+D R+W +
Sbjct: 53 RYLATASADHTVKIWNVDGFTLDKTLIGHQRWVWDCVFSVDGAYLITASSDTTARLWSMS 232
Query: 314 DGLPISVLRGHTGAVTAIA 332
G I V +GH A T A
Sbjct: 233 TGEDIKVYQGHHKATTCCA 289
Score = 38.9 bits (89), Expect = 0.002
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = +2
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGH 282
K + GH V+ +F G Y+IT S D ++WSM T + +GH
Sbjct: 122 KTLIGHQRWVWDCVFSVDGAYLITASSDTTARLWSMSTGEDIKVYQGH 265
Score = 28.9 bits (63), Expect = 1.6
Identities = 19/59 (32%), Positives = 33/59 (55%), Gaps = 1/59 (1%)
Frame = +2
Query: 298 VASSSNDYIIRVWRLPDGLPI-SVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+A++S D+ +++W + DG + L GH V FS + Y L+++S D T R+W
Sbjct: 59 LATASADHTVKIWNV-DGFTLDKTLIGHQRWVWDCVFS--VDGAY-LITASSDTTARLW 223
Score = 26.6 bits (57), Expect = 8.2
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = +2
Query: 392 IFCCAFNANGTVFVTGSSDNLARVYT 417
++ C F+ +G +T SSD AR+++
Sbjct: 149 VWDCVFSVDGAYLITASSDTTARLWS 226
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 45.8 bits (107), Expect = 1e-05
Identities = 27/99 (27%), Positives = 47/99 (47%), Gaps = 1/99 (1%)
Frame = +2
Query: 258 TGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPD-GL 316
+G D+ +W ++ A+ H ITD+ S + +A+SS D ++VW + + G
Sbjct: 11 SGGHDKKAVLWYTDSLKQKATLEEHSALITDVRFSPSMPRLATSSFDKTVKVWDVDNPGY 190
Query: 317 PISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+ GH+ +V ++ F PN + S DG R W
Sbjct: 191 SLRTFTGHSASVMSLDF--HPNKEDLICSCDSDGEIRYW 301
>TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (55%)
Length = 557
Score = 45.4 bits (106), Expect = 2e-05
Identities = 31/155 (20%), Positives = 61/155 (39%)
Frame = +1
Query: 256 VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDG 315
++TGS D V++W + + GH + +A ++ ASSS D +RV+ +
Sbjct: 109 LLTGSLDETVRLWRSDELVLERTNTGHCLGVASVAAHPLGSIAASSSLDSFVRVFDVDSN 288
Query: 316 LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDST 375
++ L V + F P+ ++ + ++WD L +P+P +
Sbjct: 289 ATLATLEAPPSEVWQMRFDPK--GALLAVAGGGSASVKLWDTSTWELVATLSIPRPEGGS 462
Query: 376 GRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSD 410
+ S + A++ +G GS D
Sbjct: 463 KPTDKSGSKKF-----VLSVAWSPDGKRIACGSMD 552
>TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit
(Beta'-coat protein) (Beta'-COP) (p102), partial (20%)
Length = 743
Score = 45.1 bits (105), Expect = 2e-05
Identities = 30/112 (26%), Positives = 53/112 (46%), Gaps = 1/112 (0%)
Frame = +1
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
V A F ++V+ G+DD +++++ T + H I +AV V SSS+
Sbjct: 385 VRSAKFIARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIRCVAVHPTLPYVLSSSD 564
Query: 304 DYIIRVWRLPDG-LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI 354
D +I++W G + + H+ V + F+P+ + S+S D T RI
Sbjct: 565 DMLIKLWDWEKGWICTQICEAHSHYVMQVTFNPKDTNTF--ASASLDRTIRI 714
Score = 37.4 bits (85), Expect = 0.005
Identities = 22/84 (26%), Positives = 37/84 (43%), Gaps = 2/84 (2%)
Frame = +1
Query: 231 MQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLAS-CRGHVGDITDL 289
M +K H + + C + YV++ SDD L+K+W E + C H + +
