

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.1 + phase: 0
(163 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
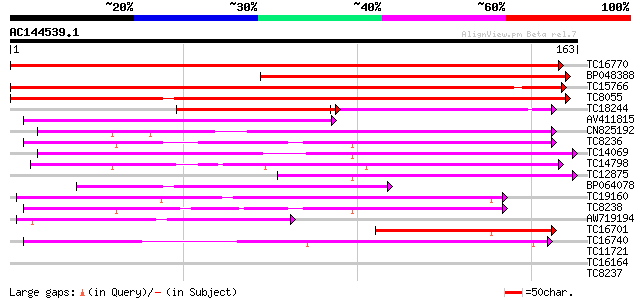
Score E
Sequences producing significant alignments: (bits) Value
TC16770 weakly similar to GB|AAL49942.1|17979251|AY070476 AT3g53... 171 4e-44
BP048388 136 2e-33
TC15766 similar to GB|AAL49942.1|17979251|AY070476 AT3g53990/F5K... 135 3e-33
TC8055 similar to UP|Q9FES5 (Q9FES5) Early nodulin ENOD18, parti... 134 8e-33
TC18244 weakly similar to UP|Q9LSP5 (Q9LSP5) Gb|AAF26101.1 (AT3g... 59 2e-21
AV411815 80 2e-16
CN825192 78 7e-16
TC8236 71 8e-14
TC14069 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fr... 68 7e-13
TC14798 similar to UP|Q84TF6 (Q84TF6) At1g09740, partial (85%) 64 2e-11
TC12875 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fr... 62 5e-11
BP064078 58 9e-10
TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%) 57 2e-09
TC8238 55 5e-09
AW719194 53 3e-08
TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome ... 51 1e-07
TC16740 similar to UP|Q9SIJ8 (Q9SIJ8) Expressed protein (RD2 pro... 49 4e-07
TC11721 36 0.004
TC16164 similar to GB|AAN28779.1|23505839|AY143840 At4g27320/M4I... 35 0.006
TC8237 33 0.024
>TC16770 weakly similar to GB|AAL49942.1|17979251|AY070476
AT3g53990/F5K20_290 {Arabidopsis thaliana;}, partial
(97%)
Length = 781
Score = 171 bits (434), Expect = 4e-44
Identities = 79/159 (49%), Positives = 109/159 (67%)
Frame = +2
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M+S R +G+A+DFS S A W VDN+++ GD L +I I P + LW TGSPL
Sbjct: 59 MSSDRNIGVALDFSKGSKTALNWAVDNLLRNGDTLYIIHINPPQDSESRNLLWSTTGSPL 238
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ + YE+ TD EVL + TA QK+V ++ K+YWGDAREK+ +A+E + L
Sbjct: 239 IPLSEFREREVMRHYEVDTDAEVLDLLDTASRQKQVTIVAKLYWGDAREKIVDAVEDLKL 418
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D L MG+RGLG ++R ++GSVS YV +NA+CPVT+VK S
Sbjct: 419 DALVMGSRGLGAIQRVLLGSVSTYVTSNANCPVTIVKDS 535
>BP048388
Length = 449
Score = 136 bits (342), Expect = 2e-33
Identities = 65/89 (73%), Positives = 77/89 (86%)
Frame = -3
Query: 73 KKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGT 132
K+Y +K +PEV+ IA TA ++K + VL+K+YWGDAREKL EAIE +PLD + MGNRGLGT
Sbjct: 447 KRYGLKPEPEVIDIAPTASQEKNIEVLLKIYWGDAREKLLEAIEHIPLDSIIMGNRGLGT 268
Query: 133 LRRAIMGSVSNYVVNNASCPVTVVKSSGQ 161
LRRAIMGSVSN+VVNNASCPVTVVKSS Q
Sbjct: 267 LRRAIMGSVSNHVVNNASCPVTVVKSSEQ 181
>TC15766 similar to GB|AAL49942.1|17979251|AY070476 AT3g53990/F5K20_290
{Arabidopsis thaliana;}, partial (98%)
Length = 995
Score = 135 bits (340), Expect = 3e-33
Identities = 65/160 (40%), Positives = 99/160 (61%)
Frame = +1
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
MA R +G+A+DFS S A +W ++N+ +GDN+ +I I P + +LW +GSPL
Sbjct: 82 MAKDRTIGVALDFSKSSKNALKWALENLADKGDNIYIIHINPNSLDESRNKLWGKSGSPL 261
