

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
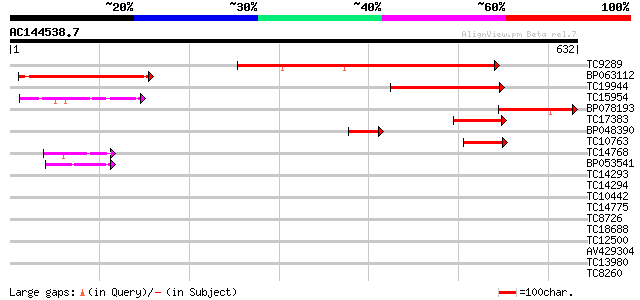
Query= AC144538.7 + phase: 1 /pseudo
(632 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9289 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MR... 281 2e-76
BP063112 258 2e-69
TC19944 similar to UP|Q9LUT8 (Q9LUT8) Genomic DNA, chromosome 3,... 184 3e-47
TC15954 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/M... 111 3e-25
BP078193 106 1e-23
TC17383 92 2e-19
BP048390 72 4e-13
TC10763 weakly similar to UP|Q94A30 (Q94A30) At1g55500/T5A14_10,... 68 5e-12
TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 52 2e-07
BP053541 42 2e-04
TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 39 0.003
TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 39 0.003
TC10442 37 0.007
TC14775 PIR|S14181|S14181 DNA-directed RNA polymerase largest c... 36 0.016
TC8726 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Ara... 33 0.11
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 33 0.14
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 32 0.24
AV429304 32 0.31
TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.3... 32 0.40
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 32 0.40
>TC9289 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (36%)
Length = 994
Score = 281 bits (719), Expect = 2e-76
Identities = 149/314 (47%), Positives = 204/314 (64%), Gaps = 22/314 (7%)
Frame = +3
Query: 255 RSFGPSAG-MASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS--------AND 305
+S+ P++ M H R + G++ R Y + +G + SG+ +N
Sbjct: 51 QSYRPNSQFMGLHHPRPMPAMGATHGFINRMYPNKLYGQYGNTVRSGMGYGTQYDSRSNG 230
Query: 306 RSFLSLENS-RRHGRETASFCRCNGTLDILSEQNRGPRASKLKNH--ISSENNAIDGSK- 361
R++L+++N + GR F N D L+E NRGPRA KN + A+ G
Sbjct: 231 RAWLAVDNKYKTRGRSGGYFGYGNENADGLNELNRGPRAKGGKNQKGFAPTILAVKGQNL 410
Query: 362 -NNASTAKFQD--------ESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTP 412
N +T + +D E N+ DF+ ++ DAKFFVIKSYSED++HKSIKY VWAST
Sbjct: 411 PANLTTEEEKDKTSNIPDREQYNKADFSEEYTDAKFFVIKSYSEDDIHKSIKYNVWASTQ 590
Query: 413 NGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSG 472
NGN+KLDAAY +A++K C +FLFFSVN S QF G+AEM GPV+F+KS+++WQQDKW+G
Sbjct: 591 NGNKKLDAAYQEAQQKPGGCPVFLFFSVNTSGQFVGLAEMTGPVDFNKSLEYWQQDKWNG 770
Query: 473 QFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSI 532
FP+KWHI+KDVPN+ RHI L+NN+NKPVTNSRDTQE+ L+ G++++ IFK Y + T I
Sbjct: 771 CFPLKWHIVKDVPNNLLRHITLDNNENKPVTNSRDTQEIMLEPGLKLVKIFKEYSSKTCI 950
Query: 533 LDDFDFYEDRQKAM 546
LDDF FYE RQK +
Sbjct: 951 LDDFGFYEARQKTI 992
>BP063112
Length = 530
Score = 258 bits (660), Expect = 2e-69
Identities = 123/152 (80%), Positives = 135/152 (87%), Gaps = 1/152 (0%)
Frame = +2
Query: 10 PSIPESSKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDN 69
P+ P++ K QGT TIGHSR+ TSQS SFGSG D PLY PNVYAPQAQAFYY GFDN
Sbjct: 89 PAEPDNLK----DQGTATIGHSRETTSQSESFGSGGDCPLYSPNVYAPQAQAFYYRGFDN 256
Query: 70 GNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGG 129
GNGEWDEYPSYVN+GG+E+GSPGVYNENQSLVFHSGYGFNPQ+PYGPYSPVT+PLPSVGG
Sbjct: 257 GNGEWDEYPSYVNSGGVELGSPGVYNENQSLVFHSGYGFNPQLPYGPYSPVTTPLPSVGG 436
Query: 130 DAQLYSPQQFPYT-PPYYNQLVPPPNLPYLNS 160