Sbjct: 472 MDKVKVFEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQICEAHSHYVMQV 651
Query: 290 AVSSNNA-LVASSSNDYIIRVWRL 312
+ + AS+S D IR+ L
Sbjct: 652 TFNPKDTNTFASASLDRTIRIRNL 723
Score = 35.8 bits (81), Expect = 0.013
Identities = 27/97 (27%), Positives = 44/97 (44%), Gaps = 5/97 (5%)
Frame = +1
Query: 265 VKIWSMETAYSLASCRGHVGDITDLAVSSNNAL-----VASSSNDYIIRVWRLPDGLPIS 319
V IW+ +T S ++T+L V S + V + ++D IRV+ +
Sbjct: 322 VCIWNYQTQTMAKSF-----EVTELPVRSAKFIARKQWVVAGADDMFIRVYNYNTMDKVK 486
Query: 320 VLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
V HT + +A P + +LSSSDD ++WD
Sbjct: 487 VFEAHTDYIRCVAVHP---TLPYVLSSSDDMLIKLWD 588
>AV409882
Length = 422
Score = 44.7 bits (104), Expect = 3e-05
Identities = 25/94 (26%), Positives = 46/94 (48%), Gaps = 1/94 (1%)
Frame = +3
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
V A F ++V+ G+DD +++++ T + H I +AV V SSS+
Sbjct: 105 VRSAKFVARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIRCVAVHPTLPYVLSSSD 284
Query: 304 DYIIRVWRLPDG-LPISVLRGHTGAVTAIAFSPR 336
D +I++W G + + GH+ V + F+P+
Sbjct: 285 DMLIKLWDWEKGWICTQIFEGHSHYVMQVTFNPK 386
Score = 34.3 bits (77), Expect = 0.039
Identities = 22/77 (28%), Positives = 37/77 (47%), Gaps = 5/77 (6%)
Frame = +3
Query: 285 DITDLAVSSNNAL-----VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNA 339
++T+L V S + V + ++D IRV+ + V HT + +A P
Sbjct: 87 EVTELPVRSAKFVARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIRCVAVHP---T 257
Query: 340 VYQLLSSSDDGTCRIWD 356
+ +LSSSDD ++WD
Sbjct: 258 LPYVLSSSDDMLIKLWD 308
Score = 33.5 bits (75), Expect = 0.067
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = +3
Query: 231 MQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAY 274
M +K H + + C + YV++ SDD L+K+W E +
Sbjct: 192 MDKVKVFEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGW 323
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 43.1 bits (100), Expect = 8e-05
Identities = 22/69 (31%), Positives = 32/69 (45%)
Frame = +1
Query: 258 TGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLP 317
+ S D VK+W +E + S GH + +A S N +AS S D I +W L +G
Sbjct: 61 SASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIWSLKEGKI 240
Query: 318 ISVLRGHTG 326
+ G G
Sbjct: 241 VKTYTGSGG 267
Score = 38.1 bits (87), Expect = 0.003
Identities = 23/62 (37%), Positives = 34/62 (54%)
Frame = +1
Query: 297 LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
++AS+S D +++W + G I L GH V ++AFS PN Y L S S D + IW
Sbjct: 52 VLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFS--PNGEY-LASGSPDKSIHIWS 222
Query: 357 AR 358
+
Sbjct: 223 LK 228
Score = 36.6 bits (83), Expect = 0.008
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVG 284
I + GH + VY F +G Y+ +GS D+ + IWS++ + + G G
Sbjct: 115 IYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIWSLKEGKIVKTYTGSGG 267
>TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1
(AT3g18860/MCB22_3), partial (21%)
Length = 595
Score = 42.0 bits (97), Expect = 2e-04
Identities = 35/107 (32%), Positives = 49/107 (45%), Gaps = 7/107 (6%)
Frame = +2
Query: 256 VITGSDDRLVKIWSMETAYSLAS--CRGHVGDITDLA-VSSNNAL----VASSSNDYIIR 308