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ KY+++ D EVL + TA QK+V ++ K+YWGDARE+L +A+E + L
Sbjct: 262 IPLKEFREPEVMTKYDVQIDIEVLDLLDTASRQKEVNIVTKIYWGDAREQLLDAVEDLKL 441
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSG 160
D L MG+RGL T++R I+ + + CP+ S G
Sbjct: 442 DSLVMGSRGLSTIQRIILXKCEQFC--DDPCPLPCDYSQG 555
Score = 32.0 bits (71), Expect = 0.054
Identities = 13/20 (65%), Positives = 17/20 (85%)
Frame = +2
Query: 140 SVSNYVVNNASCPVTVVKSS 159
SVSN+V+ +A CPVT+VK S
Sbjct: 500 SVSNFVMTHAPCPVTIVKDS 559
>TC8055 similar to UP|Q9FES5 (Q9FES5) Early nodulin ENOD18, partial (89%)
Length = 625
Score = 134 bits (337), Expect = 8e-33
Identities = 63/161 (39%), Positives = 101/161 (62%)
Frame = +1
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R++G+A DFS S A +W ++N+ +GD +I + + +W +GSPL
Sbjct: 22 MVEDRKVGVATDFSKSSNSALKWAIENMADKGDTFYIIHVMSDG---SRTNIWAKSGSPL 192
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL + Y ++TDPEVL + A QK+V + K+YWG+AR+KL ++IE + L
Sbjct: 193 IPLSILRQPEAMSNYGVQTDPEVLDMLDAAAGQKEVNFVAKLYWGEARQKLIDSIEDLKL 372
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQ 161
D L MG+RG G+++R +MGSVSN+++ +A+CPV +V+ S +
Sbjct: 373 DSLVMGSRGRGSIKRILMGSVSNFLMIHATCPVAIVRDSSK 495
>TC18244 weakly similar to UP|Q9LSP5 (Q9LSP5) Gb|AAF26101.1
(AT3g17020/K14A17_14), partial (26%)
Length = 451
Score = 58.9 bits (141), Expect(2) = 2e-21
Identities = 26/47 (55%), Positives = 37/47 (78%)
Frame = +2
Query: 49 EMQLWEVTGSPLTPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKK 95
EMQLWE TGSPL PL +F ++ + K+Y +K +PEV+ IATTA +++K
Sbjct: 47 EMQLWETTGSPLIPLSDFSDTAVLKRYGLKPEPEVIDIATTASKEEK 187
Score = 58.5 bits (140), Expect(2) = 2e-21
Identities = 31/65 (47%), Positives = 39/65 (59%)
Frame = +3
Query: 93 QKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCP 152
+K + VL+K+YWGDAREKL EAIE +PLD + MG + G N+VVN P
Sbjct: 180 KKNIEVLLKIYWGDAREKLLEAIEHIPLDSIIMGEXRPWHSHKGHXGXCHNHVVNTL-LP 356
Query: 153 VTVVK 157
VVK
Sbjct: 357 RPVVK 371
Score = 28.1 bits (61), Expect = 0.78
Identities = 38/133 (28%), Positives = 57/133 (42%), Gaps = 14/133 (10%)
Frame = +1
Query: 35 LILIIIRPEE-YEHG-EMQLWEVTGSPLTPLGEFINSDLPK------------KYEIKTD 80
LIL+IIRP+E YE G + L + T + F + L + +
Sbjct: 1 LILVIIRPQEYYERGRDAALGNHRLTSYTTV*FF*HCGLEEIWAQT*T*SD*YCHNCLQG 180
Query: 81 PEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGS 140
++L+ +I + + + YW L A L+ GN GLGTL RAI G
Sbjct: 181 RKILRCC*RSIGEMR----ARSYWKQLNTYLWTA--------LSWGNXGLGTLIRAIXGX 324
Query: 141 VSNYVVNNASCPV 153
V+ ++ SCPV
Sbjct: 325 VTT-ML*IXSCPV 360
>AV411815
Length = 423
Score = 80.1 bits (196), Expect = 2e-16
Identities = 39/90 (43%), Positives = 53/90 (58%)
Frame = +2
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R +GIAMDFSP S A +W VDN++ GD +I+I + P +H +L+ GSPL P+
Sbjct: 152 RIVGIAMDFSPTSKLALRWAVDNLINRGDQIIIINVEPPNADHTRKELFAENGSPLVPME 331