DAQLYS QQFP+T PPYY+QLV PP+LPYLNS
Sbjct: 437 DAQLYSHQQFPFTVPPYYHQLV-PPSLPYLNS 529
>TC19944 similar to UP|Q9LUT8 (Q9LUT8) Genomic DNA, chromosome 3, P1 clone:
MGD8, partial (8%)
Length = 611
Score = 184 bits (468), Expect = 3e-47
Identities = 83/127 (65%), Positives = 104/127 (81%)
Frame = +2
Query: 425 AKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDV 484
A K C IFLFFSVNAS QFCGVAEMVGPV+F+K +DFWQQDKW+G F +KWHI+KDV
Sbjct: 5 AAGKSGGCPIFLFFSVNASGQFCGVAEMVGPVDFNKDMDFWQQDKWNGSFLLKWHIVKDV 184
Query: 485 PNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQK 544
PN RHI+LENN+NKPVTNSRDTQE+ ++G+EML +FK++ TS+LDDF +YE+RQK
Sbjct: 185 PNGGCRHIILENNENKPVTNSRDTQEITYRKGLEMLRVFKSHTLKTSLLDDFIYYENRQK 364
Query: 545 AMQERKA 551
+ R++
Sbjct: 365 SHGARES 385
>TC15954 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (7%)
Length = 770
Score = 111 bits (278), Expect = 3e-25
Identities = 65/158 (41%), Positives = 86/158 (54%), Gaps = 18/158 (11%)
Frame = +2
Query: 12 IPE-SSKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPL---------YPPNVYAPQAQ- 60
IPE + K S + GT G++ + Q S+ L Y PN YA A
Sbjct: 296 IPEPTKKVSGNQYGTVDSGNAAN--GQIPSYERSVTPVLQDFIDPTMCYVPNGYASTAYY 469
Query: 61 -------AFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP 113
A+YYGG+D EWD+Y YVN G+++ S GVY +N SL++H GYG+ P
Sbjct: 470 YGGESMGAYYYGGYDGTGNEWDDYSRYVNQEGLDMTS-GVYGDNGSLLYHHGYGY---AP 637
Query: 114 YGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVP 151
YGPYSP SP+P++G D QLY PQ + Y PPY+ L P
Sbjct: 638 YGPYSPAGSPVPTMGNDGQLYGPQHYQY-PPYFQPLTP 748
>BP078193
Length = 433
Score = 106 bits (264), Expect = 1e-23
Identities = 58/90 (64%), Positives = 69/90 (76%), Gaps = 3/90 (3%)
Frame = -2
Query: 546 MQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNNEVIA---A 602
MQERK RQQSS++ GLVG +EHRSS++ST DFIKQ+ K+F+LVVRLD+NNN+ I A
Sbjct: 432 MQERKVRQQSSLVAAGLVGENEHRSSANSTNDFIKQLPKSFALVVRLDENNNKEITAANA 253
Query: 603 NRDKLASDVPTGNVFKPGEGILVTASSMQT 632
NRD LASD G V KP + I VTASS QT
Sbjct: 252 NRDNLASDGLIGKVIKPEDVISVTASSTQT 163
>TC17383
Length = 733
Score = 92.4 bits (228), Expect = 2e-19
Identities = 43/59 (72%), Positives = 48/59 (80%)
Frame = +2
Query: 495 ENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQ 553
ENND+KPVTNSRDTQEV QG+EML IFKNY TSILDDF+FYE RQK +QE+K Q
Sbjct: 2 ENNDHKPVTNSRDTQEVSFPQGVEMLNIFKNYAARTSILDDFEFYESRQKXIQEKKIXQ 178
>BP048390
Length = 126
Score = 71.6 bits (174), Expect = 4e-13
Identities = 30/39 (76%), Positives = 36/39 (91%)
Frame = -1
Query: 378 DFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNR 416
DF+ ++ DAKFFVIKSYSED++HKSIKY VWASTPNGN+
Sbjct: 123 DFSDNYSDAKFFVIKSYSEDDIHKSIKYSVWASTPNGNK 7
>TC10763 weakly similar to UP|Q94A30 (Q94A30) At1g55500/T5A14_10, partial
(10%)
Length = 544
Score = 67.8 bits (164), Expect = 5e-12
Identities = 30/49 (61%), Positives = 41/49 (83%)
Frame = +3
Query: 506 RDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQ 554
RDTQEVK ++GI+++ IFK + + TSILDDF FYE R+KA QERK+++Q
Sbjct: 3 RDTQEVKFEKGIQIIKIFKEHSSKTSILDDFGFYEAREKATQERKSKEQ 149
>TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (21%)
Length = 1036
Score = 52.4 bits (124), Expect = 2e-07