+ T S D+ V+ WS E ++S GH + L + N L VAS D ++
Sbjct: 191 IATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVGPLVWIPPNPELPQGGVASGGMDTLVL 370
Query: 309 VWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
VW L G L+GH VT+IAF ++S+S D T R W
Sbjct: 371 VWDLSTGEKAHTLKGHQLQVTSIAFDDG-----DVVSASVDCTLRRW 496
>TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle switch
protein ccs52B, partial (28%)
Length = 564
Score = 42.0 bits (97), Expect = 2e-04
Identities = 33/123 (26%), Positives = 55/123 (43%), Gaps = 5/123 (4%)
Frame = +2
Query: 259 GSDDRLVKIWSMETAYSLASCRGHV---GDITDLAVSSN-NALVASSS-NDYIIRVWRLP 313
G+ DR ++ W+ + L HV + +LA S N N +V++ + I VW+ P
Sbjct: 74 GTADRCIRFWNTTNGHQL----NHVDTGSQVCNLAWSKNVNEIVSTHGYSQNQIMVWKYP 241
Query: 314 DGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSD 373
++ L GH+ V +A SP + ++ + D T R W+ P + P P
Sbjct: 242 SLSKVATLTGHSMRVLYLAMSPDGQTI---VTGAGDETLRFWNV-----FPSMKTPAPVK 397
Query: 374 STG 376
TG
Sbjct: 398 DTG 406
>TC11835 similar to UP|Q9LG00 (Q9LG00) F20N2.10, partial (35%)
Length = 769
Score = 41.6 bits (96), Expect = 2e-04
Identities = 31/140 (22%), Positives = 60/140 (42%), Gaps = 4/140 (2%)
Frame = +3
Query: 224 PSTMVQKMQNIKRIR---GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET-AYSLASC 279
P +M+ Q K I GH + + + + G TG+ D+ +IW M + S+A
Sbjct: 39 PESMLVDSQTGKTIISLCGHLDFSFASAWHPDGMTFATGNQDKTCRIWDMRNLSKSVAVL 218
Query: 280 RGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNA 339
+G++G I + +S+ +A + + V+ + G G ++ I+FSP +
Sbjct: 219 KGNLGAIRSIRYTSDGQYLAMAEPADFVHVYDVKSGYEREQEIDFFGEISGISFSPDTES 398
Query: 340 VYQLLSSSDDGTCRIWDARY 359
++ +WD Y
Sbjct: 399 LF----------IGVWDRTY 428
>TC19190 weakly similar to PIR|T46032|T46032 WD-40 repeat regulatory protein
tup1 homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (18%)
Length = 511
Score = 41.2 bits (95), Expect = 3e-04
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 2/61 (3%)
Frame = +3
Query: 252 SGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASS--SNDYIIRV 309
+GRY+++GS+DR V +W ++ + GH + + +AS+ D +RV
Sbjct: 12 NGRYIVSGSEDRCVYLWDLQQKNMIQKLEGHTDTVISVTCHPTENKIASAGLDGDRTVRV 191
Query: 310 W 310
W
Sbjct: 192 W 194
Score = 33.9 bits (76), Expect = 0.051
Identities = 18/63 (28%), Positives = 26/63 (40%)
Frame = +3
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
+N + S S D + +W L I L GHT V ++ P N + D T
Sbjct: 9 TNGRYIVSGSEDRCVYLWDLQQKNMIQKLEGHTDTVISVTCHPTENKIAS-AGLDGDRTV 185
Query: 353 RIW 355
R+W
Sbjct: 186 RVW 194
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,792,659
Number of Sequences: 28460
Number of extensions: 147508
Number of successful extensions: 937
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 835
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 916
length of query: 417
length of database: 4,897,600
effective HSP length: 93
effective length of query: 324
effective length of database: 2,250,820
effective search space: 729265680
effective search space used: 729265680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144539.12