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQK 94
E + K+Y I DPEV+ I TA K
Sbjct: 332 ELREINFTKQYGIARDPEVIDILDTASRTK 421
>CN825192
Length = 697
Score = 78.2 bits (191), Expect = 7e-16
Identities = 47/167 (28%), Positives = 80/167 (47%), Gaps = 18/167 (10%)
Frame = +3
Query: 9 IAMDFSPCSIKAFQWTVDNI-------------VKEGDNLILII-----IRPEEYEHGEM 50
+A+D S CS A +W +DN+ +++G ++ ++ P Y G
Sbjct: 87 VAVDESDCSFHALKWALDNVLNNMTTTATPDENIEDGGGMVFLVHVEPAFHPAVYPIGTS 266
Query: 51 QLWEVTGSPLTPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREK 110
L+ + S DL +K + + L A ++ + GDARE
Sbjct: 267 ALYPASASL---------EDLMRKAQREKSTSTLSRALQMCRDNQIKAESIILTGDAREM 419
Query: 111 LCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+C+A +Q+ +D L MG+RGLG L+R +GSVS+Y ++A P+ +VK
Sbjct: 420 ICQAADQMHVDLLIMGSRGLGMLKRTFLGSVSDYCAHHAKTPILIVK 560
>TC8236
Length = 840
Score = 71.2 bits (173), Expect = 8e-14
Identities = 47/161 (29%), Positives = 79/161 (48%), Gaps = 8/161 (4%)
Frame = +3
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIV--KEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTP 62
RR+ + +D S+ A W + N+ + D+LIL+ ++P V S
Sbjct: 141 RRIMVTVDEGDESMYALSWCLKNLAFQNDKDHLILLYVKPPR----------VVYSAFDG 290
Query: 63 LGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVV------VLVKVYWGDAREKLCEAIE 116
G +SD+ E + ++A +E+ K + V VK GD R+ +C+ ++
Sbjct: 291 TGYLFSSDITATMERVSQ----QVAEGVLERAKGLCNNVENVEVKAESGDPRDVICQMVQ 458
Query: 117 QVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+ +D L MG+ G G ++RA +GSVSN+ N CPV +VK
Sbjct: 459 KWGVDVLVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVVIVK 581
>TC14069 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fragment),
partial (58%)
Length = 824
Score = 68.2 bits (165), Expect = 7e-13
Identities = 45/163 (27%), Positives = 74/163 (44%), Gaps = 8/163 (4%)
Frame = +3
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFIN 68
I +D S S+ A WT+D+ + + L+++ + + + ++
Sbjct: 162 IGIDDSEHSVYAINWTLDHFFAKNPSFKLVLVHARPSATSAVGFAGPVYAGAAEVLPIVD 341
Query: 69 SDLPKKYEIKTDPEVLKIATTAIEQKKVV--------VLVKVYWGDAREKLCEAIEQVPL 120
SDL K IA +E K + V+V+ GD R LCEA+E+
Sbjct: 342 SDLKK------------IAARVLENAKQICIKNNITDVVVEAVEGDPRNVLCEAVEKYHA 485
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
L +G+ G G L+RA++GSVS+Y ++A C V +VK H
Sbjct: 486 SVLVVGSHGYGALKRAVLGSVSDYCAHHAHCSVMIVKKPKTKH 614
>TC14798 similar to UP|Q84TF6 (Q84TF6) At1g09740, partial (85%)
Length = 1309
Score = 63.5 bits (153), Expect = 2e-11
Identities = 47/170 (27%), Positives = 79/170 (45%), Gaps = 17/170 (10%)
Frame = +3
Query: 7 LGIAMDFSPCSIKAFQWTVDNI---------VKEGDNLILIIIRPEEYEHGEMQLWEVTG 57
+ +A+D S S+ A +W + N+ G LIL + P G +
Sbjct: 282 VAVAVDGSDESMNALRWAMTNLKLRPQTPDSTTAGCFLILHVQSPPSIATG------LNP 443
Query: 58 SPLTPLGEFINSDLP------KKYEIKTDPEVLKIATTAIEQKKVVVLVK--VYWGDARE 109