Identities = 29/85 (34%), Positives = 47/85 (55%), Gaps = 4/85 (4%)
Frame = -2
Query: 38 SGSFGSGADQPLYPPNVYAPQ---AQAFYYGGFDNGNGEWDEYPSYVN-NGGIEIGSPGV 93
S G +PL P + P + +YYGG+D G +W+ Y Y+N +GG+ + GV
Sbjct: 234 SNGTAKGMTKPLTPNPSFVPNGYPSTTYYYGGYD-GQNDWNVYSRYMNLDGGM---AQGV 67
Query: 94 YNENQSLVFHSGYGFNPQMPYGPYS 118
Y ++ S +++ GYG+ PYG Y+
Sbjct: 66 YGDSCSYMYNQGYGYT---PYGAYA 1
>BP053541
Length = 500
Score = 42.4 bits (98), Expect = 2e-04
Identities = 27/81 (33%), Positives = 41/81 (50%), Gaps = 3/81 (3%)
Frame = +1
Query: 41 FGSGADQPLYPPNVYAPQAQ--AFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQ 98
F GA + + N+Y P A +Y G++ GEW+++ G +I G NEN
Sbjct: 262 FNEGAPEFVVDQNLYYPAATNYEYYCTGYETP-GEWEDHHRIXGLDGPDIQYTGAQNENF 438
Query: 99 SLVFHS-GYGFNPQMPYGPYS 118
V+++ YGF Q PY PY+
Sbjct: 439 PYVYYTPSYGF-AQSPYNPYN 498
>TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (18%)
Length = 540
Score = 38.9 bits (89), Expect = 0.003
Identities = 32/110 (29%), Positives = 42/110 (38%), Gaps = 5/110 (4%)
Frame = +2
Query: 104 SGYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTP 163
SGYG+ PY PY P S G PQ + PP PPP+ Y P P
Sbjct: 125 SGYGYGAPPPYQPYG-AAPPSQSYGAPPP---PQPYGAAPPSQPYGAPPPSQSY-GGPPP 289
Query: 164 VSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARG-----SFPVAPGSF 208
SQP + G Q + + P P S + G +P P ++
Sbjct: 290 PSQPYSASPYG---QPSAPYAAPYQKPPKDESHSHGGGGGSGYPPPPSAY 430
>TC14294 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (65%)
Length = 1192
Score = 38.5 bits (88), Expect = 0.003
Identities = 32/115 (27%), Positives = 43/115 (36%), Gaps = 10/115 (8%)
Frame = +1
Query: 104 SGYGFNPQMPYGPYSPVTSPLP-SVGGDAQLYS----PQQFPYTPPYYNQLVPPPNLPYL 158
SGYG+ PY PY P +Q Y PQ + PP PPP+ Y
Sbjct: 85 SGYGYGAPPPYQPYGAAPPSQPYGAAPPSQSYGAPPPPQPYGAAPPSQPYGAPPPSQSY- 261
Query: 159 NSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARG-----SFPVAPGSF 208
P P SQP + G Q + + P P S + G +P P ++
Sbjct: 262 GGPPPPSQPYSASPYG---QPSAPYAAPYQKPPKDESHSHGGGGGSGYPPPPSAY 417
>TC10442
Length = 564
Score = 37.4 bits (85), Expect = 0.007
Identities = 16/38 (42%), Positives = 25/38 (65%)
Frame = +3
Query: 531 SILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGGSEH 568
S+LDDFDFYE+R+K + +K+ + +S P G+ H
Sbjct: 3 SLLDDFDFYENREKLFRSQKSTKHTSPKPQVYNSGNYH 116
>TC14775 PIR|S14181|S14181 DNA-directed RNA polymerase largest chain
(isoform B1) - soybean (fragment) {Glycine
max;} , partial (22%)
Length = 848
Score = 36.2 bits (82), Expect = 0.016
Identities = 36/138 (26%), Positives = 51/138 (36%), Gaps = 5/138 (3%)
Frame = +1
Query: 108 FNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTP--PYYNQLVP--PPNLPYLNSPTP 163
++P + Y P SP SP + YSP Y+P P Y+ P P+ PY + +P
Sbjct: 133 YSPSLAYSPSSPRLSPSSPYSPTSPNYSPTSPSYSPTSPSYSPSSPTYSPSSPYNSGVSP 312
Query: 164 VSQPELTNLLGIDQQVESNFFGPRASY-PSVGSFARGSFPVAPGSFSFHESQQGFDGSRS 222
D S F P Y PS + +P S S + Q R
Sbjct: 313 ------------DYSPSSPQFSPSTGYSPSQPGY-------SPSSTSQYTPQTNDKDERG 435
Query: 223 GGLWSDSSKPSERQRSFM 240
W S + E +RS +
Sbjct: 436 KD*WRRSMRQFEARRSIL 489
>TC8726 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Arabidopsis
thaliana;}, partial (95%)
Length = 820
Score = 33.5 bits (75), Expect = 0.11