P+ P G N ++P + ++ + +L A + V+ V GD +E
Sbjct: 444 GPI-PFGGPSNLEVPAFAAAIEAHQKRITDSILDHALGICSEFNFTEKVRTHVVIGDPKE 620
Query: 110 KLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
K+CEA++ D L MG+R G ++R +GSVSNY ++A CPV ++K +
Sbjct: 621 KICEAVQDQHADVLVMGSRAFGPIKRMFLGSVSNYCAHHAECPVIIIKGN 770
>TC12875 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fragment),
partial (64%)
Length = 668
Score = 62.0 bits (149), Expect = 5e-11
Identities = 34/87 (39%), Positives = 50/87 (57%), Gaps = 1/87 (1%)
Frame = -3
Query: 78 KTDPEVLKIATTAIEQKKVV-VLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRA 136
+T +V IA QK V V V+V GDAR LC+ +E+ L +G+ G G ++RA
Sbjct: 480 RTAAKVTDIAKELCVQKSVTDVTVEVIEGDARNVLCDVVEKHHASILVVGSHGYGAIKRA 301
Query: 137 IMGSVSNYVVNNASCPVTVVKSSGQHH 163
++GSVS+Y ++A C V +VK H
Sbjct: 300 VLGSVSDYCAHHAHCTVMIVKKPKTKH 220
>BP064078
Length = 583
Score = 57.8 bits (138), Expect = 9e-10
Identities = 27/91 (29%), Positives = 48/91 (52%)
Frame = -3
Query: 20 AFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKYEIKT 79
A +W ++N+ +GD +I + + +W +GSPL PL + Y ++T
Sbjct: 578 ALKWAIENMADKGDTFYIIHVMSDG---SRTNIWAKSGSPLIPLSILRQPEAMSNYGVQT 408
Query: 80 DPEVLKIATTAIEQKKVVVLVKVYWGDAREK 110
DPEVL + A K+V + K+YWG ++++
Sbjct: 407 DPEVLDMLDAAAGPKEVNFVAKLYWG*SQDQ 315
>TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%)
Length = 557
Score = 57.0 bits (136), Expect = 2e-09
Identities = 42/150 (28%), Positives = 74/150 (49%), Gaps = 9/150 (6%)
Frame = +1
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRP------EEYEHGEMQLWEVT 56
S R++ IA+D S S A +W V N ++ GD +IL+ +RP ++ G+ +
Sbjct: 115 SHRKIAIAVDLSDESAYAVRWAVQNYLRPGDEVILLHVRPTSVLYGADWGAGDHNAEDGD 294
Query: 57 GSPLTPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIE 116
G + E L ++ T + +A +E++ + V D +E+LC +E
Sbjct: 295 GGDSS---EESRRKLEDDFDNFTSTKASDLAEPLVEEQIPFKIHIVKDHDMKERLCLEVE 465
Query: 117 QVPLDGLTMGNRGLGTLRRAI---MGSVSN 143
++ L + MG+RG G +RA +GSVS+
Sbjct: 466 RLGLSAVIMGSRGFGASKRAAKGRLGSVSD 555
>TC8238
Length = 506
Score = 55.5 bits (132), Expect = 5e-09
Identities = 42/147 (28%), Positives = 76/147 (51%), Gaps = 8/147 (5%)
Frame = +1
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIV--KEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTP 62
RR+ + +D S+ A W + N+ + D+LIL+ ++P + ++ TGS +
Sbjct: 88 RRIMVTVDEGDESMYALSWCLKNLAFQNDKDHLILLYVKPPRVVYSA---FDGTGSFVVT 258
Query: 63 LGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVV------VLVKVYWGDAREKLCEAIE 116
F +SD+ E + ++A +E+ K + V VK GD R+ +C+ ++
Sbjct: 259 RYLF-SSDITATMERVSQ----QVAEGVLERAKGLCNNVENVEVKAESGDPRDVICQMVQ 423
Query: 117 QVPLDGLTMGNRGLGTLRRAIMGSVSN 143
+ +D L MG+ G G ++RA +GSVSN
Sbjct: 424 KWGVDVLVMGSHGYGVIKRAFLGSVSN 504
>AW719194
Length = 526
Score = 52.8 bits (125), Expect = 3e-08
Identities = 27/81 (33%), Positives = 43/81 (52%), Gaps = 1/81 (1%)