Identities = 23/59 (38%), Positives = 24/59 (39%)
Frame = +2
Query: 105 GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTP 163
GYG + P G Y P P P Y PQQ PPY Q PPP P P P
Sbjct: 173 GYGKDGYPPQG-YPPQGYPPPG-------YPPQQGYPPPPYPAQGYPPPYAPQYAQPPP 325
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 33.1 bits (74), Expect = 0.14
Identities = 33/128 (25%), Positives = 48/128 (36%), Gaps = 4/128 (3%)
Frame = +3
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
P P +S P P G A +P TPP ++ PP+ NS S P
Sbjct: 162 PPSPEERFSGTPPPPPPATGSASTATP-----TPPTSSRSASPPSPSPANSHMASSPPSP 326
Query: 170 TNLLGIDQQVESNFFGPRASYP----SVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGL 225
T+ L + S P S P + + +R S PV+ FH S + S
Sbjct: 327 TSALSVSVSTPSPAHSPPTSPPAPRSATSTSSRISSPVSSRRL-FHVSP-----ALSAST 488
Query: 226 WSDSSKPS 233
W ++ P+
Sbjct: 489 WPPTTSPA 512
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 32.3 bits (72), Expect = 0.24
Identities = 27/80 (33%), Positives = 33/80 (40%), Gaps = 17/80 (21%)
Frame = +1
Query: 110 PQMPYGPYSPVTSPLPS--------VGGDAQ----LYSPQQFPYTPPYYNQLVPPPNLPY 157
P P P P T+P P+ + G+A L SP Q P PP Y PP+ P
Sbjct: 196 PPQPPPPSLPPTTPPPASVIAGSAIMSGEASFPCSLPSPPQPPPPPPKYGVTTIPPSPPL 375
Query: 158 LNS-----PTPVSQPELTNL 172
L S P P LT+L
Sbjct: 376 LMSATDRAPLPPPSTHLTSL 435
>AV429304
Length = 355
Score = 32.0 bits (71), Expect = 0.31
Identities = 26/74 (35%), Positives = 32/74 (43%), Gaps = 16/74 (21%)
Frame = +3
Query: 110 PQMPYGPYSPVTSPLPS-VGGDAQLYSPQQFPYTP----------PYY---NQLVPP--P 153
PQ P+ PY P+ PS V + SP Q P+ P P Y N PP P
Sbjct: 45 PQTPHTPYVPIPPKTPSPVYSPPNVPSPPQTPHAPYVPTPPKTPSPVYSPPNVPSPPQTP 224
Query: 154 NLPYLNSPTPVSQP 167
+ PY+ PTP P
Sbjct: 225 HAPYV--PTPSKTP 260
>TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.33), partial
(12%)
Length = 565
Score = 31.6 bits (70), Expect = 0.40
Identities = 21/55 (38%), Positives = 24/55 (43%), Gaps = 1/55 (1%)
Frame = +2
Query: 116 PYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPT-PVSQPEL 169
P SP TS P+ +P P+ PP PPP P SPT P S P L
Sbjct: 332 PCSPPTSSSPTGSSSHSTSNP---PHPPPQQQPFSPPPPPPPPPSPTAPSSHPRL 487
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 31.6 bits (70), Expect = 0.40
Identities = 23/73 (31%), Positives = 29/73 (39%), Gaps = 8/73 (10%)
Frame = +3
Query: 103 HSGYGFNPQMP-----YGPYSPVTSPLPSVGGDAQLYSPQQFPYTPP---YYNQLVPPPN 154
H Y P P + P PV SP P +Y PY P Y++ PPP+
Sbjct: 105 HYAYSSPPPPPKPYSYHSPPPPVHSPPPPYEKPHPVYHSPPPPYEKPHPVYHSPPPPPPH 284
Query: 155 LPYLNSPTPVSQP 167
PY P+P P
Sbjct: 285 KPY-KYPSPPPPP 320
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.132 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,703,765
Number of Sequences: 28460
Number of extensions: 166114
Number of successful extensions: 1069
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 950
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1014
length of query: 632
length of database: 4,897,600
effective HSP length: 96
effective length of query: 536
effective length of database: 2,165,440
effective search space: 1160675840
effective search space used: 1160675840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144538.7