Frame = +1
Query: 3 SAR-RLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLT 61
SAR ++G+A DFS S A +W ++N+ +GD +I + + +W +GSPL
Sbjct: 292 SARGKVGVATDFSKSSNSALKWAIENMADKGDTFYIIHVMS---DGSRTNIWAKSGSPLI 462
Query: 62 PLGEFINSDLPKKYEIKTDPE 82
PL + Y ++TDPE
Sbjct: 463 PLSILRQPEAMSNYGVQTDPE 525
>TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome 5, BAC
clone:F24B18, partial (24%)
Length = 691
Score = 50.8 bits (120), Expect = 1e-07
Identities = 24/55 (43%), Positives = 37/55 (66%), Gaps = 3/55 (5%)
Frame = +1
Query: 106 DAREKLCEAIEQVPLDGLTMGNRGLGTLRRAI---MGSVSNYVVNNASCPVTVVK 157
D +E+LC +E++ L + MG+RG G +RR +GSVS+Y V++ CPV VV+
Sbjct: 4 DMKERLCLEVERLGLSAVIMGSRGFGAVRRGSDGKLGSVSDYCVHHCVCPVVVVR 168
>TC16740 similar to UP|Q9SIJ8 (Q9SIJ8) Expressed protein (RD2 protein),
partial (90%)
Length = 765
Score = 48.9 bits (115), Expect = 4e-07
Identities = 38/154 (24%), Positives = 64/154 (40%), Gaps = 2/154 (1%)
Frame = +1
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R + +A+D P S AF W + ++ + D + LI
Sbjct: 178 RDIVLAIDHGPNSKHAFDWALIHLCRLADTIHLI-------------------------- 279
Query: 65 EFINSDLPKKYEIKTDPEVL-KIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGL 123
SD+ + T ++ K+A A E V + ++ GDA + +C E++ +
Sbjct: 280 -HAVSDVKNQLVYDTTQGLMEKLAVEAFEVAMVKTVARIVEGDAGKVICNEAERIKPAAV 456
Query: 124 TMGNRGLGTLRRAIMGSVSNYVVNNA-SCPVTVV 156
MG RG ++ + GSV Y V+N S PV +V
Sbjct: 457 VMGTRGRSLIQSVLQGSVGEYCVHNCKSAPVVIV 558
>TC11721
Length = 327
Score = 35.8 bits (81), Expect = 0.004
Identities = 14/27 (51%), Positives = 22/27 (80%)
Frame = +1
Query: 135 RAIMGSVSNYVVNNASCPVTVVKSSGQ 161
R +MGSVSN+++ +A+CPV +V+ S Q
Sbjct: 115 RILMGSVSNFLMIHATCPVAIVRDSSQ 195
>TC16164 similar to GB|AAN28779.1|23505839|AY143840 At4g27320/M4I22_130
{Arabidopsis thaliana;}, partial (39%)
Length = 589
Score = 35.0 bits (79), Expect = 0.006
Identities = 14/38 (36%), Positives = 25/38 (64%)
Frame = +1
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRP 42
R++G+A+D S S A +W V + ++ GD +IL+ + P
Sbjct: 238 RKIGVAVDLSDESAYAVRWAVQHYIRPGDAVILLHVSP 351
>TC8237
Length = 929
Score = 33.1 bits (74), Expect = 0.024
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +3
Query: 135 RAIMGSVSNYVVNNASCPVTVVK 157
RA +GSVSN+ N CPV +VK
Sbjct: 597 RAFLGSVSNHCAQNVKCPVVIVK 665
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,488,798
Number of Sequences: 28460
Number of extensions: 26020
Number of successful extensions: 133
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 128
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 129
length of query: 163
length of database: 4,897,600
effective HSP length: 83
effective length of query: 80
effective length of database: 2,535,420
effective search space: 202833600
effective search space used: 202833600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144539.1


